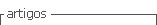Serviços Personalizados
Journal
Artigo
Indicadores
-
 Citado por SciELO
Citado por SciELO -
 Acessos
Acessos
Links relacionados
-
 Citado por Google
Citado por Google -
 Similares em
SciELO
Similares em
SciELO -
 Similares em Google
Similares em Google
Compartilhar
DYNA
versão impressa ISSN 0012-7353
Dyna rev.fac.nac.minas vol.80 no.180 Medellín jul./ago. 2013
CHARACTERIZATION OF HYDROCARBONOCLASTIC MARINE BACTERIA USING THE 16S rRNA GENE: A MICROCOSM CASE STUDY
CARACTERIZACIÓN DE BACTERIAS MARINAS HIDROCARBUROCLÁSTICAS USANDO EL GEN ARNr 16S: UN ESTUDIO DE MICROCOSMOS
DOLLY REVELO ROMO
MSc Microbiología - Grupo de Investigación en Biotecnología Microbiana - Universidad de Nariño, margo.revelo@gmail.com
MARÍA ISABEL GÓMEZ PERDOMO
Bióloga - Grupo de Investigación en Biotecnología Microbiana - Universidad de Nariño, bio_isis@hotmail.com
MARCELA CONCHA OBANDO
Bióloga - Grupo de Investigación en Biotecnología Microbiana - Universidad de Nariño, mc1226@gmail.com
DANIEL BRAVO
PhD Microbiología- Grupo de Investigación en Biotecnología Microbiana - Universidad de Nariño, daniel.bravo@unine.ch
PABLO FERNÁNDEZ IZQUIERDO
PhD Ciencias Biológicas - Grupo de Investigación en Biotecnología Microbiana - Universidad de Nariño, pabfdez@gmail.com
Received for review December 8th, 2011, accepted March 15th, 2013, final version March, 21th, 2013
RESUMEN: Algunos microorganismos marinos pueden degradar contaminantes derivados del petróleo usándolos como única fuente de carbono y energía. Este grupo heterogéneo es llamado bacterias hidrocarburoclásticas. Sin embargo, se subestima a aquellas bacterias hidrocarburoclásticas que aun no han sido caracterizadas. En este contexto, se llevo a cabo un estudio de microcosmos simulando un ambiente marino impactado con petróleo, en Biorreactores de Flujo Ascendente (BFA). A partir de estos microcosmos fueron caracterizados once aislamientos bacterianos. Se determinó la ubicación taxonómica de las bacterias identificadas en 4 géneros principales por secuenciación parcial del gen ARNr 16S: Bacillus, Pseudomonas, Halomonas y Haererehalobacter. La investigación demostró la presencia de bacterias hidrocarburoclásticas reconocidas en consorcios y proporcionó información adicional acerca de Haererehalobater como un nuevo género de bacterias marinas hidrocarburoclásticas. Desde esta perspectiva, se pretendió aportar al conocimiento de la diversidad de las bacterias marinas hidrocarburoclásticas y su uso potencial en biorremediación y otros procesos biotecnológicos.
PALABRAS CLAVE: Petróleo; bacterias marinas hidrocarburoclásticas; biodegradación; ARNr 16S; microcosmos.
ABSTRACT: Some marine microorganisms can degrade oil pollutants by using them as their sole carbon and energy sources. Members of this heterogenic group are called hydrocarbonoclastic bacteria. However, an unestimated number of hydrocarbonoclastic marine bacteria have not yet been characterized. In this context, a microcosm study was carried out, simulating a marine environment contaminated with oil in Upstream Flow Bioreactors (UFB). Eleven bacterial isolates were characterized from these microcosms. The taxonomic position of the bacteria identified was determined by partial sequencing of the gene 16S rRNA in 4 major genera: Bacillus, Pseudomonas, Halomonas and Haererehalobacter. This research demonstrated the presence of hydrocarbonoclastic bacteria recognized in consortia and provides additional information about Haererehalobater a new genera of hydrocarbonoclastic marine bacteria. The outlook for better understanding of the diversity of hydrocarbonoclastic marine bacteria and potential uses in bioremediation and other biotechnological processes is discussed.
KEYWORDS: Oil; hydrocarbonoclastic marine bacteria; biodegradation; 16S rRNA; microcosm
1. INTRODUCTION
Marine pollution by oil discharges produces highly negative effects on biotic and abiotic processes [1]. On the Colombian Pacific coast, the Bay of Tumaco-Nariño, an important port with oil activities, is frequently exposed to the environmental impacts of oil discharges [2]. Most marine environments contain hydrocarbon-degrading bacteria with different metabolic capabilities. These bacteria are categorized according to the carbon source used, as "obligate hydrocarbonoclastic" when they can grow only with a specific carbon source or "facultative hydrocarbonoclastic" when they are able to grow with alternative carbon sources. A number of these bacteria may yet be undiscovered [3].
Physical, chemical, and biological techniques have been developed to deal with the effects caused by the presence of oil in marine ecosystems. These techniques are intended to remove as much pollutant as possible and reduce the environmental impact generated from an oil spill or the progressive accumulation of oil [4]. Nowadays, of all applied techniques, bioremediation is considered the cheapest and highest yielding alternative for transforming pollutants in almost any ecosystem. In most marine environments, it is possible to use a large variety of bacteria with complex enzymatic machinery for the transformation of persistent xenobiotics. These bacteria might be in vitro isolated and may need to be improved in their metabolic capability if they have been exposed to the contaminant [5].
A bacteria’s ability to degrade petroleum hydrocarbons can be assessed from its taxonomy and physiology. The use of partial sequences of the 16S ribosomal gene (16S rRNA) allows the determination of a taxonomic location [6], while on the other hand it is possible to learn some ecophysiological properties about nutritional requirements and other factors by growing and handling the bacteria in vitro. The sequence analysis obtained might be useful for describing novel taxonomical groups with interesting physiologies for biotechnological processes useful in applied bioremediation. However, at this stage of research, even if the ecological impact and biotechnological applications are relevant, the diversity of hydrocarbonoclastic bacteria is underestimated.
The aim of this research was to simulate a marine model ecosystem contaminated with oil discharge and to characterize hydrocarbonoclastic marine bacteria using 16S rRNA sequences to contribute to the understanding of the diversity and potential uses of this metabolic group. A microcosm system for the assessment of bacterial responses to contamination by petroleum hydrocarbons was set up in the Laboratory of Microbial Biotechnology at the University of Nariño. In previous studies [7,8] a microcosm-like closed system was developed to obtain ecological data such as relevant bacterial community characteristics, in order to understand the most relevant succession processes in the bacterial community. In this research, these microcosm systems were simulated using an Upstream Flow Bioreactor (UFB), which was operated as a continuous system, supplying oil as the carbon and energy source and seawater as the mineral substrate.
2. MATERIALS AND METHODS
Bacterial strains and growth conditions
Type strains of Pseudomonas aeruginosa BUN010 and Escherichia coli BUN001 were used as positive controls for 16S rRNA PCR, available in the Laboratory of Microbial Biotechnology. The strains belong to the University of Nariño collection, Laboratory of Microbial Biotechnology, the strains were cultured in nutritive agar (NA, OXOID) and incubated at 25°C for 24 hours.
Oil degrading marine bacteria
The bacterial isolates were collected from the microcosm UFB system (Figure 1). The reactor consisted of a continuous culture operated with perfusion for an incubation time of 25 days. Then 10 mL of oil and 9250 mL of seawater were added in the ratio of 1%:0.2-0.4% as carbon and micronutrients sources respectively. The pH was measured as 7.5. The experiment was carried out at 25°C and the retention time was calculated as 5 days. The seawater and the inoculums employed in the UFB system consisted of 9250 mL of seawater not contaminated with hydrocarbons and 750 mL of water samples from different areas in the Bay of Tumaco reported to be contaminated with hydrocarbon concentrations as a result of oil discharges [8].
Bacteria were isolated from three different levels of sampling points from the UFB systems (Figure 1). Five mL samples were collected from each sampling point every five days. Isolation of marine bacteria was carried out using the serial dilution technique in physiological water 0.1%. Dilutions 10-3 to 10-7 were inoculated at 0.1 mL on the surface of plates with Brito's modified media [9]. The carbon source in this media was introduced by adding 1 mL.L-1 of diesel and gasoline, after the sterilization of the media. The inoculated plates were incubated aerobically at 25°C for 24 hrs.
The bacterial isolates were described by macroscopy in pure cultures growing in solid media and the microscopic characteristics were performed by Gram coloration and observations using optical microscopy. Bacterial isolates were purified with an inoculation loop from pure colonies in petri dishes with nutrient agar (OXOID) 15g.L-1, ensuring that the bacteria would grow to full coverage. The cultures were incubated at room temperature and were subcultured every 15 days to maintain the viability of the isolates. The strains were conserved in triplicate using three different methods: (1) In criotubes containing 500 mL of glycerol-physiological water solution 30:70 and 500 mL of strain cultures growing in nutrient broth (NB) 15g.L-1. The criotubes were stored at -20oC. (2) In tubes containing 500 mL of liquid culture in NB and sterile porcelain beads with silica gel (60:40 w/w), and (3) by adding pure colonies with a bacteriological loop to inclined tubes with nutrient agar.
Partial characterization of the 16S rRNA gene
Eleven bacterial isolates were identified using the 16S rRNA gene. The DNA template used for all PCR reactions was obtained from bacterial lysate [10] and the amplification of about 744 bp fragments, was carried out using a protocol proposed for environmental bacteria [10]. Conventional PCR amplification was performed in a final volume of 25 mL. The final concentrations of the PCR reaction mix were: 1X Buffer, 1.5 mM MgCl2, 0.18 mM dNTPs, 0.3 mM of each primer: A (forward): 5'-GGAGCAAACAGGATTAGATACCC-3' and J (reverse): 5'-TTCTCCTAGGGCTACC TTGTTAC-3' (MWG Biotech USA), and 0.05 U/mL of taq DNA polymerase (Fermentas). A total of 2.5 mL of DNA bacterial lysate were added as a template. All PCR reactions were performed in a Multigen Gradient (Labnet) thermocycler following the reference conditions [10]. Electrophoresis of PCR products was amplified using agarose gel (1%) for 130 minutes at 60 mV. Both pure DNA extracts and PCR products were conserved at -20 °C.
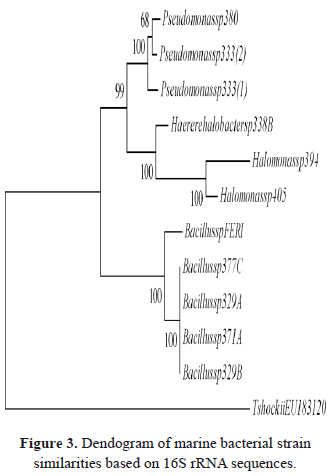
The sequencing process was carried out using purified PCR products. The amplicons were sent to the MACROGEN Company in Korea. The search for similarity amongst 16S rRNA gene partial sequences from bacterial strains was performed by alignment with the 16S rRNA sequence available in the public nucleotide databases at the National Center for Biotechnology Information (NCBI) by using its World Wide Web site (http://www.ncbi.nlm.nih.gov), as well as by using the BLAST algorithm [11], and the Ribosomal Database Project (RDP) with SeqMatch algorithm [12]. The 16S rRNA sequences from the bacterial strains were aligned using the ClustalW program [13]. The software MEGA 5.0 [14] was used to perform a dendogram using a similarity matrix as a base. The dendogram was createtd using the16S rRNA sequences by the Neighbor-Joining algorithm [15], using the P distance as the parameter. The matrix obtained was analyzed with Felsentein’ Bootstrap test.
3. RESULTS AND DISCUSSION
According to the methodology, all the bacterial isolates obtained in this study were identified as hydrocarbon-degrading marine bacteria. The bacteria were selected according to the physicochemical conditions observed in natural oil-contaminated seawaters. The simulated environment in the microcosm provided these conditions [8]: a temperature of 17-25oC, salinity (due to the basal salt medium) of 0.6%, and crude oil added at 1% as the sole carbon substrate source. The biomass remained in perfusion.
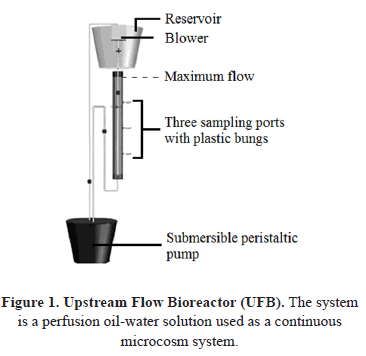
From the microcosm, 53 hydrocarbon-degrading bacterial isolates were obtained. From these, 11 strains were initially characterized according to their ability to use an aliphatic hydrocarbon (n-octane) derived from petroleum, as their sole carbon and energy source. Six of these bacteria were Gram negative and five were Gram positive. In general, the Gram negative group has been reported as the most representative group of hydrocarbon-degrading bacteria [16]. According to the literature, the lipopolysaccharide produced in bacterial membranes supports the formation and stabilization of oil in aqueous systems and contributes by increasing the attack surface on the pollutant for subsequent assimilation [17-18]. Novel metabolic pathways for polycyclic aromatic hydrocarbons (PAHs) with lower molecular weights such as phenanthrene and fluorene have recently been described in Gram negative bacteria, while on the other hand, a novel ortho-way has been identified at this point only in Gram positive bacteria [18]. Eleven bacterial isolates were characterized by 16S rRNA gene sequencing. Figure 2 shows the amplification of the gene fragment of the expected size according to the primers used. The fragment size was about 744 bp and non-specific bands were observed in the amplification.
According to the literature, there are several bacterial genera with the ability to degrade hydrocarbons in marine environments, such as Acinetobacter, Aeromonas, Bacillus, Burkholderia, Halomonas, Kocuria, Paracoccus, Pseudomonas, Rhodobacter, Rhodococcus, Staphylococcus, Vibrio [1], Alcanivorax, Marinobacter, and Cycloclasticus [19].
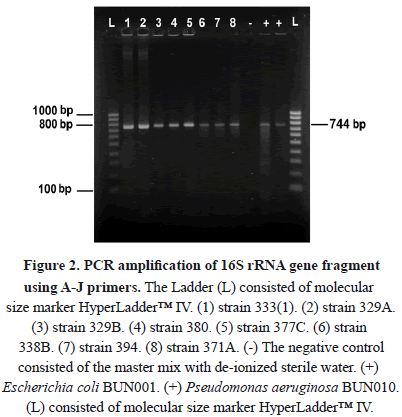
In this research, the analysis of 16S rRNA sequences indentified 4 genera (Table 1).
The strains 333(1), 333(2), and 380 showed 99% of similarity with the genus Pseudomonas (Table 1) and most probably correspond to P. aeruginosa for the strain 333(1), P. stutzeri for the strain 333(2), and P. putida for the strain (380). Pseudomonas species have been widely studied in the degradation of hydrocarbons. These species are known to degrade crude oil, diesel, and a variety of PAHs. P. aeruginosa, P. alcaligenes and P. putida are known for their efficiency in bioremediation [20-22]. Interestingly, P. aeruginosa is one of the species most frequently used in bioremediation research and presents several metabolic ways to deal with xenobiotics [23]. Pseudomonas bacteria are also rhamnolipid biosurfactant producers and are involve in oil decontamination [24].
The strains 329A, 329B, 377C, 371A, and FER1 presented similarities of between 99 and 100% in relation with species of the genus Bacillus, particularly B. simplex for the strains 329A, 329B, 371A, and 377C, and B. subtilis for the FER1 strain (Table 1). These species are widely distributed in areas with oil discharges due to their low nutritional requirements [25]. Bacillus is other dominant genera associated with hydrocarbon degradation besides the genus Pseudomonas [26, 27 y 28]. Some species of Bacillus have also been used in the bioremediation of contaminated marine environments [29], i.e. B. simplex is a well-known PAH reducer [30-31]. B. subtilis might reduce other complex hydrocarbons and paraffins [32] and produce rhamnolipid biosurfactants [33].
The strains 394 and 405 showed similarities of 87% and 89% respectively with Halomonas sp. (Table 1). Halomonas sp. is considered a "facultative hydrocarbonoclastic bacteria" and degrades phenanthrene or chrysene. However, Halomonas sp. doesn’t have the capability to degrade alkanes and some aromatic compounds such as toluene and naphthalene [34]. Interestingly this genus is reported to be an exopolymer synthesizer, and they can emulsify different hydrocarbons and use them as carbon and energy sources [35-36]. It has been shown that strains like Pseudomonas, Bacillus and Brevibacterium were more efficient as hydrocarbon-degrading marine bacteria, but on the other hand, Halomonas strains produced the highest amount of biopolymers with emulsifying activity using the same substrate [36].
The oil-degrading marine bacteria included here were grouped into 2 clusters (bootstrap tests presented 98% and 99% confidence values for each cluster), corresponding to the Proteobacteria and Firmicutes phylum (Figure 3). The distribution of marine oil-degrading bacteria agreed with the grouping recognized in the references [37-38]. The strains of Halomonas, Pseudomonas and Haererehalobacter obtained in this research were grouped into three clusters belonging to Proteobacteria phylum (the Gamma-proteobacteria class), which is now the largest and most diverse phylogenetic lineage. The strain identified as Bacillus sp. belongs to the Firmicutes phylum, and future studies are required to establish possible intraspecific diversity.
The strain 338B has shown 98% similarity to Haererehalobacter sp. (Table 1). While, there are a few studies reporting this genus to be a halophilic bacteria found in several marine-like ecosystems, with optimal growth using a salt solution of approximately 15% as a mineral source [39], no reports have yet been made about Haererehalobacter sp. as a hydrocarbonoclastic marine bacteria. For this reason, this research permits us to report for the first time the strain Haererehalobacter sp. 338B to be a marine hydrocarbonoclastic bacteria, isolated from the Colombian Pacific coast as a novel genus representative of this metabolic group.
4. CONCLUSIONS
This research characterized the Pseudomonas sp. strains 333(1), 333(2) and 380, Bacillus sp. strains 329A, 329B, 371A, 377C, and FER1, Halomonas sp. strains 394 and 405, and the strain Haererehalobacter sp. 338B. Therefore, the bioreactors provided the necessary conditions for the development of bacterial community growth. The presence of oil-degrading bacteria was observed in this community, and their potential biosurfactant production was deduced by hydrocarbon degradation. This study contributes significantly to the limited studies carried out in Colombia with respect to hydrocarbon-degrading bacteria in marine environments.
The strains selected in a mixed culture might have metabolic capabilities to tolerate high concentrations of hydrocarbons, absorb them, and degrade them. These characteristics are the starting points for further research about the metabolic diversity and intraspecific microbial interactions that may occur in a mixed culture. The mixed culture is likely related to the degradation of hydrocarbon pollutants; more research on this bioremediation application is necessary.
Through 16S rRNA sequentiation it was possible to identify a genus of marine hydrocarbonoclastic bacteria that has not been mentioned previously in the literature. We report in this research the strain Haererehalobacter sp. 338B as a new hydrocarbonoclastic marine bacteria, which may play an important role in the bioremediation of oil-contaminated seawaters on the Colombian Pacific coast.
ACKNOWLEDGMENTS
The authors thank engineer Roberto Garcia for his assistance in building the UFBs and Dr. Mariela Burbano for her advice on issues with partial sequences of the 16S rRNA gen. This research was mainly supported by COLCIENCIAS through Grant Project Number 110445221299 and the Vicerrectoría de Investigaciones, Posgrados y Relaciones Internacionales (VIPRI), University of Nariño.
REFERENCES
[1] Pucci, G. N., Acuña, A., Tonin, N., Tiedemann, M. C. and Pucci, O. H., Diversity of culturable bacteria capable of degrading hydrocarbons from the beach of Caleta Córdova, Argentina, Rev. Peru. Biol., 17, pp. 237-244, 2010. [ Links ]
[2] Garay, J.A., Gómez, D.I. y Ortiz, J. R., (Eds). Diagnóstico integral del impacto biofísico y socioeconómico relativo a las fuentes de contaminación terrestre n la bahía de Tumaco, Colombia y lineamientos básicos para un Plan de Manejo. Proyecto del Programa de las Naciones Unidas para el Medio Ambiente (PNUMA - Programa de Acción Mundial PAM) y Comisión Permanente del Pacífico Sur CPPS. Instituto de Investigaciones Marinas y Costeras INVEMAR- Centro Control Contaminación del Pacífico CCCP- Corporación Autónoma Regional de Nariño CORPONARIñO. Santa Marta, P. 290, 2006. [ Links ]
[3] Harayama, S., Kasai, Y. and Hara, A., Microbial communities in oil-contaminated seawater, Curr. Opin. Biotechnol., 15, pp. 205-214, 2004. [ Links ]
[4] Narváez-Flórez, S., Gómez, M. L. and Martínez, M. M., Selection of bacteria with hydrocarbon degrading capacity isolated from Colombian Caribbean sediments., Bol. Invest. Mar. Cost., 37, pp. 63-77, 2008. [ Links ]
[5] INVEMAR. Bacterias marinas nativas degradadoras de compuestos orgánicos persistentes en Colombia. Serie de publicaciones generales No. 19, Santa Marta: Colombia. Instituto de Investigaciones Marinas y Costeras, 2006. [ Links ]
[6] Janssen, P. H., Identifying the dominant soil bacterial taxa in libraries of 16S rRNA and 16S rRNA genes, Appl. Environ. Microbiol., 72, pp. 1719-1728, 2006. [ Links ]
[7] Syakti, A. D., Mazzela, N., Nerini, D., Guiliano, M., Bertrand, J. C., Doumenq, P., Phospholipid fatty acids of a marina sedimentary microbial community in a laboratory microcosm: Responses to petroleum hydrocarbon contamination, Organic Geochemistry., 37, pp. 1617-1628, 2006. [ Links ]
[8] Gómez, M. I., Concha, M. J., Bravo, D., Villota, G., Revelo, D. M. and Fernández, P., Variación de la comunidad bacteriana hidrocarburoclástica proveniente de la Ensenada de Tumaco- Nariño, bajo un sistema de fermentación continuo, Revista de la Asociación Colombiana de Ciencias Biológicas., 22, pp.148, 2010. [ Links ]
[9] Brito, E. M. S., Guyoneaud, R., Goñi-Urriza, M., Ranchou-Peyruse, A., Verbaere, A., Crapez, M.A.C., Wasserman, J. C. A. and Duran, R., Characterization of hydrocarbonoclastic bacterial communities from mangrove sediments in Guanabara Bay, Brazil, Res. Microbiol., 157, pp. 752-762, 2006. [ Links ]
[10] Revelo Romo, D, M., Grosso, M, V., Moreno, C, N. and Montoya, D., A most effective method for selecting a broad range of short and medium-chain-length polyhidroxyalcanoate producing microorganisms, Electron. J. Biotechnol., 10, pp. 48-357, 2007. [ Links ]
[11] Altschul, S. F., Madden, T. L., Schäffer, A. A., Zhang, J., Zhang, Z., Miller, W. and Lipman, D. J., Gapped BLAST and PSI-BLAST: a new generation of protein database search programs, Nucleic Acids Research., 25, pp. 3389-3402,1997. [ Links ]
[12] Cole, J. R., Wang, Q., Cardenas, E., Fish, J., Chai, B., Farris, R. J., Kulam-Syed- Mohideen, A. S., Mcgarrell, D. M., Marsh, T., Garrity, G. M., Tiedje, J. M., The Ribosomal Database Project: improved alignments and new tools for rRNA analysis, Nucleic Acids Res., 37, D141-145, 2009. [ Links ]
[13] Thompson, J. D., Higgins, D. G., Gibson, T. J. and Clustal, W., improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice, Nucleic Acids Research., 22, pp. 4673-4680, 1994. [ Links ]
[14] Tamura, K., Peterson, D., Peterson, N., Stecher, G., Nei, M., Kumar, S., MEGA5: Molecular Evolutionary Genetics Analysis using Maximum Likelihood, Evolutionary Distance, and Maximum Parsimony Methods. Mol. Biol. Evol., 28, pp. 2731-2739, 2011. [ Links ]
[15] Saitou, N. and Nei, M., The neighbor-joining method: a new method for reconstructing phylogenetic tres, Mol. Biol. Evol., 4, pp. 406-425, 1987. [ Links ]
[16] Ruberto, L., Vázquez, S. and Mac Cormack, W., Effectivenness of the natural bacterial flora. Biostimulation and bioaugmentation on the bioremediation of a hydrocarbon contaminated antartic soil, International Biodeterioration & Biodegradation., 52, pp. 115-125, 2003. [ Links ]
[17] Sikkema, J., Bont, J. and Poolman, B., Mechanisms of membrane toxicity of hydrocarbons, Microbiol. Rev., 59, pp. 201-222, 1995. [ Links ]
[18] Van Hamme, J. D., Singh, A. and Ward, O. P., Recent advances in petroleum microbiology, Microbiol. Mol. Biol. Rev., 67, pp. 503-549, 2003. [ Links ]
[19] Yakimov, M. M., Timmis, K. N. and Golyshin, P. N., Obligate oil-degrading marine bacteria, Curr. Opin. Biotechnol., 18, pp. 257-266, 2007. [ Links ]
[20] Zhang, G-L., Wu, Y-T., Qian, X-P. and Meng, Q., Biodegradation of crude oil by Pseudomonas aeruginosa in the presence of rhamnolipids, J. Zhejiang. Univ. SCI., 6B, pp. 725-730, 2005. [ Links ]
[21] Ting, A. S. Y., Fang, M. T. and Tee, C. S., Evaluation on the efficacy of formulated Pseudomonas cells in degrading the aromatic hydrocarbon toluene, Aust. J. Basic & Appl. Sci., 4, pp. 3122-3127, 2010. [ Links ]
[22] Ting, A. S, Y., Tan, C. H. C., Aw, C.S., Hydrocarbon-degradation by isolate Pseudomonas lundensis UTAR FPE2, Mal. J. Microbiol., 5, pp. 104-108, 2009. [ Links ]
[23] Benavides, J. B., Quintero, G., Guevara, V. A., Jaimes, D. C., Gutiérrez, S. M. y Garcia, J. M., Biorremediación de suelos contaminados con hidrocarburos derivados del petróleo, Nova. Publicación Científica., 4, pp. 82-90, 2006. [ Links ]
[24] Benincasa, M., Contiero, J., Manresa, M. A. and Moraes, I. O., Rhamnolipid production by Pseudomonas aeruginosa LBI growing on soapstock as the sole carbon source, J. Food. Eng., 54, pp. 283-288, 2002. [ Links ]
[25] Cybulski, Z., Dziurla, E., Kaczorek, E., Olszanowski, A., The influence of emulsifiers on hydrocarbon biodegradation by Pseudomonadacea and Bacillacea strains, Spill Science & Technology Bulletin., 8, pp. 503-507, 2003. [ Links ]
[26] Queiroga, C., Nascimento, L. R. and Serra, G. E., Evaluation of paraffins biodegradation and biosurfactant production by Bacillus subtilis in the presence of crude oil, Brazilian J. Microbiol., 34, pp. 321-24, 2003. [ Links ]
[27] Zhao, J., Jian, L., Liang, H. Y., Qiang, J., Ying, Z. W., Xia, H., Biodegradation of toxic organic pollutants by Bacillus sp. LY with heterotrophic nitrogen removal ability, Huanjing-Kexue., 28, pp. 2838-42, 2007. [ Links ]
[28] Gómez, W., Gaviria, J., Cardona, S., Assessment of the biostimulation against bioaugmentation and natural attenuation on contaminated soil with diesel-gasoline mixing, Dyna., 160, pp. 83-93, 2009. [ Links ]
[29] Chikere, C. B., Okpokwasil, I G. C., Ichiakor, O., Characterization of hydrocarbon utilizing bacteria in tropical marine sediments, Afr. J. Biotechnol., 8, pp. 2541-2544, 2009. [ Links ]
[30] Juhasz, L. and Naidu, R., Bioremediation of high molecular weight polycyclic aromatic hydrocarbons: a review of the microbial degradation of benzo[a] pyrene, International Biodeterioration & Biodegradation., 45, pp. 57-88, 2000. [ Links ]
[31] Nair, D., Fernández, F. J., García, E., Riba, I. and Del Valls, T. A., Isolation and characterization of naphthalene-degrading bacteria from sediments of Cadiz area (SW Spain), Environ. Toxicol., 23, pp. 576-582, 2008. [ Links ]
[32] Naggar, A. Y., Kamel, M. M., Aladly, A. A. and Ismail, N. A., Bioremediation of paraffinic and polynuclear aromatic hydrocarbons using laser irradiated Bacillus amyloliquefaciens, Journal of American Science., 6, pp. 661- 670, 2010. [ Links ]
[33] Nwaogu, L. A., Onyeze, G. O. C., Nwabueze, R. N., Degradation of diesel oil in a polluted soil using Bacillus subtilis. Afr. J. Biotechnol., 7, pp. 1939-1943, 2008. [ Links ]
[34] Melcher, R. J., Apitz, S. E. and Hemmingsen, B. B., Impact of irradiation and polycyclic aromatic hydrocarbon spiking on microbial populations in marine sediment for future aging and biodegradability studies, Appl. Environ. Microbiol., 68, pp. 2858-2868, 2002. [ Links ]
[35] Calvo, C., Martinez, F., Toledo, F. L., Porcel, J. and Quesada, E., Characteristics of bioemulsifiers synthesised in crude oil media by Halomonas eurihalina and their effectiveness in the isolation of bacteria able to grow in the presence of hydrocarbons, Appl. Microbiol. Biotechnol., 60, pp. 347-351, 2002. [ Links ]
[36] Uad, I., Silva, G. A., Pozo, C., González, J. and Calvo, C., Biodegradative potential and characterization of bioemulsifiers of marine bacteria isolated from samples of seawater, sediment and fuel extracted at 4000 m of depth (Prestige wreck), International Biodeterioration & Biodegradation., 64, pp. 511-518, 2010. [ Links ]
[37] Kersters, K., De Vos, P., Gillis, M., Swings, J., Vandame, P. and Stackebrandte, E., Introduction to the Proteobacteria, Prokaryotes., 5, pp. 3-37, 2006. [ Links ]
[38] Velmurugan, N., Kalpana, D., Cho, J-Y., Lee, G-H., Park, S-H. and Lee, Y-S., Phylogenetic analysis of culturable marine bacteria in sediments from South Korean Yellow Sea, Microbiol. 80, pp. 261-272, 2011. [ Links ]
[39] Wang, X., Chi, N-Y., Zhang, Q-F., Research on the isolation, identification and characteristics of a moderately halophilic bacterium wich from Liaodong Bay, Advanced Materials Research., 183-185, pp. 1085-1089, 2011. [ Links ]