Introduction
Alzheimer's disease (AD) is a progressive and complex neurodegenerative disorder. It is the main cause of dementia in aging populations 1-4. It is estimated that approximately 35 million people suffer from AD, affecting around 6% of the population over 65 years old worldwide 3. The main symptoms of the disease include a decline in memory, cognitive functions, language skills, self-care, and behavioral changes. In advanced stages, individuals may even lose the capability to perform basic activities of daily living 4-7. In the central nervous system, AD is characterized by the deposition of β-amyloid (Aβ) plaques and intracellular neurofibrillary tangles (NFT) formed due to hyperphosphorylation of the Tau protein. It is also associated with oxidative stress, neuroinflammation, and neuronal loss in the brain 8-11. The neuropathology of this disease is not completely understood. Despite efforts to find a suitable treatment for AD, current treatments only provide symptomatic relief. However, their efficacy remains unsatisfactory 3,5-6,12-14.
Currently, pharmacological treatment for Alzheimer's disease (AD) focuses on acetylcholinesterase (AChE) inhibitors, which provide beneficial effects on the cognitive, functional, and behavioral symptoms of the disease 15. Therapeutic approaches also target several key pathways, including inhibition of Aβ protein aggregation (anti-amyloid strategy), modulation of amyloid transport, regulation of secretase enzymes, prevention of amyloid aggregation, amyloid-based vaccination, inhibition of Tau phosphorylation, blocking Tau oligomerization, enhancing Tau degradation, targeting intracellular signaling cascades, modulation of GABAergic neurons, and NMDA receptor antagonism. Pin1, highly expressed in adult neurons, regulates key proteins like Tau and APP, maintaining their functionality 8. The Tau protein, associated with microtubules, forms aggregates when excessively phosphorylated, disrupting the cytoskeleton and leading to cell death. This phosphorylation, mediated by kinases such as Cdk5, precedes the formation of helical filaments, a critical event in neurodegeneration 8. The production of Aβ peptide occurs through the amyloidogenic pathway of APP, where BACE1 (beta-secretase) and gamma-secretase cleave APP to generate Aβ peptides 8. Among these, multi-target directed ligands (MTDLs) are a promising approach in the search for new AD drugs, as they involve single chemical entities capable of simultaneously modulating multiple targets implicated in AD pathology 3,7,17. Additionally, numerous in silico studies have explored various computational approaches for AD treatment. These studies include constructing structural models to elucidate critical protein receptors involved in AD 18, conducting molecular docking studies to predict the 3D structure of complexes between macromolecules of interest 18-19, and employing ADMET profiling to assess pharmacokinetic properties such as blood-brain barrier permeability, hepatotoxicity, druggability, mutagenicity, and carcinogenicity 20. The aim of this study was to perform an in silico search for small molecules with ADME properties and capable of crossing the blood-brain barrier (BBB), which present multiple interactions with the proteins gamma-secretase, beta-secretase, pin1 and Cyclin-dependent kinase 5 (Cdk5) involved in the development of Alzheimer's disease, and of synthesizing a molecule to evaluate its neuroprotective effect.
Materials and methods
Equipment and Software
Open BabelGUI and Drug Likeness Tool (DruLiTo 1) were used to detect molecules with ADME properties. AutoDock Vina and autoDock 4.2 were the primary docking programs used in this work 21. A molecular format conversion program, Open BabelGUI, was used to transform geometries to SDF and mol2 format for their subsequent processing 22. The preparation of the PDBQT files and determination of the grid box size were carried out using Auto-Dock Tools version 1.1. SYBYL 8.1.1 was utilized to prepare protein structures 21. PyMol (DeLano Scientific LLC, USA) was employed to visualize protein-ligand complex structures. The identification of protein residues that interact with small molecules was carried out employing Discovery Studio. Computational studies were carried out using machine running on Inter Core i7 4.0GHZ, processor with 8GB RAM and 1TB hard disk.
Melting points were determined on a Büchi melting point apparatus and are uncorrected. IR spectra were recorded on a Shimadzu FTIR 8400 spectrophotometer in KBr disks. Mass spectra were run on a SHIMADZU-GCMS 2010-DI-2010 spectrometer (equipped with a direct inlet probe) operating at 70 eV. Microanalyses were performed on an Agilent elemental analyzer and the values are within 0.4% of the theoretical values. The starting compounds were purchased from Aldrich, Fluka and Acros (analytical reagent grades) and were used without further purification.
Selection of molecules with properties drug likeness
The database PubChem was used to download 10 million molecules deposited during 2015 and 2016. The molecules were downloaded in sdf format. These chemicals were filtered considering those have characteristics of drugs-like, to filter molecules were used free software Open BabelGUI and Drug Likeness Tool (DruLiTo 1). The rules Lipinski, that predicts if a compound is more likely to be membrane permeable and easily absorbed by the body 23. Veber rule, for predict good oral bioavailability 24. Ghose filter that defines drug-likeness constraints 25 and BBB rule, which evaluates the ability of compounds to cross the blood brain barrier. The molecules that passed these filters were selected for molecular docking, and previously prepared with Auto-Dock Tools. The molecules that passed the filters were converted to pdbqt format for performing molecular docking. The ligands were prepared using the GaussView 5.08 interface, and energy minimization was performed with the PM6 semi-empirical method in Gaussian09. Tautomers and ionization states at physiological pH (7.4) were also considered to obtain stable conformations suitable for docking.
Proteins structure preparation
The three-dimensional structure of AD proteins: Pin1, γ-secretase, β-secretase and Cyclin-dependent kinase 5 (Cdk5) were downloaded from Protein Data Bank (PDB: 4U84, 4UPC, 3VF3 and 3O0G, respectively) and prepared with SYBYL 8.1.1 package. The selection of protein structures was based on quality and structural diversity criteria. Proteins resolved by X-ray diffraction (resolution <2.0 Å) and electron microscopy (resolution ~3.40 Å) were included, prioritizing models with R-Value Free ≤ 0.25 to ensure reliability. Proteins were minimized using atomic partial charges by Kollman method, which describes the potential of the system in terms of the energy positions of the atoms and is parameterized for proteins and nucleic acids 26. MGLTools 1.5.0 software was utilized to convert structures from PDB to PDBQT format, adding polar hydrogens and assigning Kollman partial charges 26.
Protein-ligand docking
Molecular docking was performed using AutoDock Vina. The docking site for ligands on evaluated proteins was defined by establishing a cube with a sufficient dimension to cover the complete protein, with a grid point spacing of 1 Å for 4U84 and 4UPC proteins. For 3VF3 and 3O0G proteins, docking was performed at the active site of the protein. Three runs were carried out by ligand, and for each run the best pose was saved. Finally, the average binding affinity for best poses was accepted as the binding affinity value for a particular complex. The ten best protein-ligand complexes were selected for docking with AutoDock 4.2.
Refinement of molecular docking with AutoDock and identifying interactions
The docking program Autodock 4.2 was used to estimate accuracy, repeatability, and reliability of docking results. This program uses a semi-empirical free energy force field to predict binding free energies of protein-ligand complexes of a known structure and binding energies for both the bound and unbound states 27-29. Four ligand structures were selected and coupled with four proteins related to the development of Alzheimer's disease (PDB: 4U84, 4UPC, 3VF3 and 3O0G). The four best protein-ligand complexes were selected to identify residues of proteins that interact with small molecules, using Discovery Studio software 30. This program propose the number and type of primary existing ligand-residue interactions on the protein active site.
Docking validation
The validation of the docking process was carried out by redocking with β-secretase in complex with (3S,4S,5R)-3-(4-amino-3-bromo-5-fluorobenzyl)-5-{[3-(1,1-difluoroethyl)benzyl]amino}tetrahydro- 2H-thiopyran-4-ol 1,1-dioxide into your binding site; and cyclin-dependent kinase 5 in complex with {4-amino-2-[(4-chlorophenyl)amino]-1,3-thiazol- 5-yl}(3-nitrophenyl) methanone into your binding site. To examine the broader predictability of the docking, we compared the affinity values reported and ours for each ligand. The program of identification of protein residues Discovery Studio was used to do post-docking analyses.
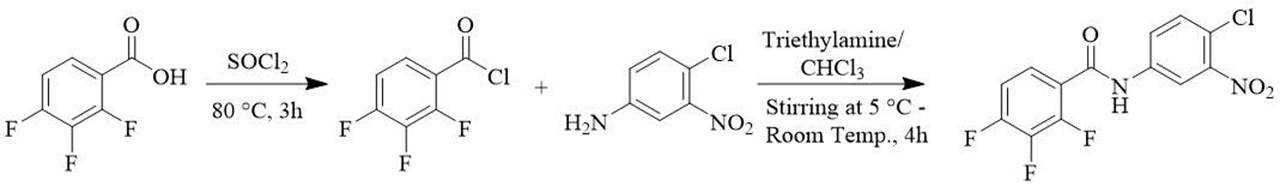
Synthesis of of N-(4-chloro-3-nitrophenyl)-2,3,4-trifluorobenzamide (84378305) (Scheme 1)
Compound 84378305 was selected for synthesis because it was one of the molecules that showed the highest affinity value with the proteins studied and it was possible to purchase the raw material for the synthesis.
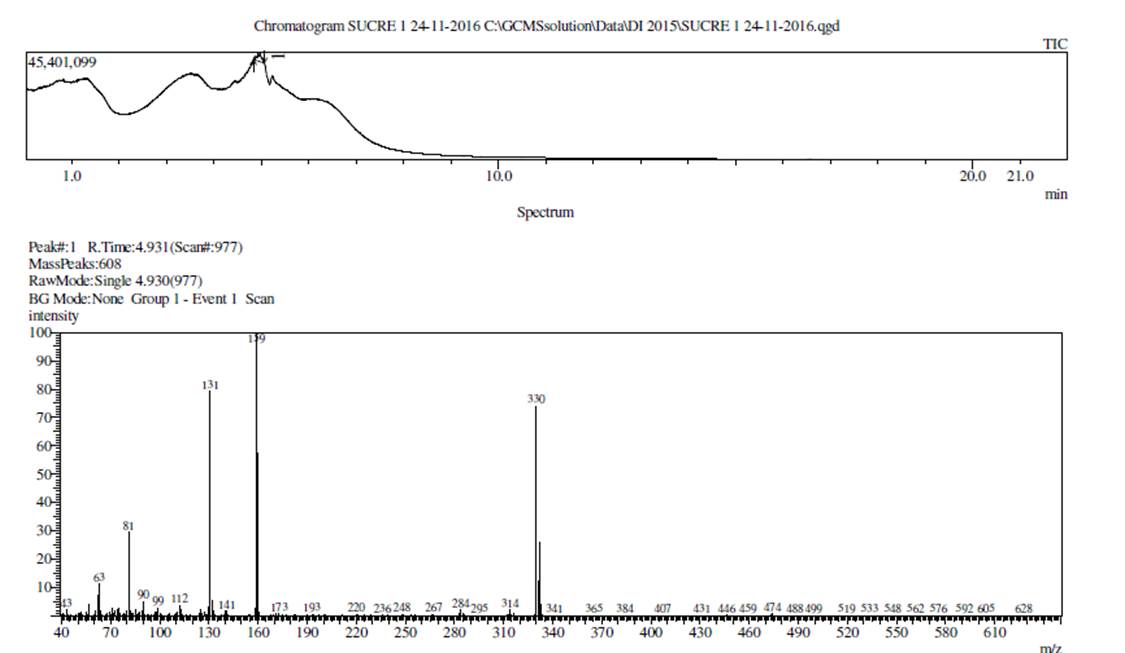
A mixture of 2,3,4-trifluorobenzoic acid (330 mg, 1 mmol) with SOCl2 (3 mL) was heated at 80 °C with stirring for 3 hours (Scheme 1). The reaction progress was monitored by TLC and after complete disappearance of the starting material the solvent was concentrated under vacuum to one half of the original volume. The residue was dissolved in 10 mL of chloroform and cooled to 5 °C in a water-ice bath for 5 min. Subsequently, a mixture of the 4-chloro-3-nitroaniline, (172 mg, 1 mmol) was slowly added with 3 equivalents of triethylamine (TEA) and stirred at room temperature for 4 h, (reaction monitored by TLC). The mixture was then washed with a saturated solution of KHCO3, the organic layer was extracted with chloroform (3 x 2 mL). The organic extract was dried with anhydrous Na2SO4, and the solvent was evaporated under reduced pressure, to afford compound 3. Beige solid, 85% yield; m.p. 178−180ºC; IR (KBr) (ν max, cm−1): 3387 (NH) 3118, 3081, 1659 (C=O), 1627, 1594 (C=C), 1531, 1315 (NO2), 1044. MS (EI) m/z (%): 330 (73, M+), 159 (100, M-171), 131 (79, M-198) 81 (29) 63 (11). Anal. Calcd. for C13H6ClF3N2O3: C, 48.05; H, 1.91; Cl, 10.02; F, 18.11; N, 8.25; O, 14.13. Found: C, 47.22; H, 1.83; Cl, 10.72; F, 17.24; N, 8.47; O, 14.52.
Pharmacological evaluation
The SK-N-SH cell line (ATCC # HTB-11), which is derived from human neuroblastoma, was introduced. These cells were cultured in DMEM culture medium containing 10% heat inactivated fetal bovine serum (FBS) and 0.5 mg / mL Gentamicin, incubated at 37 ° C and 5% CO2. For the maintenance of the cells, the medium was changed every 3 days and were passed through trypsin treatment once they reached approximately 80% confluence 11.
The selected molecule was subjected to cytotoxicity evaluation in the SK-N-SH cell line by decomposition test of 3-(4,5-dimethyl-2-thiazolyl)-2,5-diphenyl-2H-tetrazolium bromide (MTT). Briefly, 96 well plates were seeded at a density of 2.5x10 4 cells / well and incubated for 24 hours at 37 °C and 5% CO2, or until a confluence of 80-90% was reached. Subsequently, 100 μL of at least five serial (10-fold) dilutions were added in triplicate in DMEM culture medium and incubated for 48 hours again; after this time the medium was removed and 100 μL per well of 0.5 mg / mL MTT solution was added and an additional incubation was carried out for 1.5 hours.
The MTT solution was removed and 100 μL of DMSO was added, shaken gently for 10 minutes at room temperature; finally, the plate was read in a spectrophotometer at 492 nm with differential filter of 630 nm. The percentage of cell viability was calculated using the equation: (Absx * 100) / AbsMock, where Absx will correspond to the absorbance obtained in each of the wells in which the molecules and AbsMock were added to the geometric mean of the absorbance of the untreated wells (Controls). The percent viability calculation was performed for each replicate / dilution / molecule and estimates of cytotoxic concentration 50 (CC50) were performed by interpolation on a sigmoid dose-response curve generated from the data of each dilution used. The cytotoxicity assay was performed to ensure that cell viability remains above 70% at the concentrations used to assess neuroprotective effects, confirming that the molecule is not toxic to cells at these levels. This step is essential to attribute any observed neuroprotective effect directly to the molecule itself, thereby avoiding false-negative results due to cytotoxicity.
Measurement of neuroprotective effect: SK-N-SH cells were seeded at a density of 2.5 x 104 cells / mL in 96-well plates for 24 h at 37 °C in a CO2 incubator and exposed for 2 hours at different concentrations previously established in the cytotoxicity assay of the molecule evaluated. After this time, it was treated in the presence and absence of 10 μM of the Aβ25-35 peptide for 24, 48 and 72 hours. Once each incubation period was completed, cell viability was measured with the aid of the MTT reagent, according to the manufacturer's instructions. The absorbance was measured at a wavelength of 492 nm in a microplate reader 11,13. In all experiments untreated cells were used as control and each concentration of the compound in triplicate was evaluated. The IC50 value was calculated by regression analysis of the dose-response curve generated from the data.
Statistical Analysis
The statistical analysis of the data was performed using GraphPad Prism software. A Student's t-test was used to compare the results of the control group with the other experimental groups, in order to assess whether there were significant differences between the means of the groups regarding cell viability values or other relevant variables. The results are presented as the mean ± standard error of the mean (SEM). A difference was considered statistically significant if the p-value was less than 0.05.
Results
In silico study
The molecules utilized in this study were obtained from PubChem. A total of 10 million molecules were downloaded from the database and filtered based on drug-like characteristics. Only 3,968 molecules passed the Lipinski rule, Veber rule, Ghose filter, and BBB rule. Molecular docking was conducted using AutoDock Vina with the 3,968 molecules and proteins associated with Alzheimer's disease. The results for the top 30 ligand-protein complexes are shown in Table S1. To select the best protein-ligand complexes, a refinement process was performed using AutoDock. Additionally, commercially available molecules were selected. The results of this molecular docking are presented in Table 1. These molecules possess ADME properties and have high affinity for all proteins.
Table 1 Drug-like properties and AutoDock-calculated affinities obtained for docking of best molecules on proteins related to Alzheimer's disease.
| Compounds | Drug-like properties | Protein | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Affinity (Kcal/mol) | ||||||||||
| MW | Log p | LRV | GFV | VRV | BBBLR | 4U84 | 4UPC | 3VF3 | 3O0G | |
| 84554447 | 394.92 | 2.536 | 0 | 0 | 0 | 0 | -6.8±0.2 | -9.1±0.2 | -7.4±0.1 | -7.2±0.0 |
| 84577234 | 368.96 | 2.306 | 0 | 0 | 0 | 0 | -7.9±0.1 | -8.5±0.0 | -8.1±0.0 | -8.3±0.2 |
| 84577855 | 368.96 | 2.306 | 0 | 0 | 0 | 0 | -7.4±0.0 | -8.5±0.1 | -7.8±0.0 | -7.8±0.2 |
| 84378305 | 330.0 | 1.966 | 0 | 0 | 0 | 0 | -7.1±0.2 | -8.9±0.1 | -7.3±0.0 | -7.2±0.0 |
| (3S.4S.5R)-3-(4-amino-3-bromo-5-fluorobenzyl)-5-{(3-(1.1-difluoroethyl)benzyl)amino}tetrahydro-2H-thiopyran-4-ol 1.1-dioxide | 521.4 | 3.79 | 1 | 1 | 0 | 0 | - | - | -7.5±0.0 | - |
| {4-amino-2-((4-chlorophenyl)amino)-1.3-thiazol-5-yl}(3-nitrophenyl)methanone | 374.80 | 2.98 | 0 | 0 | 1 | 0 | - | - | - | -7.3±0.0 |
MW: Molecular weight; Logp: Logarithm of octanol/water partition coefficient; LRV: Lipinski rule of 5 violations: Maximum is 4 violations; GF: Ghose Filter violation; VR: Vebers Rule violation; BBBLR: BBB Likeness Rule violation.
The four compounds with higher affinity values interact with the proteins pin1 (pdb: 4U84), gamma secretase (pdb: 4UPC), beta secretase (pdb: 3VF3), and Cyclin-dependent kinase 5 (Cdk5) (pdb: 3O0G) in their different bonding sites, which are shown in Figures 1, 2, 3 and 4.
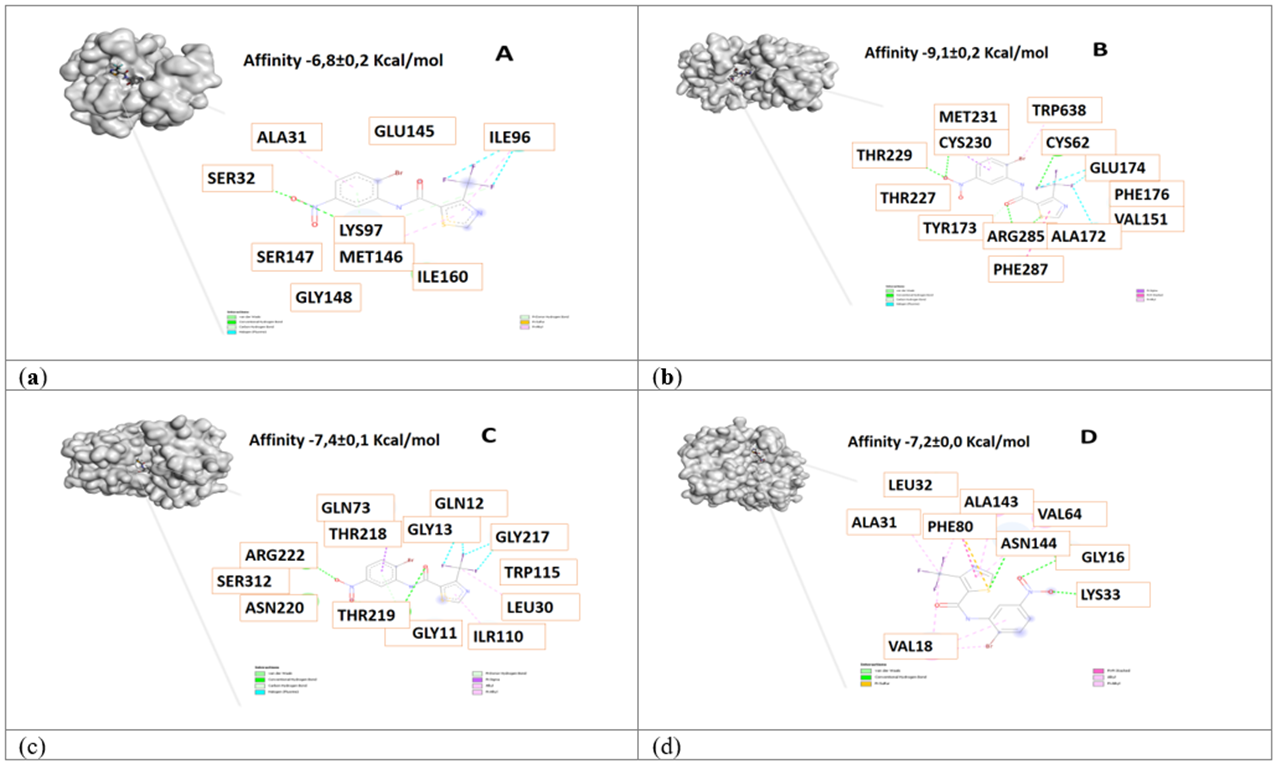
Figure 1 3D view and interacting residues present in proteins related to Alzheimer's disease. A. Pin1-84554447, B. γ-Secretase -84554447, C. β-Secretase -84554447, D. Cdk5-84554447 complexes. The molecules with the highest affinity for protein are represented.
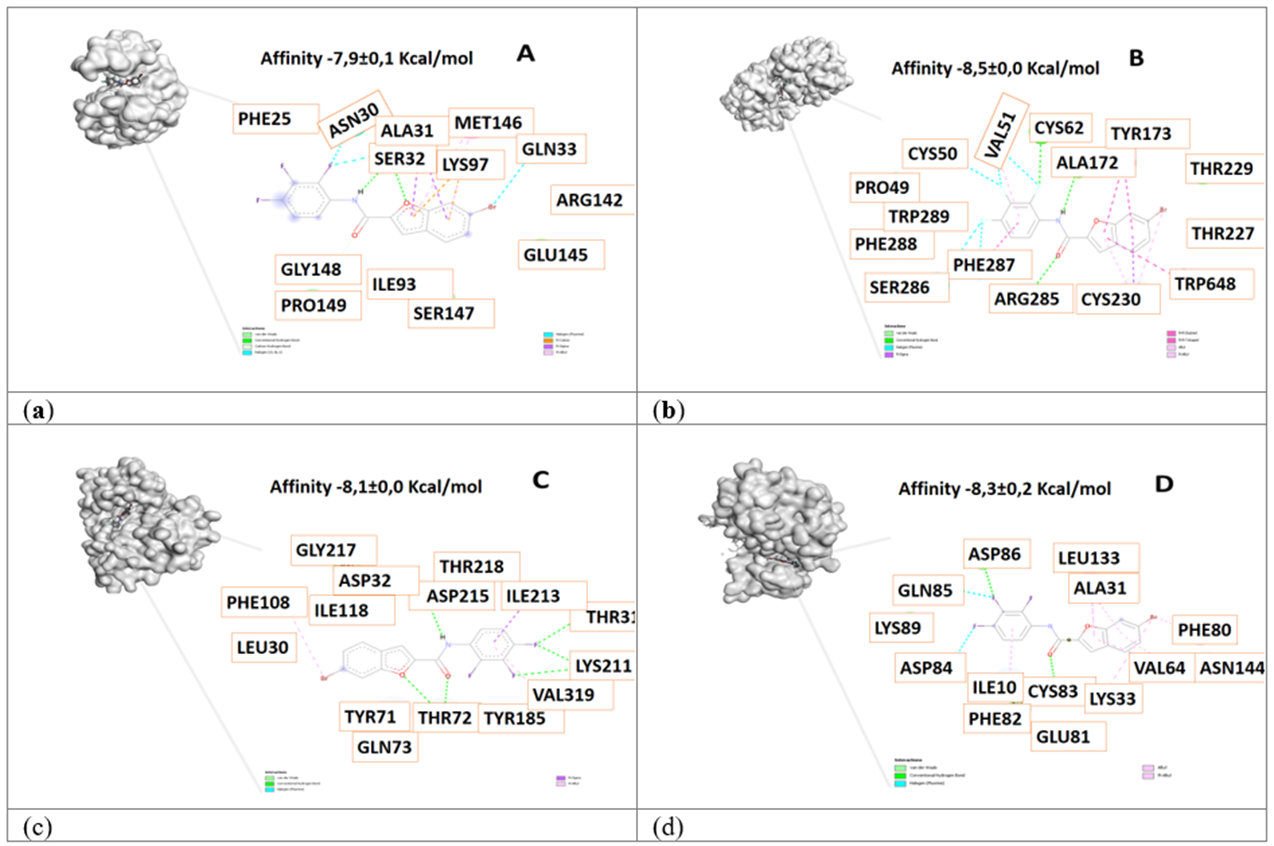
Figure 2 3D view and interacting residues present in proteins related to Alzheimer's disease. A. Pin1-84577234, B. γ-Secretase -84577234, C. β-Secretase -84577234, D. Cdk5 -84577234 complexes. The molecules with the highest affinity for protein are represented.
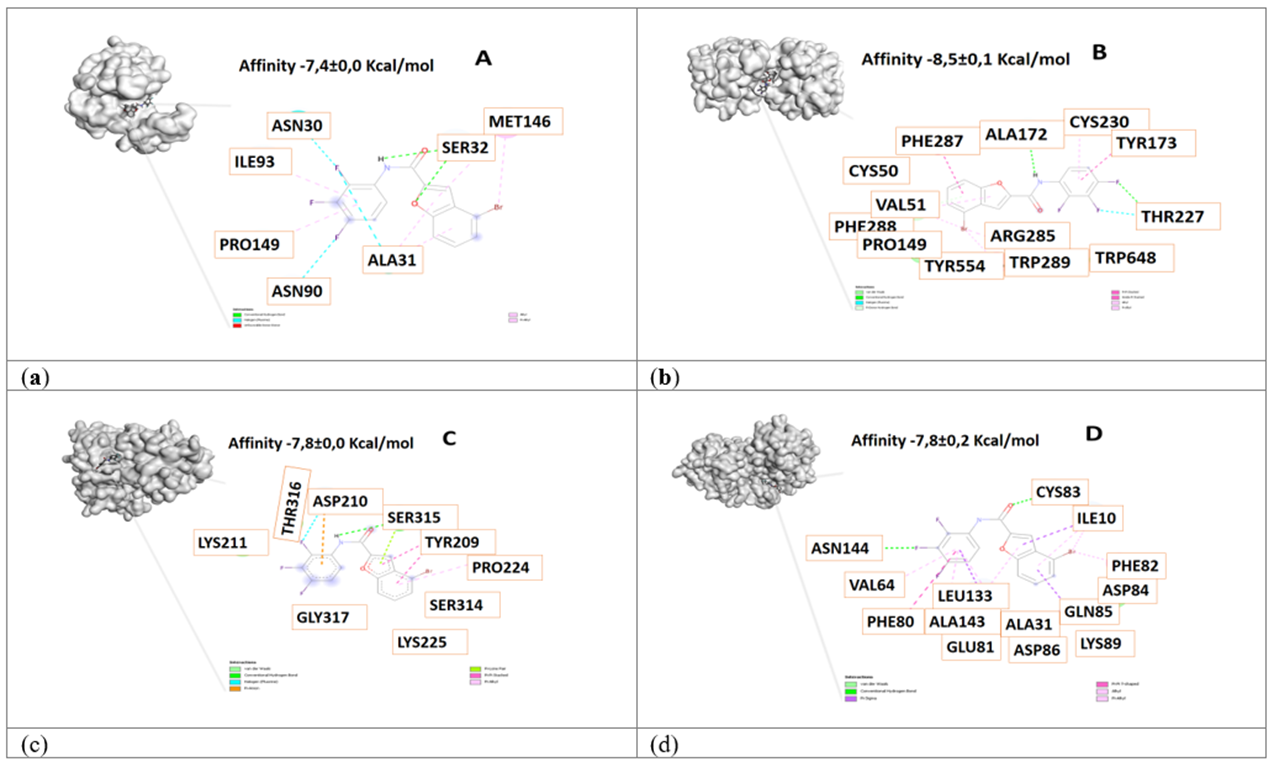
Figure 3 3D view and interacting residues present in proteins related to Alzheimer's disease. A. Pin1-84577855, B. γ-Secretase -84577855, C. β-Secretase -84577855, D. Cdk5-84577855 complexes. The molecules with the highest affinity for protein are represented.
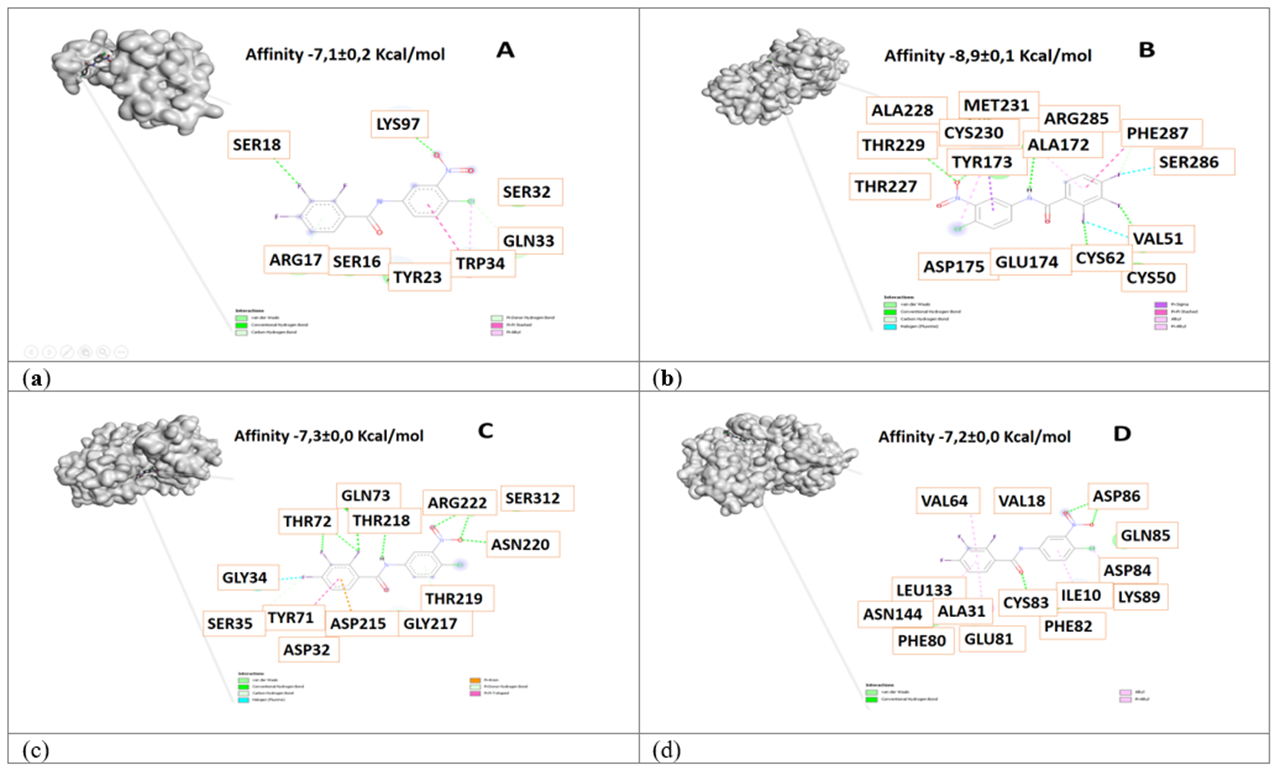
Figure 4 3D view and interacting residues present in proteins related to Alzheimer's disease. A. Pin1-84378305, B. γ-Secretase -84378305, C. β-Secretase -84378305, D. Cdk5 -84378305 complexes. The molecules with the highest affinity for protein are represented.
To validate the molecular docking, the binding mode of the inhibitor (3S,4S,5R)-3-(4-amino-3-bromo-5-fluorobenzyl)-5-{[3-(1,1-difluoroethyl)benzyl]amino} tet-rahydro-2H-thiopyran-4-ol 1,1-dioxide with the 3VF3 protein was analyzed, along with the amino acids involved in the interaction (Figure 5, Table 2).
Table 2 Interacting residues 3VF3 protein - inhibitor complex in comparison with ligands studied
| Interaction β-secretase (3VF3) in complex with inhibitor | Interaction identified in the protein - ligand complexes | |||
|---|---|---|---|---|
| 84554447 | 84577234 | 84577855 | 84378305 | |
| TYR185 | - | X | - | - |
| ASP215 | - | X | - | - |
| SER35 | - | - | - | X |
| ASP32 | - | X | - | X |
| PHE108 | - | X | - | - |
| TRY71 | - | X | - | X |
| THR72 | - | X | - | X |
| GLN73 | - | X | - | X |
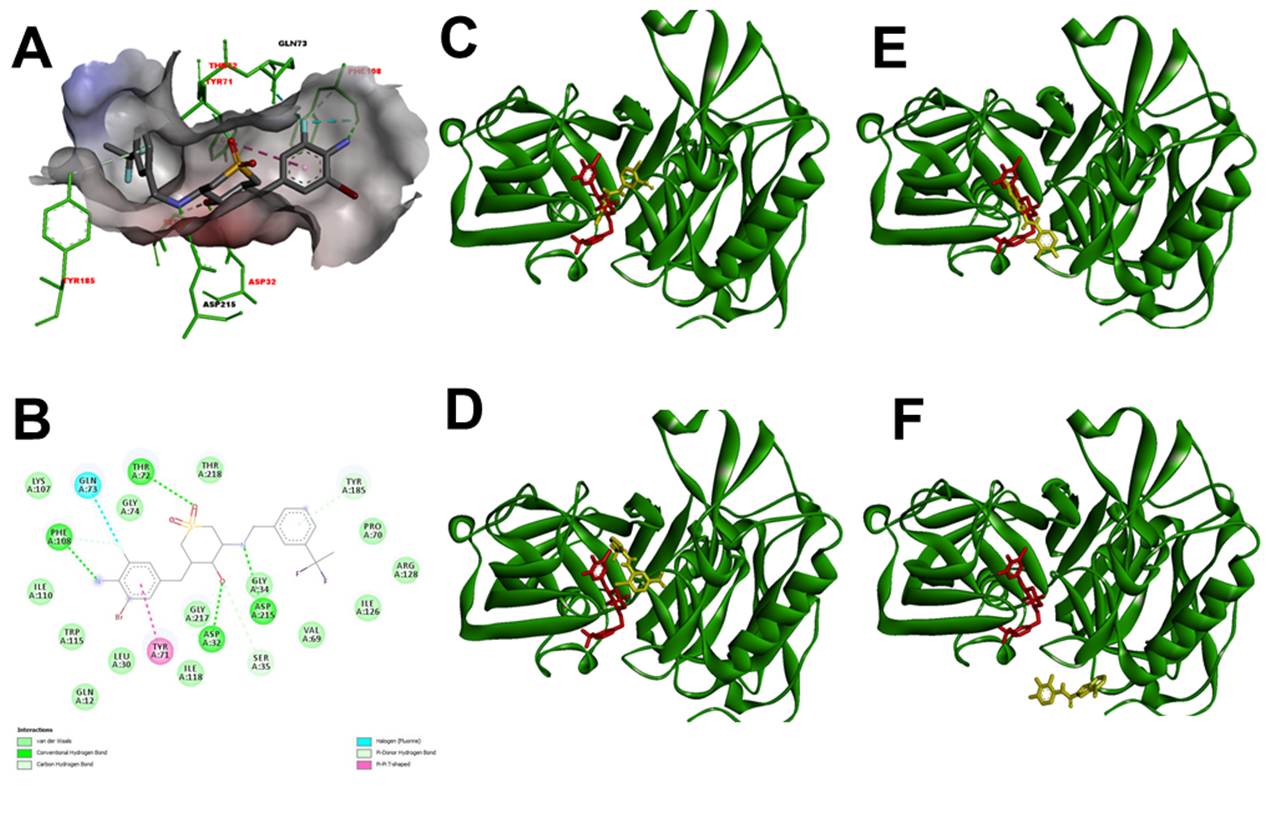
Figure 5 β-secretase (3VF3) in complex with (3S,4S,5R)-3-(4-amino-3-bromo-5-fluorobenzyl)- 5-{[3-(1,1-difluoroethyl)benzyl]amino}tetrahydro- 2H-thiopyran-4-ol 1,1-dioxide (Red). A. 3D view protein-inhibitor complex. B. interacting residues protein-inhibitor complex. C. 3D view protein-inhibitor (Red) - ligand 84378305 (Yellow) complex. D. 3D view protein-inhibitor (Red) - ligand 84554447 (Yellow) complex. E. 3D view protein-inhibitor (Red) - ligand 84577234 (Yellow) complex. F. 3D view protein-inhibitor (Red) - ligand 84577855 (Yellow) complex.
Validation was also performed with Cdk5 protein and its inhibitor {4-amino-2-[(4-chlorophenyl)amino]-1,3-thiazol- 5-yl}(3-nitrophenyl)methanone, the results are shown in Table 3 and Figure 6.
Table 3 Interacting residues 3O0G protein - inhibitor complex in comparison with ligands studied
| Interaction cyclin-dependent kinase 5 (3O0G) in complex with inhibitor | Interaction identified in the protein - ligand complexes | |||
|---|---|---|---|---|
| 84554447 | 84577234 | 84577855 | 84378305 | |
| GLU81 | - | X | X | X |
| ALA31 | X | X | X | X |
| LEU133 | - | X | X | X |
| CYS83 | - | X | X | X |
| ILE10 | - | X | X | X |
| PHE82 | - | X | X | X |
| ALA143 | X | - | X | - |
| VAL18 | - | - | - | X |
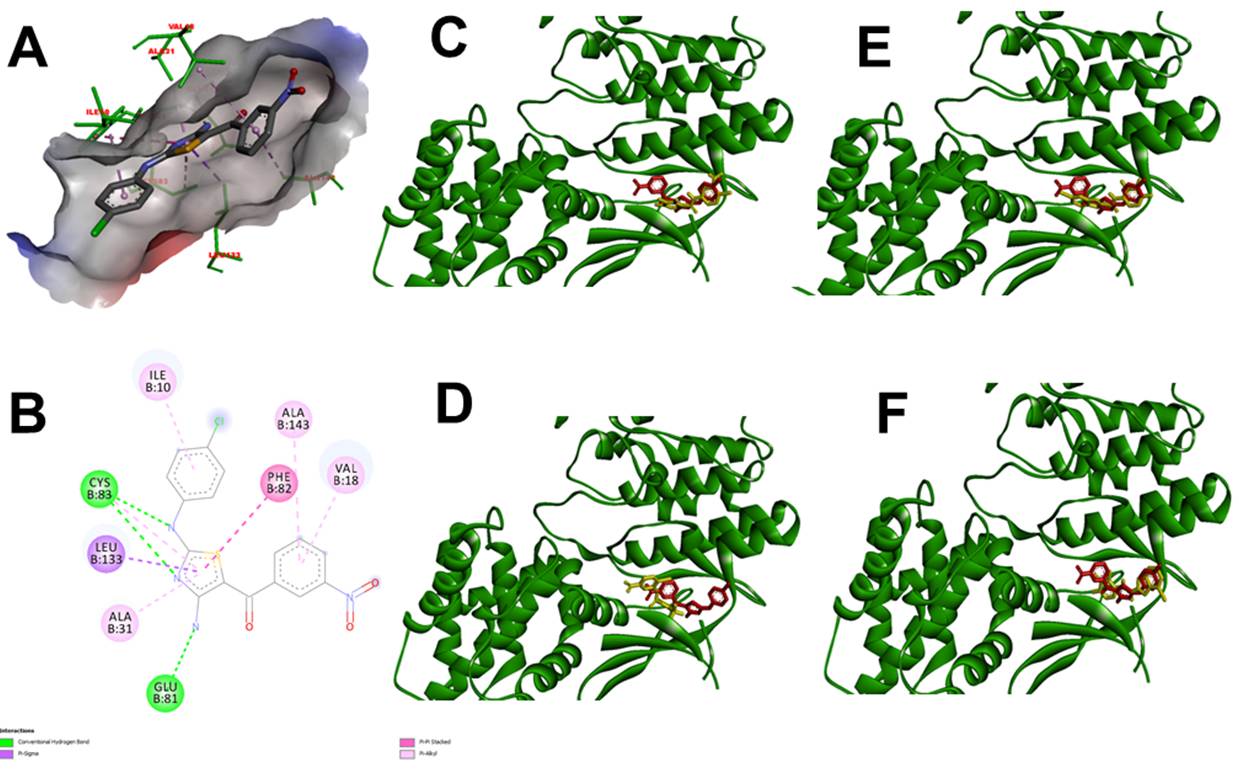
Figure 6 Cyclin-dependent kinase 5 (3O0G) in complex with {4-amino-2-[(4-chlorophenyl)amino]-1,3-thiazol- 5-yl}(3-nitrophenyl)methanone (Red). A. 3D view protein-inhibitor complex. B. interacting residues protein-inhibitor complex. C. 3D view protein-inhibitor (Red) - ligand 84378305 (Yellow) complex. D. 3D view protein-inhibitor (Red) - ligand 84554447 (Yellow) complex. E. 3D view protein-inhibitor (Red) - ligand 84577234 (Yellow) complex. F. 3D view protein-inhibitor (Red) - ligand 84577855 (Yellow) complex.
Synthesis and experimental evaluation
The compound with code 84378305 was synthesized (Figure 7), and its cytotoxicity was assessed. Figure 8 presents the cytotoxic activity of the synthesized molecule on neuroblastoma cells over a 48-hour period. The compound 84378305 was found to significantly reduce cell viability in SK-N-SH cells to concentrations more than 10 µM. Exposure to concentrations ranging from 0.001 µg/mL to 10 µM did not affect viability, as shown in Figure 8. The corresponding IC50 value for SK-N-SH cells at 48 h were 84.7 µM (52 to 136 µM). Concentrations below 1 µM were used to evaluate the neuroprotective effect, as cell viability remained above 70%.
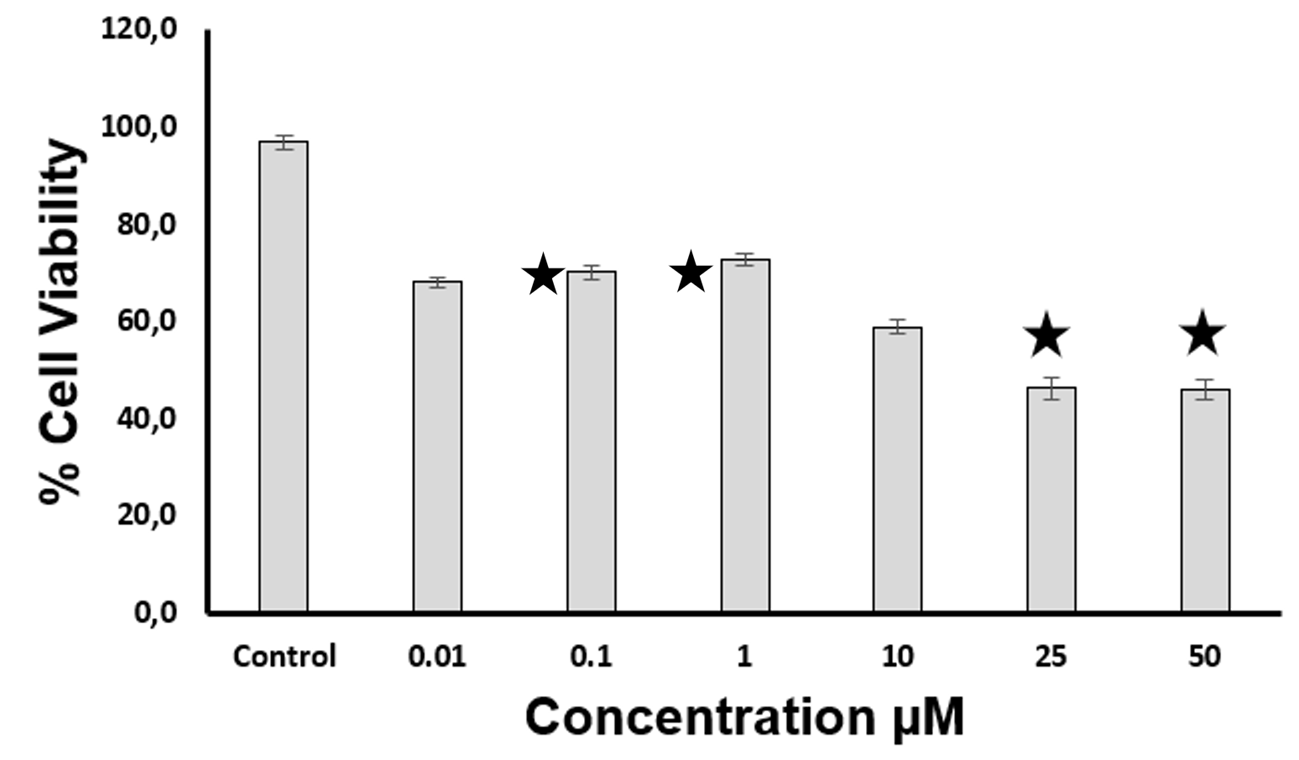
Figure 8 Cytotoxic effect of compound 84378305. The asterisks indicate a significant difference compared to the control. p < 0.05
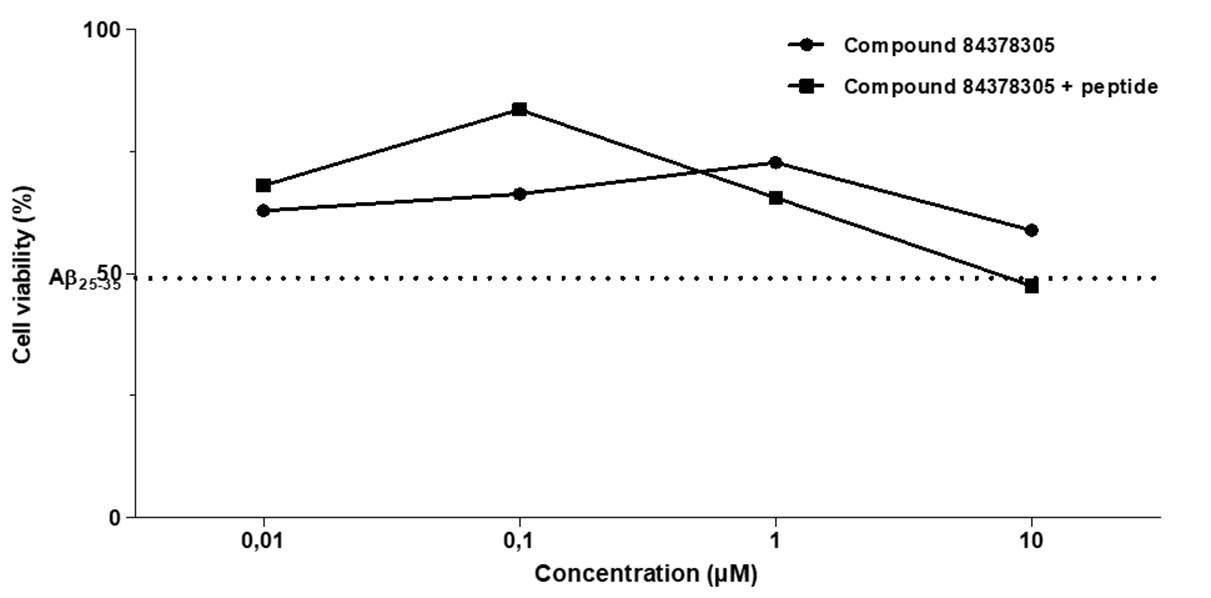
The neuroprotective effects of compound 84378305 against cytotoxicity in SK-N-SH cells induced by Aß25-35 peptide after 48 h exposures, are shown in Figure 9. The results showed that when SK-N-SH cells were simultaneously treated with Aß25-35 peptide (5 µM) and compound 84378305 (0,1 µM), the molecule exhibited significant neuroprotection (33%) after the 48 h of incubation.
Discussion
In silico study
The proteins analyzed were Pin1, gamma-secretase, beta-secretase, and cyclin-dependent kinase 5 (Cdk5), all implicated in Alzheimer's pathology. Pin1, highly expressed in adult neurons, regulates proteins like Tau and APP, preserving their functionality. Dysregulation of these pathways contributes to neurodegeneration, including Tau aggregation and Aβ peptide production through amyloidogenic APP processing 31,8. The selected ligands interact with all the analyzed proteins, making them potential MTDLs. Pin1 (4U84) is a prolyl isomerase whose regulation is still not fully understood 32. The oxidation of CYS113 in Pin1 inactivates its isomerization ability and its regulation of proteins like Tau and APP. A network of hydrogen bonds involving CYS113-SER115-HIS59-HIS157-THR152 has been proposed as key to its activity 32. In this study, ligands 84554447, 84577234, 84577855, and 84378305 interacted primarily with the amino acids SER32, ALA31, and LYS97 (Figures 1-4). The highest affinity was observed with ligand 84577234 (-7.9 kcal/mol), due to hydrogen bonds and pi-sigma and pi-cation interactions. Although strong interactions were detected with all the ligands, it cannot be confirmed whether the isomerization activity is inhibited.
Gamma-secretase (4UPC) is a multiprotein complex composed of presenilin, nicastrin, Aph-1, and Pen-2, whose combined action regulates Aβ peptide production; all these proteins are necessary for its complete proteolytic activity 8,33. Presenilins (PS1 and PS2) provide the proteolytic activity, while nicastrin facilitates substrate recognition, and Aph-1 acts as scaffolding in the assembly of the complex 34. In this study, the key residues in 4UPC were CYS230, VAL51, ALA172, PHE287, ARG285, TYR173, and THR227 (Figures 1-4). Interactions with ligands 84554447 and 84378305 included hydrogen bonds, Van der Waals interactions, and pi-pi stacking, while ligands 84577234 and 84577855 showed pi-sigma and pi-allyl interactions. It is important to note that the residues ASP257 and ASP385 in the transmembrane domains are crucial for the catalytic activity of the protease 35, and the residue GLU332 in nicastrin is critical for γ-secretase assembly and maturation 36.
Several compounds have been identified as γ-secretase inhibitors, including BMS-708163, GSM-1, and E2012, which target presenilin-1, the catalytic core of this complex. Cross-competition studies show that each of these inhibitors binds to different sites within presenilin-1, and GSM-1 binding induces a conformational change in the active site of γ-secretase. Furthermore, the affinity of E2012 increases in the presence of an active-site transition inhibitor 37. Puentes et al., 2022 38 reported that curcumin analogs interact with γ-secretase with affinities up to -8.9 kcal/mol. In this study, the four identified compounds presented affinities greater than -8.5 kcal/mol (Table 1), acting as potential allosteric modulators of γ-secretase, with interacting amino acids close to the key residues for its activity. Additionally, allosteric γ-secretase inhibitors (AGSIs), such as the coumarin-dimer class, selectively inhibit Aβ42 production in vitro and cell models, showing greater selectivity for Aβ42 over Aβ40, Aβ38, or Notch by binding to an allosteric site on γ-secretase 39.
Regarding beta-secretase (BACE-1, 3VF3), this is an aspartic protease type I with two aspartate residues in its active site 40. Several BACE-1 inhibitors have been developed, though many have high molecular weight peptide structures and limiting properties for blood-brain barrier penetration 41. In this study, the interaction of ligands 84554447, 84577234, 84577855, and 84378305 with protein 3VF3 revealed key amino acids involved, namely GLN73, THR218, and GLY217. Beta-secretase showed a high affinity for 84577234 (-8.1 kcal/mol), facilitated by hydrogen bonds formed by ASP215, THR72, THR316, and LYS211 with the ligand molecule. Additionally, a pi-sigma interaction involving ILE213 and a pi-allyl interaction involving VAL319 and PHE108 contributed to the favorable binding. For ligand 84577855, the interaction with SER315 was due to hydrogen bonds, pi-allyl interaction with PRO224, pi-pi stacking with TYR209, and halogen bonding with ASP210. For ligand 84378305, Van der Waals interactions (SER35, THR219), hydrogen bonds (THR72, GLN43, THR218, ARG222, ASN220), pi-pi interactions (THR71), and halogen interactions (GLY34) were observed. For ligand 84554447, hydrogen bonds (ARG222), Van der Waals interactions (THR219), halogen interactions (GLN12, GLY217), and pi-allyl interactions (ILE110, LEU30) were identified. The docking study revealed the significant role of Van der Waals forces in stabilizing the protein-ligand complex. The Van der Waals interaction and hydrogen bond formed by reactive amino acid residues of protein 3VF3 with the molecules lead to the binding of BACE1 with 84378305 and 84554447. Other studies have reported a binding affinity of -10 kcal/mol for ginsenoside Rb1 toward BACE1, with interacting residues such as ILE126, ARG235, ARG307, LYS321, SER325, PHE108, ILE118, GLN73, and others 42. Some of these residues were also identified in our complexes, such as PHE108 and ILE118 with 84577234, and GLN73 with 84554447, 84577234, and 84378305 via Van der Waals interactions. We studied the docking of protein 3VF3 with the inhibitor (3S,4S,5R)-3-(4-amino-3-bromo-5-fluorobenzyl)-5-{[3-(1,1-difluoroethyl)benzyl]amino}tetrahydro-2H-thiopyran-4-ol 1,1-dioxide (Figure 5) 44 (-7,5±0,0 Kcal/mol) and were able to identify the binding site of the inhibitors and the interacting amino acids, noticing that ligands 84378305 and 84577234 bind at the same site as the inhibitor and interact with the amino acids (Table 3) and with similar affinity value. Therefore, these ligands may act as inhibitors of beta-secretase.
Cyclin-dependent kinase 5 (Cdk5) is a key target in the research of therapies for neurological disorders, particularly Alzheimer's disease, where it is involved in forming neurofibrillary tangles 45,46. This study evaluated potential Cdk5 inhibitors through molecular docking, highlighting relevant interactions between the protein and ligands 84554447, 84577234, 84577855, and 84378305, with high-affinity values in all cases. Figures 1 to 4 present the specific amino acid interactions in each complex. The residues ALA31, ASN144, and VAL64 were recurrently identified in the complexes, acting via hydrogen bonds and pi-pi stacking. In the Cdk5-84554447 complex, key interactions with VAL18, ALA31, ALA143, VAL64 (pi-allyl), ASN144, GLY16, and LYS33 (hydrogen bond), and PHE80 (pi-sulfur) were observed. In the Cdk5-84577234 complex, residues ILE10, LYS33, VAL64, THR80, ALA31, LEU133 (pi-allyl), CYS83, and ASP86 (hydrogen bond), and GLN83 and ASP84 (halogen interaction) were critical. Similarly, in the Cdk5-84577855 complex, interactions with ASN144, CYS83 (hydrogen bond), LEU133, GLN85, ILE10 (pi-sigma), PHE80 (pi-pi stacking), VAL64, ALA143, ALA31, and PHE82 (pi-allyl) were identified. Finally, in the Cdk5-84378305 complex, residues ASP86, CYS83 (hydrogen bond), VAL64, LEU133, ALA31, ILE10, and LYS89 (pi-allyl) were important. Comparing these findings with known inhibitors, such as {4-amino-2-[(4-chlorophenyl)amino]-1,3-thiazol-5-yl}(3-nitrophenyl)methanone (Figure 6) (-7,3±0,0 Kcal/mol), it was observed that similar residues involved in binding to Cdk5 were present in our results and with similar affinity value, specifically in complexes with 84554447, 84577234, and 84378305. This reinforces the potential of these ligands as Cdk5 inhibitors.
Recently, the crystal structures of the active CDK5/p25 kinase complexed with three inhibitors-(R)-roscovitine, aloisin-A, and indirubin-3’-oxime-were reported. These inhibitors bind to the well-conserved catalytic pocket of the kinase, interacting with residues LEU55, VAL64, PHE80, ASN144, PHE145, and ALA143, providing a potential space for rational drug design 45. New Cdk5 inhibitors have been reported, including olomoucine, roscovitine, butyrolactone I, purvalanols, indirubins, hymenialdisine, and paullones, which are ATP competitors 47. Other studies have identified a series of potent and selective 2-aminothiazole inhibitors of Cdk5/p25 as potential therapeutic agents for Alzheimer's disease (AD) treatment 48.
The molecular docking validation using inhibitors reported in the literature 44-45 was carried out with proteins 3VF3 and 3O0G. It was observed that these inhibitors bind to the same active site as reported and interact with the previously identified amino acids (Figures 5 and 6, Tables 3 and 4). Rueeger et al. (2012) reported amino acids ASP32, THR72, and GLN73 were involved in the interaction between protein 3VF3 and the inhibitor, which was confirmed in our validation 44. Additionally, Ahn et al. (2005) reported that the amino acids ILE10, VAL18, ALA31, PHE82, LEU133, GLU81, CYS83, and ALA143 were involved in the interaction between protein 3O0G and the inhibitor, which was also confirmed in our validation. This confirms the reliability of the molecular docking process 45.
Currently, available drugs for the treatment of Alzheimer's disease are insufficient for the proper management of this pathology, underscoring the urgent need for more advanced drug development 12. The MTDL (multi-target drug design) strategy is an attractive approach for developing drugs targeting disorders involving complex pathological mechanisms, such as multifactorial neurodegenerative AD 49. Indeed, virtual screening through drug-like ADMET filtering, the best pharmacophore model, and molecular docking analyses have been used to identify new inhibitors of AGSI, AChE, BACE-1, and Cdk5 4,37,42. In recent years, many MTDLs with improved pharmacological profiles have been described, such as β-secretase inhibitors with AChE inhibitory activity or metal-chelating properties, including the design of compound 46, which incorporates a 1,4-benzoquinone functionality as a radical scavenger in the polyamine backbone series of cholinergic derivatives. It is a potent inhibitor of AChE activity (IC50 = 1.55 nM) and also inhibited β-secretase activity in a concentration-dependent manner (IC50 = 108 nM) 40. Other compounds described include loganin, morroniside, and 7-O-galloil-D-sedoheptulosa, potential candidates for AD treatment by inhibiting AChE and BACE1 activities 43. In this study, four candidates were identified that act on the proteins Pin1, gamma-secretase, beta-secretase, and Cdk5, making them potential MTDLs.
Synthesis
Synthesis of N-(4-chloro-3-nitrophenyl)-2,3,4-trifluorobenzamide (84378305): The amidation of carboxylic acids using acyl chlorides is typically a two-step process, involving first the conversion of the acid to the acyl chloride, followed by the coupling with the amine. In our study, we aimed to determine the viability of this process by subjecting benzoic acid (1) to thionyl chloride (SOCl2) at 80°C for 3 hours. TLC analysis of the reaction mixture confirmed the complete conversion of benzoic acid. To recover the reaction amide (4), the solvent was evaporated under reduced pressure to remove any remaining unreacted thionyl chloride (Scheme 1). The resulting residue was dissolved in dichloromethane and stirred overnight at room temperature in the presence of arylamine and triethylamine. The reaction progress was monitored by TLC. Once the reaction was complete, the resulting mixture was processed to yield the desired product (4) with an overall yield of 85%.
Cytotoxicity and Neuroprotective Effect
Aβ aggregation is a key hallmark of AD 50. Molecules capable of interacting with multiple receptors and acting as agonists or antagonists are highly promising as potential treatments for Alzheimer's disease 51. The synthesized molecule exhibits a neuroprotective effect and low cytotoxicity at the evaluated concentrations. To determine the non-cytotoxic dose ranges of compound 84378305 for SK-N-SH cells, cytotoxicity was assessed after exposure to varying molecule concentrations up to 50 μM. After 48 hours of culture, compound 84378305 showed minimal or no cytotoxic effects in SK-N-SH cells at concentrations up to 10 μM (Figure 7), with cell viability reaching 70% compared to the vehicle-treated control group. Considering that the IC50 value for SK-N-SH cells at 48 hours was 84.7 µM, this indicates that a high concentration of compound 84378305 is required to induce 50% cell death. Based on these findings, concentrations ranging from 10 to 0.01 µM were selected for the neuroprotective assay, evaluating Aβ25-35 cytotoxicity in SK-N-SH cells using the MTT reduction assay. As shown in Figure 8, a concentration of 5 µM of Aβ25-35 resulted in decreased cell viability, indicating Aβ25-35-induced toxicity in SK-N-SH cells. Moon et al. (2019) 52 report a study evaluating the neuroprotective effect of the ethyl acetate fraction of the methanol extract of Ophiophogon japonicus using a similar methodology, demonstrating strong protective effects against Aβ25-35-induced cytotoxicity in PC12 cells.
To investigate whether compound 84378305 could inhibit neuronal cell death induced by beta-amyloid peptide, SK-N-SH cells were co-exposed to 5 μM of Aβ25-35 and compound 84378305 at various concentrations for 48 hours. As illustrated in Figure 7, when SK-N-SH cells were cultured for 48 hours following treatment with 5 μM Aβ25-35 alone, cell viability decreased to approximately 50% compared to vehicle treatment. However, when SK-N-SH cells were cultured for 48 hours after co-treatment with 5 μM Aβ25-35 and compound 84378305, all concentrations of compound 84378305 demonstrated a strong ability to mitigate Aβ25-35-induced cytotoxicity. Specifically, at a concentration of 0.1 μM, compound 84378305 significantly and effectively prevented Aβ25-35-induced cell death, exhibiting significant neuroprotection (33%). Other studies have reported the neuroprotective effect of other molecules. For example, the neuroprotective effects of quercetin have been widely studied. At low micromolar concentrations, it antagonizes cell toxicity due to oxidative stress in neurons. It suppresses neuroinflammatory processes by negatively regulating pro-inflammatory cytokines, such as NF-kB and iNOS, while stimulating neuronal regeneration 50. Limonene (+) has a neuroprotective function against Aβ42 neurotoxicity and is thus a potential therapeutic agent for AD, though the detailed mechanism of this neuroprotective action was not elucidated. However, our results suggest that the antioxidant and anti-inflammatory properties of limonene (+) may play an important role in neuroprotection 53. Additionally, neuropeptides exert significant neuroprotective effects against Aβ-induced neuronal toxicity at nanomolar and micromolar concentrations 54. Finally, it has been reported that genistein (10, 30, and 50 μM) significantly increases the survival rate of SH-SY5Y cells treated with Aβ25-35 55.
Conclusions
We identified small molecules that interact with key proteins associated with Alzheimer's disease, including one with promising neuroprotective activity. These molecules function as multi-target directed ligands (MTDLs), showcasing the potential for treating Alzheimer's, a multifactorial neurodegenerative disorder. However, further detailed analysis of the underlying molecular pathways, and the evaluation of all four identified compounds, is essential to fully assess their neuroprotective effects and validate the overall potential of this platform. The results highlight the efficacy of combining computational methodologies with pharmacological evaluation in the drug discovery process, underscoring the promise of this approach for developing novel therapeutic agents.