Introduction
The admission of patients to the intensive care unit (ICU) has an effect on the frequency of infections caused by strains of multi-resistant Gram-negative bacilli (MRGN). Some risk factors increase the susceptibility to MRGN infections, including those caused by extended spectrum beta-lactamases (ESBL) producing strains. Some of these risk factors include immunosuppressive treatments in diseases such as cancer 1, transplant interventions 2, the implementation of central or urinary catheters 3, hemodialysis procedures 4, and mechanical ventilation devices. 5
Klebsiella pneumoniae and Escherichia coli are Gram-negative bacilli that are present in the intestinal tract of mammals, and their colonization rates are trebled in hospital environments in a direct proportion to the length of the stay. Furthermore, their colonization degree has been associated with the selective pressure exerted by antibiotics on intestinal microbiota. 6,7
These microorganisms are potentially pathogenic for ICU patients undergoing mechanical ventilation since, after colonizing the supraglottic region, they adhere to both inert surfaces and epithelial cells and trigger ventilator-associated pneumonia (VAP) in order to resist desiccation and β-lactam antibiotics respectively. 8,9
Currently, four pathogenic pathways for VAP to occur have been described: by aspiring secretions through the oropharynx, by contiguity, by entering into contact with blood (hematogenous route), and through breathing circuits. Aspiration of secretions through the oropharynx is the most common pathway 10, therefore biofilms containing respiratory pathogenic microorganisms from the supraglottic region constitute a source of pneumonia associated with health care.
Microbiological analysis by molecular methods performed on the tongue and from bronchoalveolar lavage has evidenced the presence of a several bacterial species. 11 Some tests have suggested that these microorganisms are able to act as pathogens that affect the lower respiratory tract. 11 The isolation of Staphylococcus aureus, Pseudomonas aeruginosa, Acinetobacter spp, and enterobacteria from dental plaque in patients with pneumonia associated with health care, indistinguishable of isolates obtained from bronchoalveolar lavage, reinforces this idea. 11
This study was made from the data obtained in a prior research where the effectiveness of chlorhexidine mouthwash was compared to toothpaste mouthwash in terms of decontamination of the bacterial microbiota in the supraglottic region. In addition, reduction of the supraglottic region bacterial microbiota in patients undergoing mechanical ventilation at the ICU was evaluated. After 72 hours in the ICU, colonization of the oropharynx by normal microbiota was reduced, while pathogenic microbiota increased, being K. pneumoniae and E. coli the most common microorganisms. 12 Some of these strains were preserved in order to determine, as it is proposed here, their susceptibility against antibiotics, as well as their molecular profiles.
Considering the above mentioned, this study focused on the molecular characterization and detection of ESBL-producing K. pneumoniae and E. coli strains isolated from the supraglottic region of patients undergoing mechanical ventilation in an ICU.
Materials and methods
Type of study and sample unit
A descriptive study was carried out. Microorganisms were preserved in the Collection of Microorganisms of Universidad Católica de Manizales (CM-CUM).
The data was collected from a prior study conducted in patients undergoing mechanical ventilation at the ICU of a hospital in Manizales, Colombia), between 2013 and 2014. The sample consisted of 18 isolates of K. pneumoniae and 7 isolates of E. coli obtained from the supraglottic region of said patients. In addition, 5 isolates of K. pneumoniae obtained from blood cultures, external secretion and peritoneal fluid of patients admitted to the same ICU, but not included in the first study, were also considered for molecular analysis.
Assessment of response to antimicrobials and ESBL production
The preserved strains were initially grown on nutrient agar and were subsequently inoculated on a selective medium (MacConkey agar). These preserved strains were confirmed in terms of gender and species through the automated system VITEK® 2 compact (BioMérieux) at the UCM laboratory.
After confirming the identification, antimicrobial susceptibility was determined by means of the disk diffusion method on selective medium (Mueller Hinton Agar), in accordance with the recommendations of the Clinical and Laboratory Standards Institute (CLSI), 13 including the following antibiotics for detecting possible ESBL-producing strains: aztreonam, cefotaxime, cefotetan, ceftazidime and ceftriaxone.
For preparing the inoculum, 4 to 5 colonies of similar morphology were selected from each culture and were homogenized in 5mL of saline solution, while the resulting turbidity in the solution was adjusted by using sterile saline solution until a density equivalent to 0.5 on the McFarland standard was achieved. Within 15 minutes after adjusting the inoculum, the spread plate technique on a Mueller Hinton agar was performed by using a sterile swab, and then, antibiotic disks were placed on the surface of the inoculated agar through sterile forceps. The disks were distanced at a 24mm average length. Afterwards, plates were incubated at 37°C for 16 to 18 hours. Finally, diameters of inhibition zones were measured according to the CLSI criteria. 13 To confirm ESBL-producing strains, double-disk diffusion method was used in accordance with the the CLSI standard and the following breakpoints: cefpodoxime (≤22mm), aztreonam (≤27mm), ceftazidime (≤22mm), cefotaxime (≤27mm) or ceftriaxone (≤25mm) 13.
ESBL production by the strains studied was confirmed through growth measurement against cefotaxime and ceftazidime alone and in combination with clavulanic acid. A ≥5mm difference between the diameter of the disk where ceftazidime + clavulanic acid were used and the disk where only ceftazidime was used, or between the disk were cefotaxime + clavulanic acid were applied and the one where only cefotaxime was used, was considered as ESBL production.
Genotyping by BOX-PCR system
Bacterial DNA was extracted by using the UltraClean® Blood DNA Isolation Kit (NON-SPIN), which was provided by MOBIO laboratories, Inc. Subsequently, the DNA obtained was stored in Eppendorf tubes at -20°C until its amplification by polymerase chain reaction (PCR) was done.
BOX-PCR typing was performed with the AR1-5'-CTACGGCAAGGCGACGCTGACG-3' primer under the following conditions: 100ng of DNA were amplified in a mini thermal cycler (Bio Rad®) and were arranged in a reaction volume of 25μL, (14,15) which contained 0.2mM of DNTPs, 2mM of MgCl2, 1.5μM of primer, 0.10mg/mL of BSA, 10% of DMSO and 1U/μL of Taq DNA polymerase (BIOLINE®). For the BOX-PCR amplification, first the DNA was denatured for 5 minutes at 95°C and then underwent 30 denaturation cycles (92°C, 30 sec), an association cycle (60°C, 1 min), and one extension cycle (65°C, 8 min), which was followed by a final extension cycle at 65°C for 8 min. Amplified products were separated by electrophoresis on 2% agarose gel with 1.0 X TBE buffer, for 3.5 h at 4.6 V/cm. Banding patterns were recorded in photos 14,15.
Finally, visual inspection of electrophoretic profiles was done and matrices of presence and absence of bands for the K. pneumoniae and E. coli isolates were constructed by using Microsoft Office Excel software.
Statistical analysis
The analysis of the matrices was done by using NTSYSpc 2.2 version software. A dendrogram with the unweighted pair group method with arithmetic averages algorithm and Jaccard similarity coefficient was created.
The similarity coefficient (S) was obtained based on the following equation: S=2*Nab/a+b, where Nab is the total number of similar bands between 2 isolates and a+b is the total sum of the number of bands of isolates a and b.
Ethical considerations
The microorganisms used in this study were collected in a prior research that was approved by the Research Ethics Committee of the Universidad Católica de Manizales on April 18, 2012, this approval was updated in September 2018 by this committee, it should be noted that due to the university policies the approvals issued by said ethics committee are not numbered. In addition, participants' informed consent was duly obtained in said research.
Results
K. pneumoniae response to antimicrobials and ESBL production
Out of the 18 K. pneumoniae strains analyzed, sensitivity to aztreonam, cefotaxime, ceftriaxone, cefotetan and ceftazidime was observed in 11 (61 %), while the 7 remaining strains (3 9%) were resistant to these antibiotics. Intermediate sensitivity was not observed in any of the isolations (Figure 1).
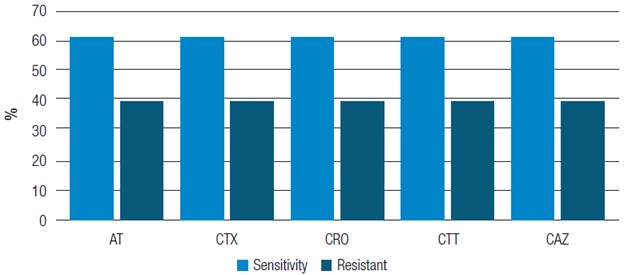
Source: Own elaboration
Figure 1 Percentage of sensitivity of Klebsiella pneumoniae strains against different antibiotics. At: Aztreonam; CTX: Cefotaxime; CRO: Ceftriaxone; CTT: Cefotetan; CAZ: Ceftazidime.
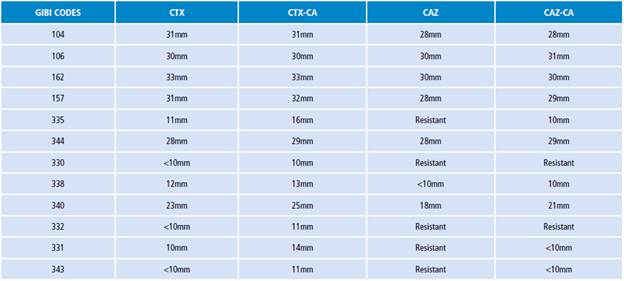
Then, 12 potentially ESBL-producing strains were selected and they were assigned the following GIBI codes: 104, 106, 162, 157, 335, 344, 330, 338, 340, 332, 331, and 343. Through the disk diffusion technique, an ESBL test was done for each of the strains, the results are as follows: 11 negative isolates and 1 ESBL-producing strain (Table 1).
E. coli response to antimicrobials and ESBL production
Out of 7 E. coli strains analyzed, 6 (86%) showed sensitivity to aztreonam, cefotaxime, ceftriaxone, cefotetan, and ceftazidime. Intermediate sensitivity was not observed in any of the isolations (Figure 2).
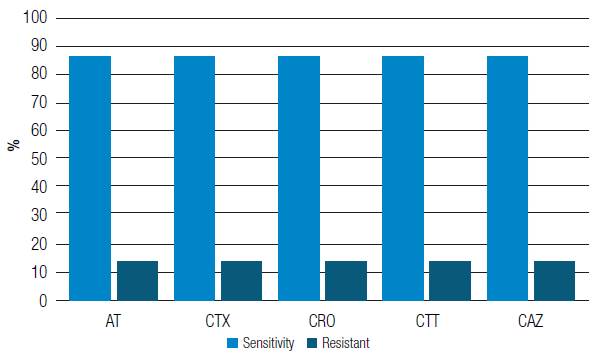
Source: Own elaboration
Figure 2 Percentage of sensitivity of Escherichia coli strains against different antibiotics. At: Aztreonam; CTX: Cefotaxime; CRO: Ceftriaxone; CTT: Cefotetan; CAZ: Ceftazidime.
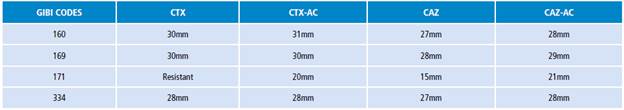
Later, 4 potentially ESBL-producing strains were selected and were given the following GIBI codes: 160, 169, 171, and 334. Results after performing the ESBL production confirmatory test were as follows: negative for 3 isolates, and positive for 1 strain (Table 2).
Electrophoretic profile of K. pneumoniae strains
Regarding the results obtained through the BOX-PCR fingerprinting technique, out of23 K. pneumoniae isolates, 2 groups with a divergence greater than 50% were observed. In the first group, constituted by 12 isolates, 3 subgroups were identified: subgroup 1, with isolates 62 and 344; subgroup 2, with isolates 72 and 331, and subgroup 3, where isolates 73, 104, 102, 92, 332, 335, 330, and 112 were found. It is noteworthy that isolate 112 had a divergence greater than 50% in comparison with the other isolates of this subgroup. In addition, a 100% similarity between isolates 73 and 104 was observed, as well as between isolates 92, 332, and 335, while similarity among other isolates was less than 90% (Figure 3).
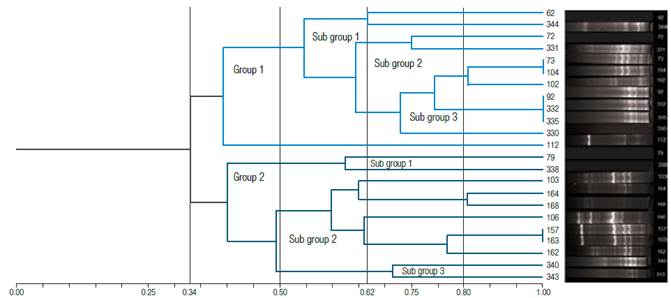
Source: Own elaboration
Figure 3 Cluster analysis to determine similarity among Klebsiella pneumoniae strains and electrophoretic profiles according to the BOX-PCR test.
On the other hand, in the second group, made up by 11 isolates, 3 subgroups were found: isolates 79 and 338 in subgroup 1, isolate 103 in subgroup 2, and strains 164, 168, 106, 157, 163, 162, 340, and 343 in subgroup 3. A 100% similarity between isolates 157 and 163 was observed, while similarity among other isolates in the remaining subgroups was less than 90% (Figure 3).
Electrophoretic profile of E. coli strains
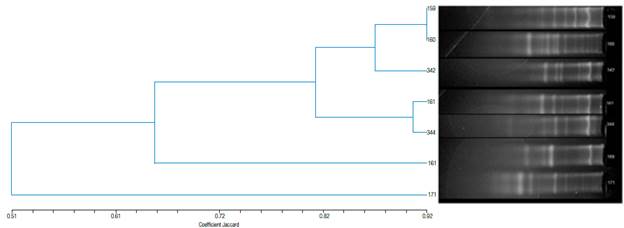
According to the results obtained after performing the BOX-PCR test, a divergence greater than 50% in the 7 isolates was found. A 92% similarity between isolates 159 and 160 was observed, while for isolates 161 and 334 a 91% similarity was reported (Figure 4).
Discussion
In this study one type of ESBL-producing strain was identified. In Colombia, higher percentages of ESBL-producing K. pneumoniae and E. coli strains have been reported. For example, in a study conducted in 22 K. pneumoniae isolates obtained from patients with a hospital-acquired infection, 15 were resistant to ceftazidime, cefotaxime, ceftriaxone and aztreonam. Besides, all the 15 strains were confirmed as ESBL-producing. 16 Likewise, another research conducted on 144 isolates of E. coli and K. pneumoniae, obtained from patients staying at several hospitalization services, reported that 48.6% of the total sample were resistant to any cephalosporin, while 25.6% of the E. coli and 48.4% of the K. pneumoniae strains were ESBL-producing. 17 Another study carried out in a hospital in Valledupar (Colombia) reported a 12.3% ESBL-producing enterobacteria, with E. coli and K. pneumoniae as the most frequent strains (55.6% and 23.2%, respectively). 18
Prior studies in Colombia have focused on strains with an infectious origin, while in this research isolates were obtained from the microbiota of the supraglottic region. Considering their origin, the isolates were not expected to be resistant to third generation cephalosporins or ESBL producers, which explains why the results obtained here indicate that the prevalence of ESBL-producing strains was lower than those reported in previous studies. Furthermore, resistance to third generation cephalosporins by strains from the supraglottic region may imply that there is a possibility of other resistance mechanisms such as alteration of target sites related to the penicillin-binding protein (PBP), reduction of porin-mediated outer membrane permeability, and efflux pumps. 9
It has also been reported that dissemination in the hospital environment occurs mainly because of the presence of strains of endogenous origin that are associated with each patient's microbiota. 16 Moreover, the resistance observed in the strains obtained from the supraglottic region could be explained by the fact that treatments against various infections are made through empiric therapy, namely, without knowing the etiology and antimicrobial susceptibility of the pathogen involved in the infection. Likewise, other inappropriate practices such as self-medication promote the development of resistance mechanisms. When these factors come together, any type of resistance to be acquired will have an effect on the effectiveness of antimicrobial treatments, thus generating therapeutic limitations in hospitals.
On the other hand, the 100% genotypic similarity between some of the strains studied here suggests a common intrahospital contamination source, as well as some cross contamination events, since patients' dates of admission and stays at the ICU were different.
The results of this study indicate the need for controlling ESBL-producing strains from the supraglottic microbiota of patients referred to the ICU. One strategy to control said strains is to use routine surveillance cultures, which also includes an educational intervention for the staff involved. 19 Regarding this approach, it is necessary to raise the awareness on the possibility of using microbial culture on a regular basis as an alert tool, especially if there is enough evidence to consider that some patients may be hosts of ESBL-producing strains prior to their admission to the hospital or the ICU.
In this sense, Boyer et al.20 reported the need to eliminate circulating strains in the environment. Therefore, monitoring potential sources of contamination that are usually underestimated, such as sinks, is an appropriate practice, for sometimes these sources may contain ESBL-producing bacteria and their elimination helps reducing the incidence of multiresistant strains in any ICU. 21 In that regard, the installation of self-disinfecting siphons in sinks of ICUs has been proved to remove biofilm formation and reduce transmission of ESBL-producing Gram negative bacteria. 22
Experts on this matter agree that genotyping by BOX-PCR technique offers an adequate discriminatory power, as well as reproducibility, and that is a fast typing alternative to get fast results. Furthermore, it requires less investment in comparison with other techniques such as gel electrophoresis by pulsed fields (PFGE) and DNA sequencing. 3 Somehow, a more efficient approach involving the identification of differential resistance genes could include matrix-assisted laser desorption/ionization time-of-flight mass spectrometry (MALDI-TOF MS), considering the reduced times it requires and the possibility of obtaining specific mass spectra of genus and species it offers. 23
Conclusions
A greater than 50% divergence in the molecular profiles among the strains groups of K. pneumoniae and among those of E. coli was determined. This polyclonality indicates that most of the strains belonged to the microbiota of each patient.
The isolation of K. pneumoniae and E. coli resistant to third generation cephalosporins and ESBL-producing strains in the ICU had as its main source the microbiota of each patient. Cases of genotypic similarity of 100% suggest a common intrahospital contamination source. These cases involved cross contamination events, since the date of admission and the time of permanence of patients in the ICU were different.
Considering their pathogenicity mechanisms, the presence of E. coli and K. pneumoniae resistant to β-lactam antibiotics strains in the supraglottic region could trigger persistent infections such as VAP, which is considered as a major financial challenge for the maintenance of public health.