INTRODUCTION
Tooth decay is still the most prevalent chronic disease in humans.1 Its frequency steadily increases with age, affecting almost 100% of the Chilean population aged 65 to 70 years; it affects 85% school-age children, in Chile.2)(3)(4
This disease has a multifactorial etiology, which is microbiologically induced and associated to the fermentation of carbon hydrates. It is defined as an imbalance of proteins and minerals in teeth, with irreversible loss of minerals when lesions are in their cavitated state. Factors such as time, frequency, and type of food intake, deficiencies in techniques of dental biofilm control, low levels of fluoride in the oral environment, and low rate of salivary secretion are some of the ecological determinants influencing variations in biofilm.5
The oral biofilm in caries-free patients contains more than one thousand different species; however, patients with active caries have a reduction of more than 70% of species, with dominance of the acidogenic microbiota associated with the onset and progression of the disease.5) The bacterium Streptococcus mutans is one of the species traditionally responsible for the onset of lesions, along with other acid-producing species, such as Streptococcus sobrinus, Veillonella and Lactobacillus spp.( 6)(7)(8 Other alkalis-producing microbiota has been identified as it influences pH balance and the oral microbiome, such as Streptococcus sanguinis, Streptococcus gordonii, Streptococcus salivarius and A naeslundii, to name just a few.9)(10)(11
The deleterious activity of S. mutans spp. is associated with its ability to produce lactic acid from sucrose, reaching levels capable of causing the loss of minerals in teeth.7)(12)(13)(14
Streptococcus is the most abundant bacterial genus in the oral cavity. According to its phylogeny, it is classified into three groups (bovis, pneumonia and viridans),15 and the viridans group is divided into five groups: mitis, mutans, salivarius, anginosus, and sanguinis.15 The Mutans Streptococci group (MS) is classified, according to the nucleotide sequence of its 16S ribosomal RNA, in eight species (mutans, sobrinus, cricetus, dawnei, ferus, macacae, ratti, and hyovaginalis); of these, S. mutans is considered as the predominant species in the oral cavity.16
S. mutans is a Gram-positive coccus which groups itself in chains; it is motionless, catalase negative, a fast producer of lactic acid, and its usual habitat is the dental surface, although it has also been detected in heart valves and in atheromatous plaques of large blood vessels.17
The virulence of S. mutans is characterized by its acidogenesis, aciduria, acidophile, synthesis of extracellular polysaccharides (EPS), synthesis of intracellular polysaccharides, synthesis of glucan-binding proteins (Gpbs), adhesins, wall-associated protein A (WapA), and bacteriocins. The synthesis of extracellular polysaccharides is a prominent factor of its virulence, since even before the sucrose enters the cell, a percentage is transformed by exoenzymes, glucosyltransferases (Gtfs) and fructosiltransferasas (Ftf), which metabolize sucrose in monosaccharide molecules, transferring each fraction to a receiving molecule and forming polymers that can be spread in the biofilm or remain associated with S. mutans.18)(19
S. mutans has three different types of gtf genes expressing glucosyltransferase activity. Genes gtfB and gtfC share a same operon, encoding enzymes GtfB and GtfC as well as gene gtfD, which is not in the gftBC locus, encoding for enzyme GtfD. These enzymes (Gtfs) condition the aggressive role of biofilm through the production of glucans from sucrose, which adhere to the enamel, providing sites of colonization and insoluble matrix.16) GtfB synthesizes insoluble glucans that adhere to the dental surface. GtfC produces soluble and non-soluble glucans that are easily metabolizable. S. mutans produces three different enzymes to act on the same substrate to form polysaccharides; this fact allows S. mutans to contribute to the formation and composition of cariogenic biofilm, unlike other oral microorganisms.18)(20
Traditionally, high counts of S. mutans in saliva and dental biofilm have been linked to carious activity. However, while this evidence has been challenged over the past years, there are numerous reports claiming that higher counts of this bacteria are detected in individuals with caries, and thus the count of S. mutans has been used as an indicator of cariogenic risk.21)(22)(23
In the past, quantification was conducted through traditional cultivation, but technological advances have demonstrated that real-time polymerase chain reaction (PCR) can be a sensitive, quick methodology, revolutionizing the field of quantification methods.24)(25)(26)(27)(28 Unlike quantification by cultivation, this fluorescence-based technique 15)(29 does not require trained staff to detect colonies compatible with S. mutans, which needs an additionally step of species confirmation, which can be done through biochemical or molecular tests; this arduous process of cultivation, isolation, and identification requires more time and resources, and compared against quantification by qPCR is faster, inexpensive, accurate, and sensitive; its only drawback is that it does not discriminate quantification between living and dead cells.24)(30)(31
On the other hand, this improved technique of conventional PCR, which is a qualitative test, allows conducting qualitative and quantitative measurements of specific genes in a sample of nucleic acid. An example of this is SYBR-Green, a fluorometric detection method designed to amplify a DNA sequence not exceeding 250 pb,24) where the reagent non-specifically binds to a double DNA strand producing quantitative fluorescence proportional to the initial number of molecules,29)(32)(33)(34)(35) resulting in an accurate, sensitive, and fast quantification.
The use of molecular biology in cariology is intended to develop tools for the early detection of risk factors that destabilize the oral ecosystem, in order to implement preventive approaches and to establish protective measures to prevent the appearance of carious lesions and their possible transmission.
Based on this information, the objective of this research project was to implement the methodology of real-time qPCR to detect and quantify Streptococcus mutans spp. in frozen samples of saliva and dental biofilm from children 8 years of age, based on the amplification of a fragment of the gtfB gene.
MATERIALS AND METHODS
The protocol of this research project was approved by the Scientific Ethical Committee of Universidad de Chile School of Dentistry, through Affidavit 012-13 W.
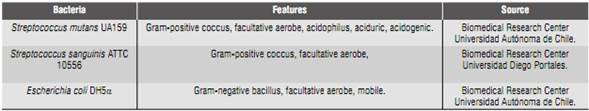
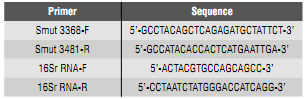
This study used the bacterial strains described in Table 1, and the amplification primers for both conventional PCR (end-point) and qPCR as described in Table 2.
Table 2 Sequences of primers used for PCR and qPCR, previously reported by Yoshida et al36 ) and Gordan et al.9
Sample
This study involved a random selection (Random Numbers Generation, Microsoft 2007, Seattle, Washington, USA) of 27 children of both sexes aged 8 years, from local schools in the northern metropolitan area of Santiago, Chile, who met the following criteria:
Inclusion criteria: a) 8 years of age, b) systemically healthy, c) with no contraindications to participate in a dental examination.
Exclusion criteria: a) patients with reduced salivary flow, b) patients with systemic diseases preventing them from dental treatment, c) patients who have been on antibiotic therapy, with steroids or mouthwash, during the past three months.
The parents of children meeting the inclusion criteria signed an informed consent form accepting their children’s participation in the study.
Sample collection and processing
The examined children were required to refrain from any oral hygiene procedure and food intake during the 12 hours prior to sampling.9
Saliva collection: 5 mL of non-stimulated saliva were collected in the morning in a 15 mL sterile plastic tube, which was maintained at 4 °C until brought to the Chemistry Laboratory (School of Dentistry, Universidad de Chile). The saliva was transferred to 1.5 mL centrifuge tubes for further analysis in the Microbiology Laboratory (School of Dentistry, Universidad de Chile).
Collection of supragingival dental biofilm: biofilm was collected from all the mesio-vestibular surfaces of temporary and permanent anterior and posterior teeth, using sterile curettes (GR 4/5, Gracey finishing curettes, Ransom and Randolph, Toledo, OH, USA). This pool was brought to 1.5 mL sterile centrifugal tubes containing 500 µL of K2HPO4 10 mM solution (pH 7) and stored at 4 °C until being transferred to the laboratory.
Conservation of samples: the saliva and dental biofilm samples adequately labeled were stored at -80°C for further processing.
Extraction of genomic DNA from saliva and biofilm
DNA extraction was conducted with the MasterPureTM DNA purification system, Epicentre® (Ilumina Co, San Diego, CA, USA). The samples of saliva and dental biofilm were defrosted to room temperature and re-suspended by agitation in vortex (Cole-Parmer Vortex Mixer, Vernon Hills, IL, USA) for 30 sec. 150 µL of saliva or dental biofilm were put in 1.5 mL centrifuge tubes, adding 150 µL of solution (2lx) T & C (MasterPureTM) and 5 µL of lysozyme (20 mg/mL) (Sigma-Aldrich, St. Louis, MO, USA) and incubating for 1 hour at 37 °C. Then, 1µL of proteinase K was added (50 mg/mL) (Sigma-Aldrich, St. Louis, MO, USA), incubating at 65 °C for 1 hour and stirring every 5 min; 2 µL of RNase were added (5 mg/mL) (Sigma-Aldrich, St. Louis, MO, USA), incubating for 5 min at 37 °C. The tubes were later deposited on ice for 3 to 5 min before adding 150 µL of MPC buffer (MasterPureTM), stirring in vortex for 10 sec and centrifuging to 13,000 rpm for 10 min at 37 °C; 400 µL of the supernatant were transferred to a new tube. 400 µL of pure isopropanol were added, mixing by inversion and centrifuging at 13,000 rpm for 10 min at 4 °C. Then the isopropanol was removed, rinsing with 150 µL of ethanol (75%), centrifuging again at 13,000 rpm for 5 min at 4 °C. The precipitate was left to dry at room temperature and rehydrated in 45 µL ultrapure sterile water. The obtained DNA was stored at -20 °C.
Extraction of genomic DNA of Streptococcus mutans UA159
To obtain a standard quantification curve, Streptococcus mutans UA159 was cultivated. This bacterium grew in microaerophilic conditions at 37 °C in TSB broth (Oxoid, Hampshire, UK) for 48 hours. 3 mL of culture were centrifuged at 13,000 rpm for 5 min. The supernatant was removed adding 300 µL of T & C solution (MasterPure™ system). The protocol continued as described above.
Extraction of genomic DNA of S. sanguinis
Genomic DNA of Streptococcus sanguinis ATTC10556 strain was used as control of specificity of primers in the PCR reactions. This bacterium grew in microaerophilic conditions at 37 °C in TSB broth (Oxoid) for 48 hours, and genomic DNA of S. sanguinis was extracted using an identical procedure as described for S. mutans.
Extraction of genomic DNA of E. coli
Genomic DNA of Escherichia coli DH5α strain was used as negative control for the PCR reactions. This strain was obtained from a culture of E. coli in Mc Conkey agar (Becton Dickinson®, NJ, USA). A culture was selected and inoculated in 5 mL of nutrient broth, and grown by shaking at 37 °C for 12 hours. Subsequently, using 3 mL of culture, genomic DNA was extracted by following the protocol described for S. mutans.
Separation of DNA by electrophoresis
The analysis of integrity of DNA samples was made by electrophoretic separation (Major Science MJ 105S/MT108, Saratoga, CA, USA) in 1% agarose gel using TAE 0,5X buffer (Winkler, Lampa, Santiago, Chile). Once completed, it was stained with ethidium bromide for 5-10 min observing in UV transilluminator at 254 nm.
Verification of primers in end-point PCR
To verify that the amplified DNA corresponded to bacterial DNA, end-point PCR was conducted with universal primers for bacteria 16Sr RNA (table 2), which produce a fragment of DNA of 296-350 pb.8
This procedure implied making a master mix for PCR (5 µL of reaction buffer 10X, 4 µL of MgCl2 25 mM, 1 µL of dNTPs 10 mM each, 1µL of primer 16Sr RNA-F 25 pmol/µL, 1µL of primer 16Sr RNA-R 25 pmol/µL, 33.5 µL ultrapure water, 0.5 µL Taq polymerase 5UI/µL (Thermo Fisher Scientific, Waltham, MA, USA) and 4 µL of purified DNA: genomic DNA of S. mutans, S. sanguinis, E. coli, saliva and dental biofilm). The PCR reactions were performed in thermal cycler multigene gradient (Labnet® Edison, NJ, USES), using the following plan: 94 °C during 5 min, followed by 40 cycles of 94 °C during 30 sec. 50 °C during 1 min, 72 °C during 1 min, and a final extension at 72 °C during 10 min.
The same procedure was used to determine the specificity of primers for S. mutans (table 2), using the following program: 94 °C for 5 min, followed by 40 cycles of 94 °C for 30 sec, 60 °C for 30 sec, 72 °C for 1 min, and a final extension at 72 °C for 10 min, obtaining a fragment of 114 pb.
Quantification of total DNA of saliva and dental biofilm
To determine the total concentration of DNA of samples of genomic DNA from saliva, dental biofilm, and S. mutans UA159, the quantification was carried out with spectrophotometry by microplates (Take3 Model Epoch, BiotekR, Winooski, VT, USA). 2 µL of each sample were taken in duplicate and the obtained values of total DNA were brought to a final volume of 20 ng/µL.
Quantification of S. mutans using qPCR
For the standard curve, a serial dilution of factor 10 was prepared and 1 µL of DNA of S. mutans UA159 was taken five times, in addition to 9 µL of miliQ water. A master mix was prepared for qPCR reaction (10 µL of Mix 2X SYBR Green (Thermo Fisher Scientific, Waltham, MA, USA), 0.5 µL of primer Smut 3368-F 25 pmol, 0.5 µl of primer Smut 3481-R 25 pmol, 4 µL of miliQ water, and 5 µL of DNA from the samples). 5 µL of miliQ water were used for No Template Controls (NTC).
Each recipient was loaded with 15 µL of master mix plus 5 µL of DNA of S. mutans; the measurements were duplicated for controls (NTC), the construction of the standard curve, and the quantification of each sample. Finally, the plaque was covered with optical sealing, according to the manufacturer's instructions.
DNA amplification was performed in an ECO Real Time thermocycler (Illumina®, SD, Cal, USA), using the following thermal profile: 95 °C for 5 min, followed by 30 cycles of 95 °C during 30 sec., 66 °C during 30 sec., 72° C for 1 min, and a final extension at 72 °C for 10 min.
RESULTS
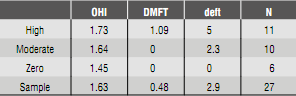
The sample in the present study included 27 children of 8 years whose oral health, expressed in averages, was: average Loë Silness oral hygiene index = 1.45; DMFT = 1.09 and deft = 2.9, of four sites (mesial, distal, vestibular and palatal or lingual) of teeth 1.6, 1.2, 2.4, 3.6, 3.2, and 4.4 (Table 3).
Table 3 Average Oral Hygiene Index, DMFT and deft, sorted out in groups per cariogenic risk index (high, moderate, zero).
The genomic DNA of Streptococcus mutans UA159 showed integrity in 1% agarose gel as shown in Figure 1.
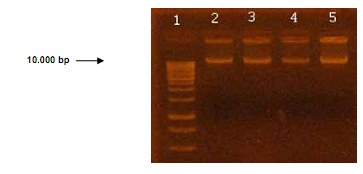
Figure 1 Electrophoresis of genomic DNA of S. mutans UA159 in 1% agarose gel. Lane 1: marker of 1 kb in molecule size. Lanes 2 to 5 show genomic DNA of S. mutans UA159.
The use of universal primers demonstrated the presence and amplification of bacterial DNA by conventional PCR (end-point) with universal primers for 16Sr RNA.
The use of Smut 3368-F and Smut 3481-R primers (Figure 2) showed specificity by obtaining an amplicon of expected size (114 pb),20 since these primers were only detected in the DNA samples of S. mutans UA159 and in DNA from saliva and dental biofilm.
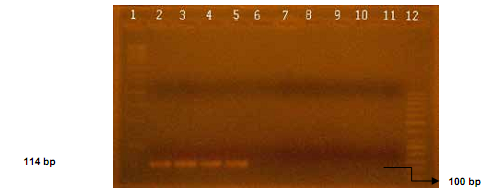
Figure 2 Electrophoretic separation of a fragment of 114 bp of gene gtfB in 2% agarose gel amplified by means of PCR. Lane 1 corresponds to a standard molecular size of 1 kb. Lanes 2 and 3 correspond to amplification using DNA of S. mutans UA159. Lanes 4 and 5 correspond to amplifications using DNA from saliva and dental biofilm samples (S C6, C6 BF). Lanes 6 and 7 correspond to amplifications using DNA of S. sanguinis. Lanes 8 to 10 correspond to amplifications using DNA from E. coli. Lane 11 corresponds to amplification using ultrapure water (miliQ). Lane 12 corresponds to a standard molecular size of 50 bp.
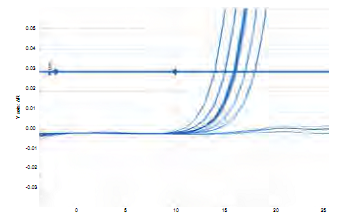
The standard curve was designed based on the dilution of DNA isolated from S. mutans UA159, whose initial concentration was 128 ng/µL (figure 8). The melting curve for the amplification of DNA of S. mutans UA159 (Figure 3) shows a single amplification pick, to the same temperature for all the dilutions. Duplicates issue fluorescence, achieving the quantification of samples with a delta of 3.36; i.e. between each amplification cycle in which fluorescence is detected there is a difference of 3.36 cycles (Figure 4).
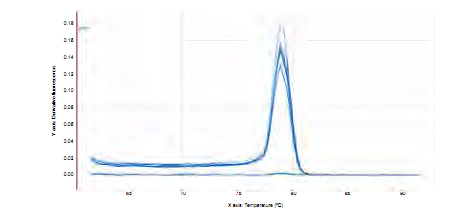
Figure 3 Melting curve for a fragment of 114 bp of gtfB gene. Note the only amplification temperature of serial dilutions of DNA of S. mutans UA159.
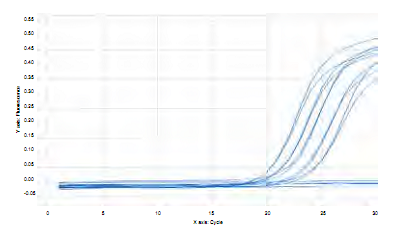
Figure 4 Graph of concentration per amplification cycle for a fragment of 114 bp of gtfB gene using serial dilutions of genomic DNA of S. mutans UA159 .
Quantification of S. mutans in saliva and dental biofilm samples by means of qPCR
After isolating the total DNA of samples from the 27 patients came the analysis of DNA integrity of saliva and purified biofilm. Figure 5 corresponds to a representative gel of integrity analysis of DNA extracted from samples of saliva and dental biofilm. The electrophoretic separation showed the presence of DNA bands larger than 10,000 bp, with integrity of the isolated DNA and minimal degradation (Figure 5).
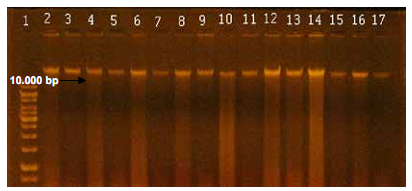

Figure 5 Electrophoretic separation of genomic DNA from saliva and dental biofilm in 1% agarose gel. Lane 1 corresponds to the standard of molecular size of 1 kb. Lanes 2 to 17 correspond to DNA of samples C1 S (saliva), C1 BF (biofilm), C2 S (saliva), C2 BF, C3 S, C3 BF, C4 S, C4 BF, C5 S, C5 BF, C6 S, C6 BF, C7 S, C7 BF, C8 S and C8 BF, respectively.
The values of total DNA concentrations were distributed ranging from 633 to 25 ng/μL for saliva and 406 to 34 ng/μL for dental biofilm. To standardize the initial concentrations of samples before conducting qPCR, the obtained values were diluted to a standardized final concentration of 20 ng/µL per sample.
The melting curve achieved in identifying gene gtfB of S. mutans in saliva and biofilm samples in the 27 patients showed a single pick at a same amplification temperature for all samples (Figure 6). Amplification of each duplicate is similar, achieving a quantification of samples with a delta of 3.36 in difference between each cycle, and an optimal repeatability of duplicates (Figure 7).
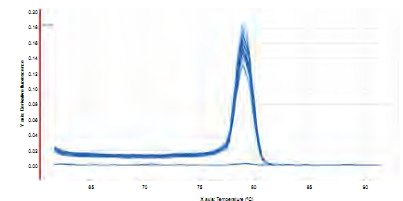
Figure 6 Melting curve for the amplification of a fragment of 114 bp of the gtfB gene of S. mutans in saliva and dental biofilm samples from 27 children of 8 years of age .
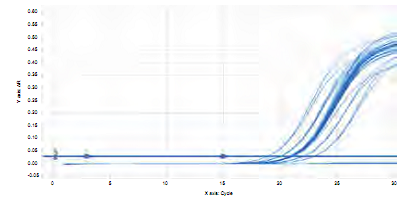
Figure 7 Graph of concentration per amplification cycle for a fragment of 114 bp of gtfB gene in saliva and dental biofilm samples from 27 children of 8 years of age.
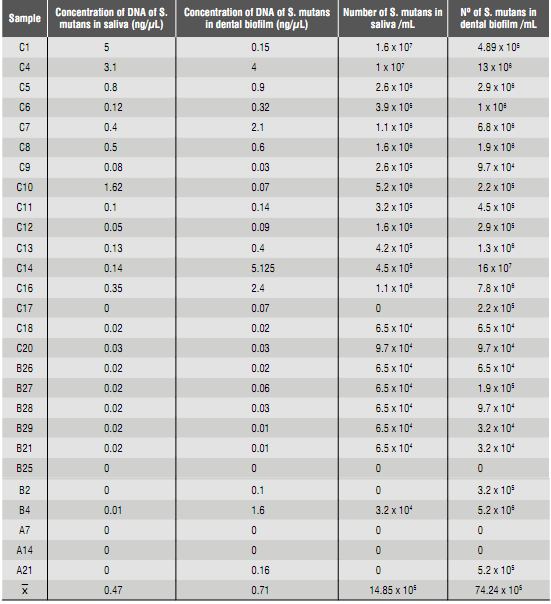
The values obtained by quantifying DNA of S. mutans in saliva and dental biofilm samples by qPCR are shown in table 4. The number of bacteria in mL of saliva and dental biofilm was calculated based on the size of the S mutans genome.35
Table 4 Quantification of S. mutans in saliva and dental biofilm samples from 8-year-old children through the qPCR technique.
The linearity achieved for the standard curve used in quantifying by qPCR (Figure 8) accounts for 98% technique efficiency. On the other hand, the behavior of No Template Controls (NTC) shows fluorescence far below the minimum threshold in which the templated samples were quantified, validating the quantification results obtained with this technique (Figure 9).
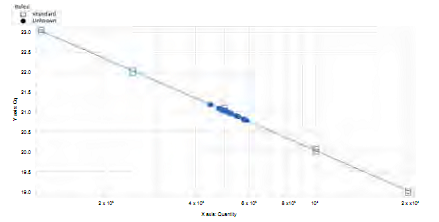
Figure 8 Standard curve. Graph of concentration of DNA starting from the value of cycle of quantification (cq) of the unknown sample.
DISCUSSION
The findings yielded by qPCR amplification showed a high percentage of efficiency and a reasonable delta in between cycles. The analysis of these parameters allows validating the results thanks to the sensitivity and accuracy of this technique, as previously published,24)(25)(26)(27) with similar methodology. It is therefore possible to claim that this methodology identifies and quantifies Streptococcus mutans spp. in saliva and dental biofilm.
The traditional methods for microbial identification and quantification used the method of cultivation on plaque to detect the presence or quantity of S. mutans, which implies processing samples immediately without the possibility of storing them for long periods of times or freezing them for later analysis. In addition, the culture method has limitations that affect accuracy and consistency in evaluating infection by S. mutans. Cultures show certain disadvantages, such as extended time for microbiological processing and a lower level of sensitivity. These limitations make it difficult for traditional culture methods to provide accurate microbial evaluations to identify and quantify S. mutans associated with caries susceptibility in individuals at risk.37
In addition, there is the drawback related to storage or preservation of culture agents, which must ensure viability over time; contamination risks should be minimal ensuring that culture purity remains unaffected.38
The transportation of frozen samples is the preferred method for moving and storing large collections of microbial species. These conditions increase complexity as well as the risk of losing the viability of cells for quantification with the traditional culture technique.39
On the other hand, conventional PCR is a qualitative technique that only allows indicating the presence or absence of a specific fragment of DNA and therefore associating it with presence or absence of a particular microorganism. Instead, qPCR is a quick technique that easily provides qualitative and quantitative results that can come from samples stored for long periods of time.40
Despite being more complex and requiring costly and more advanced equipment in comparison to the traditional method of cultivation in dilution series, once stablished, qPCR results in lower cost and greater speed and accuracy.24) These advantages also allow reducing laboratory costs when patients are requested this test as a complement to determine individual cariogenic risk. However, determining the initial sample mass (collected saliva and bacterial plaque) is an essential stage for establishing bacterial count-a fact that appears as a weakness of the present study-.
The extraction of DNA from saliva and biofilm samples showed some level of degradation, contrasting with the observations in samples of pure culture of S. mutans UA159. This phenomenon may be explained by the diversity and abundance of the oral ecosystem. RNAasa was used to overcome this problem, thus increasing the integrity of samples.
According to Ono et al,41) the white gen is chosen because the spaP gene is described as the one with the greatest sensitivity to detect S mutans. However, a recent study on strains in Chilean population reported the presence of the gtfB gene in 100% of S. mutans strains ,( 45) while only 63% shows the presence of the spaP gene,42)(43)(44) which explains the choice of the gtfB gene for this study. However, this fact minimizes, but does not guarantee, detection defects, due to the possibility of finding polymorphism.
The etiology of dental caries has been traditionally linked to imbalances in dental biofilm;1)(12) however, counts of CFU of S. mutans in saliva are used to define disease risk. This dichotomy used to be based on the ease of volumetric measurement of saliva samples compared to the difficulty in determining the amount of dental biofilm. Currently, molecular techniques allow quantifying such risk in dental biofilm; therefore, it is recommended to obtain biofilm samples, since this is the natural niche of S. mutans in the oral cavity. The present study showed greater amount of S. mutans in biofilm (average 74.24 x 105) that saliva (average 14, 85 by 105), making it a tool of greater precision for the identification and quantification of S. mutans in biofilm during the stage of evaluation of cariogenic risk.
Finally, the appropriate distribution of dilutions on the standardization curve showed the procedural and methodological precision in creating the graph (the five recommended points were used), which allowed validating subsequent quantifications performed for each sample. The same accuracy was observed in the behavior of fluorescence under the threshold in analyzing No Template Controls, demonstrating the absence of external contaminations during the processes, as well as the absence of primer dimers.
CONCLUSIONS
The used methodology allows identifying and quantifying S. mutans in saliva and dental biofilm samples stored in cold, in a specific, sensitive and quick manner, using specific primers. The qPCR technique enables processing a greater volume of samples, which saves time in obtaining results.











 text in
text in