Services on Demand
Journal
Article
Indicators
-
 Cited by SciELO
Cited by SciELO -
 Access statistics
Access statistics
Related links
-
 Cited by Google
Cited by Google -
 Similars in
SciELO
Similars in
SciELO -
 Similars in Google
Similars in Google
Share
Biomédica
Print version ISSN 0120-4157On-line version ISSN 2590-7379
Biomédica vol.31 no.1 Bogotá Jan./Mar. 2011
COMUNICACIÓN BREVE
1 Grupo de Microbiología, Instituto Nacional de Salud, Bogotá, D.C., Colombia
2 Hospital Erazmo Meoz, Cúcuta, Colombia
Recibido: 04/03/10; aceptado:26/08/10
Introduction. In Cúcuta, Cryptococcus gattii serotype B is commonly recovered from immunocompetent patients with cryptococcosis, but it has not been recovered from the environment in spite of its high incidence which is 77% out of reported cases.
Objective. The aim of this work was to carry out an extensive environmental sampling in Cúcuta, in an attempt to isolate C. gattii serotype B and to expand our knowledge about the ecology and epidemiology of this important yeast.
Materials and methods. Samples associated with 3,634 trees from 40 zones of Cúcuta were collected and processed with 28 samples collected near the houses of four patients with cryptococcosis caused by C. gattii serotype B. The serotype of the recovered isolates was done using multiplex PCR, molecular patterns were determined by RFLP of the URA5 gene and mating type was determined using the primers MfαU, MfαL, MFa2U and MFa2L.
Results. In total, 4,389 samples were processed and one isolate of C. gattii serotype B (VGI/a), two isolates of C. gattii serotype C (VGIII/α) and three isolates of C. neoformans var. grubii, serotype A (VNI/α), were recovered. The density of the recovered isolates varied from 50 to 350 cfu/g of soil.
Conclusions. This is the first report on the environmental isolation of C. gattii serotype B from Cúcuta. However, because of the low rate of recovery of isolates from soil only, the environmental niche of C. gattii has not been established and further environmental studies in Cúcuta are necessary, owing that this serotype is not only causing cryptococcosis but also has shown a higher virulence after the Vancouver outbreak.
Key words: Cryptococcus, mycoses, environment, ecology, Colombia, sampling studies.
Primer aislamiento ambiental de Cryptococcus gattii de serotipo B, en Cúcuta, Colombia
Introducción. En Cúcuta, Cryptococcus gattii de serotipo B se recupera comúnmente de pacientes inmunocompetentes con criptococosis, pero no ha sido recuperado del ambiente a pesar su alta incidencia, que es de 77 % de los casos reportados.
Objetivo. El objetivo de este trabajo fue realizar un extenso muestreo ambiental en Cúcuta, para aislar C. gattii de serotipo B y expandir nuestro conocimiento sobre la ecología y la epidemiología de esta importante levadura.
Materiales y métodos. Se recolectaron y se procesaron muestras asociadas con 3.634 árboles de 40 zonas de Cúcuta y 28 muestras cercanas a las casas de cuatro pacientes con criptococosis causada por C. gattii de serotipo B. El serotipo de los aislamientos recuperados se estableció usando PCR múltiple, el patrón molecular se determinó por RFLP del gen URA5 y la pareja sexual se determinó usando los iniciadores MfαU, MfαL, MFa2U y MFa2L.
Resultados. Se procesaron 4.389 muestras y se recuperó un aislamiento de C. gattii de serotipo B (VGI/a), dos aislamientos de C. gattii de serotipo C (VGIII/α) y tres aislamientos de C. neoformans var. grubii de serotipo A (VNI/α). La densidad de los aislamientos recuperados varió entre 50 y 350 UFC/g de suelo.
Conclusiones. éste es el primer reporte de la recuperación ambiental de C. gattii de serotipo B en Cúcuta. Sin embargo, por la baja tasa de recuperación de aislamientos únicamente en suelo, el nicho ambiental de C. gattii no se pudo establecer, por lo que es necesario realizar posteriores estudios ambientales en Cúcuta, debido a que este serotipo, no sólo está causando criptococosis, sino también muestra una gran virulencia después del brote ocurrido en Vancouver.
Palabras clave: Cryptococcus, micosis, ambiente, ecología, Colombia, muestreo.
Cryptococcus neoformans/Cryptococcus gattii species complex comprises a group of capsulated yeasts causing cryptococcosis in humans and animals worldwide (1). C. neoformans var. grubii, serotype A and C. neoformans var. neoformans, serotype D, have been mostly isolated from immunocompromised host with disseminated cryptococcosis, while C. gattii, serotypes B and C is most often associated with neurological disorders in hosts with normal immunity (2-4) and with disease in domestic animals (5-7). Infections are mainly acquired via inhalation of basidiospores from the environment, which are the principal infectious propagules for the C. neoformans/ C. gattii species complex (8).
In Colombia, C. neoformans var. grubii is the most important cause of cryptococcosis, followed by C. gattii serotype B (9), which is mainly recovered in patients from Cúcuta, where this serotype is the most frequently isolated in immunocompetent patients (10). However, while C. gattii serotype C, has been isolated in Cúcuta from almond trees (Terminalia catappa) (11), C. gattii serotype B has not been recovered from the environment in spite of the high incidence of this serotype associated with cryptococcosis, which is 77% out of the reported cases in immunocompetent patients (9,10). Additionally, the vast majority of clinical isolates of C. gattii serotype B recovered in Cúcuta since 1990, belong to the molecular type VGII (12), the same molecular type of the high virulence isolates associated with the Vancouver Island outbreak (13), where an increasing number of cases of cryptococcosis in both human and animals caused by C. gattii serotype B have been reported (14).
On the other hand, C. gattii serotype B has been isolated several times from different environmental sources in the world (14-17); for that reason, the aim of this work was to carry out an extensive environmental sampling survey in different areas of the city of Cúcuta, in an attempt to isolate C. gattii serotype B from different trees, to determine its circulation and to expand our knowledge about the ecology and the epidemiology of this pathogenic yeast in our country.
Materials and methods
Study area
Cúcuta is the capital city of the department Norte de Santander, in the north-east of Colombia, limiting with Venezuela. It is situated at 7°54´48" north latitude and 72°30´36" west longitude, at 320 m above sea level and the average temperature is 28°C with an annual rainfall of 763 mm approximately (18,19).
Sample collection
Samples including soil (n= 3,634), bark (n= 358), detritus (n= 225), leaves (n= 103) and fruits (n= 69) were collected and processed from 3,634 trees from Cúcuta, including, mainly, the species T. catappa (almond tree), Ficus sp., Licania tomentosa (oiti), Pithecellobium dulce, Fagara rhoifolia, Melicoccus bijugatus (mamón), Enterolobium cyclocarpum and Mangifera indica (mango). No swab samples were taken. Extensive collections were made in January, May and August 2008, from 40 zones of the city, which were defined by map grids (18) and included both urban and rural areas. Data of temperature, climatic conditions as rainfall and relative humidity, and season of the year were considered for the analysis of the results (19). In the zones where either C. neoformans or C. gattii was found, a sampling was made monthly from August 2008 to February 2009, to increase the number of isolates recovered. Additionally, 28 samples were collected from 22 trees near the houses of four patients with cryptococcosis caused by C. gattii serotype B molecular type VGII.
Sample processing
Five grams of each sample were suspended in 25 ml of phosphate buffer saline (PBS), mixed and filtered with sterile gauze. Fifteen ml of the filtrate were mixed with 50 µl of a solution of penicillin (20 U) and streptomycin (40 U) and 100 µl of this suspension was plated in Guizotia abyssinica agar, supplemented with penicillin and streptomycin (20). Plates were incubated at 27°C for one month, with weekly observation. Brown colonies grown in the media were isolated and transferred to a new G. abyssinica agar and next to Sabouraud dextrose agar for their identification using conventional techniques (21). Colony forming units (cfu) were counted and their number calculated and expressed per gram of sample, considering the volume plated and the dilutions of the filtrates. Additionally, all colonies from the same sample were taken to identify C. neoformans/ C. gattii isolates.
Typing
DNA of the isolates of the C. neoformans/C. gattii species complex recovered from the environment was extracted as previously described (22). Serotype was determined using the multiplex PCR described by Ito-Kuwa et al., with some modifications (23). The primers LAC1 No 2 and LAC1 No 4 and the new primers LAC1 No 5 (5´-CAGCATTGTGAGCGCTTTTCCG) and LAC1 No 6 (5´-GCATTACTGTGAGTGTACTTATAG) were used and PCR amplification was initiated at 94°C for 4 min, followed by 40 cycles of 94°C for 30 s, 46°C for 40 s and 72°C for 90 s, and then 7 min of extension at 72°C, in a thermal cycler MJ Mini (Bio-Rad Laboratories). The amplified products were separated by 2% agarose gel electrophoresis at 100 V for 1 h and the gels were stained with ethidium bromide and photographed under ultraviolet light. Strains WM 628 serotype A, WM 629 serotype D, H0058-I-455 serotype C and H0058-I-2976 serotype B were used as controls.
Molecular type and mating type determination
Restriction fragment length polymorphism (RFLP) of the URA5 gene was used as the molecular typing technique as previously reported by Meyer, et al (24). The URA5 gene was amplified with two primers: URA5 (5´ATGTCCTCCCAAGCCCTCGACTCCG3´) and SJ01 (5´TTAAGACCTCTGAACACCGTACTC3´). The amplification products were double digested with SAU961 (10 U/µl) and HhaI (20U/µl) overnight and separated by 2.5% agarose gel electrophoresis at 80 V for 3.5 h using a 50 bp molecular size marker (Promega). RFLP patterns were assigned by comparing them with the eight molecular types VNI -VNIV and VGI - VGIV (24,25). Mating type was determined using the primers MfαU, MfαL, MFa2U and MFa2L and PCR reactions and amplifications were performed as previously published (26).
Results
In total, 4,389 samples were processed from the 3,634 trees studied and among all samples processed, three (0.07%) were positive for C. gattii and three (0.07%) for C. neoformans.
From the 3 samples positive for C. gattii,seven C. gattii serotype C, molecular type VGIII isolates and one C. gattii serotype B, molecular type VGI isolate were recovered (Figure 1). From the 3 samples positive for C. neoformans, 13 C. neoformans var. grubii, serotype A, molecular type VNI isolates were recovered. Additionally, all C. neoformans var. grubii and C. gattii serotype C isolates were mating type alpha, while the only C. gattii serotype B isolate was mating type a.
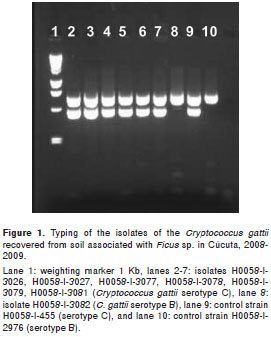
Among the recovered isolates, in January 2008, five colonies (250 cfu/g) of C. neoformans var. grubii and one colony (50 cfu/g) of C. gattii serotype B were recovered simultaneously from the same sample of soil associated with a Ficus sp. tree in the San Eduardo zone. In May 2008, one colony (50 cfu/g) of C. gattii serotype C was recovered in La Libertad zone from soil, and, in the Estadio General Santander, one colony (50 cfu/g) of C. neoformans var. grubii and six colonies (300 cfu/g) of C. gattii serotype C, were found simultaneously in the same sample of soil in a tree of Ficus sp. Due to these findings, the Estadio General Santander, San Eduardo and La Libertad zones were sampled again in August 2008. In this last sampling, only seven colonies (350 cfu/g) of C. neoformans var. grubii were recovered from soil in the Estadio General Santander (Table 1), while C. gattii was not found. The San Eduardo zone was sampled one more time in February 2009, because of the finding of C. gattii serotype B, but no isolates were recovered, and among the 28 samples collected from the houses of the four patients with cryptococcosis produced by C. gattii serotype B, no isolates were recovered either.
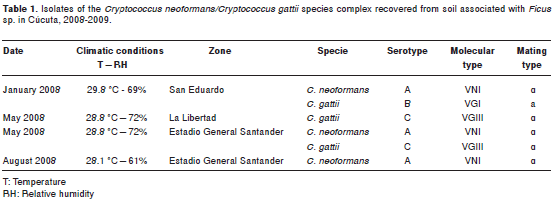
There was no association between temperature, climatic conditions and season of the year for the recovering of isolates of the C. neoformans/C. gattii species complex (Table 1). Additionally, no isolates were recovered directly from bark, detritus, leaves or fruits from the trees sampled; all isolates recovered were collected from soil.
Discussion
Although the number of isolates of the C. neoformans/C. gattii species complex recovered in this study was very small, this is the first report about the isolation of C. gattii serotype B from the environment in Cúcuta. This report confirms the presence of C. gattii in a terrestrial ecosystem, and although the isolate recovered was molecular type VGI, in contradistinction to the clinical isolates that have been recovered in Cúcuta, which are the majority molecular type VGII (12), the isolation of C. gattii serotype B helps us to clarify how the incidence of human cases of cryptococcosis reported in this Colombian city, could be probably due to the environmental exposure to this fungus, due to the fact that the recovering of C. gattii from the environment in Cúcuta had been only reported for serotype C isolates (11). Additionally, this suggests that it is necessary to do further studies in order to recover environmental isolates belonging to the same molecular pattern as the clinical isolates.
It is very important to mention that the serotype identification of the serotype B isolate recovered was done using a multiplex PCR described by Ito-Kuwa S, et al., which is a technique that helps us predict the serotype within the C. gattii isolates, and in no manner replaces the serotyping done with the specific antisera, since it does not directly detect the capsular serotype itself. However, it is worth mentioning that this multiplex PCR has been proven to be an excellent choice for C. neoformans serotypes A and D. An important fact, is that all C. neoformans var. grubii isolates recovered were molecular type VNI and most of the C. gattii isolates were molecular type VGIII (67%), which is in agreement with other studies made in Colombia, where these molecular types had been the most prevalent within these species (12,25).
In addition, the findings of this study increase the number of reports of the recovering of C. gattii from the environment, although it was not possible to associate the isolate with a specific tree species, because the isolate was recovered from soil, in contrast to previous studies of ecological reservoirs of C. gattii, in which associations between the yeast and different trees species had been found due to the recovering of the yeast from vegetable parts (14-17,27,28). However, our results show that the soil is the sample that is harboring the fungus, while the other tree material was negative, possibly because most of the samples were taken from soil (82.8%), but suggesting that the soil is one of the main reservoirs of colonization for this pathogen, and therefore, the fungus may be interacting with non-vertebrate hosts as nematodes and amoebae, as suggested by Casadevall et al (29).
Our results also reinforce that C. gattii is not associated with a particular species of tree, or a specific niche, because it was recovered from soil, which could be possible due to the contamination with avian dropping, which is considered an important substrate for the presence and maintenance of C. neoformans in nature (30) and although there are no reports for the recovering of C. gattii from avian dropping, this source could also be important, considering that there are some reports where species of Cryptococcus had been recovered from birds excreta (31) and in Cucuta there is a wide population of pigeons (Columba livia) and no sampling of these birds have been done.
Another factor that influenced the isolation of C. gattii from soil is the fact that all samples were recovered near to the soil surface, which is in agreement with other reports, where C. gattii has been recovered in the first 15 cm of the humic layer of soil, where the conditions of temperature, moisture, and nutrient requirements, are enough for the colonization of the fungus (32).
Compared with other studies where C. gattii had been found in 27.5% and 99% of the sampled trees, and the recovery was between 50 and 15 x 106 cfu/g of sample (16,33) and C. neoformans had been recovered among 7.9% and 14% of the samples, with a recovery between 50 and 1x107 cfu/g of sample (33,34), in our study, only 0.07% of the samples processed were positive for C. neoformans/C. gattii species complex. However, the recovery of only one isolates of C. gattii serotype B from all of the sampled areas, including the houses of patients with cryptococcosis, and given its prevalence in Cúcuta, could suggest that the fungus exists in few amounts in the environment in comparison with another species of bacteria or fungi which makes it difficult to be recovered, and further sampling over an extended period of time is required to find new ecological niches of C. gattii in this city. Besides, it is necessary to recover a higher number of isolates to associate the distribution of the fungus with climatic conditions, season of the year and temperature of the city, because in this study no association was found, due to the small number of isolates.
On the other hand, the isolation of C. gattii serotypes B and C with C. neoformans var. grubii from the same place, like the co-distribution of different genotypes or serotypes found in previous studies (27,28,32) could suggest a related introduction and dispersal mechanisms or synergistic colonization considering that in some cases the populations could be very complex.
In conclusion, and in spite of the low recovery rate of C. gattii serotype B, this study shows the importance of continuing doing environmental samplings in Cúcuta where C. gattii is causing cryptococcosis, due to the fact that this pathogen has shown a higher virulence after the outbreak in Vancouver Island (13) and ecological studies are relevant in the epidemiology of a pathogenic agent. In addition, the identification of new habitats of C. gattii is very important and it is possible that the presence of yeasts in soil is the source of contamination of immunocompetent individuals, because the yeast can be dispersed by the air and patients acquire the cryptococcal infection from environmental sources (8,15). For further studies, other environmental sources such as pigeon droppings or biodegraded wood could be included to increase the possibility of recovering more isolates.
Acknowledgments
To the Grupo de Microbiología of the Instituto Nacional de Salud for the collaboration in the development of this work. To Elizabeth Castañeda for the significant revision done to this manuscript.
Conflicts of interests
The authors declare that there are no conflicts of interest.
Financial support
This study was financed by the Instituto Nacional de Salud and the Departamento Administrativo de Ciencia, Tecnología e Innovación-Colciencias, grant 2104-343-19163.
Corresponding autor: Patricia Escandón, Grupo de Microbiología, Instituto Nacional de Salud, Avenida Calle 26 Nº 51-20, Bogotá, D.C., Colombia.
Phone: (571) 2200920; fax: (571) 2200934 pescandon@ins.gov.coReferencias
1. Bovers M, Hagen F, Boekhout T. Diversity of the Cryptococcus neoformans-Cryptococcus gattii species complex. Rev Iberoam Micol. 2008;25:S4-12. [ Links ]
2. Peachey PR, Gubbins PO, Martin RE. The association between cryptococcal variety and immunocompetent and immunocompromised hosts. Pharmacotherapy. 1998;18:255-64. [ Links ]
3. Speed BR, Dunt D. Clinical and hosts differences between infections with the two varieties of Cryptococcus neoformans. Clin Infect Dis. 1995;21:28-34. [ Links ]
4. Sorrell TC. Cryptococcus neoformans variety gattii. Med Mycol. 2001;39:155-68. [ Links ]
5. Byrnes EJ, Bildfell RJ, Dearing PL, Valentine BA, Heitman J. Cryptococcus gattii with bimorphic colony types in a dog in western Oregon: Additional evidence for expansion of the Vancouver Island outbreak. J Vet Diagn Invest. 2009;21:133-6. [ Links ]
6. Castellá G, Abarca ML, Cabañes FJ. Cryptococcosis and pets. Rev Iberoam Micol. 2008;25:S19-24. [ Links ]
7. Duncan C, Schwantje H, Stephen C, Campbell J, Bartlett K. Cryptococcus gattii in wildlife of Vancouver Island, British Columbia, Canada. J Wildl Dis. 2006;42:175-8. [ Links ]
8. Velagapudi R, Hsueh YP, Geunes-Boyer S, Wright JR, Heitman J. Spores as infectious propagules of Cryptococcus neoformans. Infect Immun. 2009;77:4345-55. [ Links ]
9. Lizarazo J, Linares M, de Bedout C, Restrepo A, Agudelo CI, Castañeda E, et al. Results of nine years of the clinical and epidemiological survey on cryptococcosis in Colombia, 1997-2005. Biomédica. 2007;27:94-109. [ Links ]
10. Lizarazo J, Mendoza M, Palacios D, Vallejo A, Bustamante A, Ojeda E, et al. Criptococosis ocasionada por Cryptococcus neoformans variedad gattii. Acta Med Colomb. 2000;25:171-8. [ Links ]
11. Callejas A, Ordóñez N, Rodríguez MC, Castañeda E. First isolation of Cryptococcus neoformans var. gattii, serotype C, from the environment in Colombia. Med Mycol. 1998;36:341-4. [ Links ]
12. Escandón P, Sánchez A, Martínez M, Meyer W, Castañeda E. Molecular epidemiology of clinical and environmental isolates of the Cryptococcus neoformans species complex reveals a high genetic diversity and the presence of the molecular type VGII mating type a in Colombia. FEMS Yeast Res. 2006;6:625-35. [ Links ]
13. Kidd SE, Hagen F, Tscharke RL, Huynh M, Bartlett KH, Fyfe M, et al. A rare genotype of Cryptococcus gattii caused the cryptococcosis outbreak on Vancouver Island (British Columbia, Canada). PNAS. 2004;101:17258-63. [ Links ]
14. Stephen C, Lester S, Black W, Fyfe M, Raverty S. Multispecies outbreak of cryptococcosis on southern Vancouver Island, British Columbia. Can Vet J. 2002;43:792-4. [ Links ]
15. Ellis DH, Pfeiffer TJ. Natural habitat of Cryptococcus neoformans var. gattii. J Clin Microbiol. 1990;25:1642-4. [ Links ]
16. Escandón P, Quintero E, Granados D, Huérfano S, Ruiz A, Castañeda E. Aislamiento de Cryptococcus gattii serotipo B a partir de detritos de Eucalyptus spp. en Colombia. Biomédica. 2005;25:390-7. [ Links ]
17. Sorrell T, Brownlee A, Ruma P, Malik R, Pfeiffer T, Ellis DH. Natural environmental sources of Cryptococcus neoformans var. gattii. J Clin Microbiol. 1996;34:1261-3. [ Links ]
18. Alcaldía de San José de Cúcuta. [Cited: Jun 24, 2009]. Available:http://www.alcaldiadeCúcuta.gov.co/ [ Links ]
19. Instituto de Hidrología, Meterología y Estudios Ambientales (IDEAM). Clima. [Cited: May 21, 2009]. Available: http://www2.ideam.gov.co/atlas/mclima.htm [ Links ]
20. Staib F, Seibold M, Antweiler E, Frohlich B, Weber S, Blisse A. The brown colour effect (BCE) of Cryptococcus neoformans in the diagnosis, control and epidemiology of C. neoformans infections of AIDS patients. Zentralbl Bakteriol Hyg.1987;266:167-77. [ Links ]
21. Kwon-Chung KJ, Polachek I, Bennett JE. Improved diagnostic medium for separation of Cryptococcus neoformans var. neoformans (serotype A and D) and C. neoformans var. gattii (serotype B and C). J Clin Microbiol. 1982;15:535-7. [ Links ]
22. Casali A, Goulart L, Rosa e Silva L, Ribeiro A, Amaral A, Alves S, et al. Molecular typing of clinical and environmental Cryptococcus neoformans isolates in the Brazilian state Rio Grande do Soul. FEMS Yeast Res. 2003;3:405-15. [ Links ]
23. Ito-Kuwa S, Nakamura K, Aoki S, Vidotto V. Serotype identification of Cryptococcus neoformans by multiplex PCR. Mycoses. 2007;50:277-81. [ Links ]
24. Meyer W, Castañeda A, Jackson S, Huynh M, Castañeda E, IberoAmerican Cryptococcal Study Group. Molecular typing of IberoAmerican Cryptococcus neoformans isolates. Emerg Infect Dis. 2003;9:189-95. [ Links ]
25. Meyer W, Mitchell TG. Polymerase chain reaction fingerprinting in fungi using single primers specific to minisatellites and simple repetitive DNA sequences: Strain variation in Cryptococcus neoformans. Electrophoresis. 1995;16:1648-56. [ Links ]
26. Halliday CL, Bui T, Krockenberger M, Malik R, Ellis DH, Carter DA. Presence of alpha and a mating types in environmental and clinical collections of Cryptococcus neoformans var. gattii strains from Australia. J Clin Microbiol. 1999;37:2920-6. [ Links ]
27. Girish CP, Prabu D, Mitani H, Mikami Y, Menon T. Environmental isolation of Cryptococcus neoformans and Cryptococcus gattii from living trees in Guindy National Park, Chennai, South India. Mycoses. 2010;53:262-4. [ Links ]
28. Randhawa HS, Kowshik T, Chowdhary A, Preeti Sinha K, Khan ZU, Sun S, et al. The expanding host tree species spectrum of Cryptococcus gattii and Cryptococcus neoformans and their isolations from surrounding soil in India. Med Mycol. 2008;46:823-33. [ Links ]
29. Malliaris SD, Steenbergen JN, Casadevall A. Cryptococcus neoformans var. gattii can exploit Acanthamoeba castellanii for growth. Med Mycol. 2004;42:149-58. [ Links ]
30. Lazéra MS, Salmito MA, Londero AT, Trilles L, Nishikawa MM, Wanke B. Possible primary ecological niche of Cryptococcus neoformans. Med Mycol. 2000;38:379-83. [ Links ]
31. Rosario I, Acosta B, Colom F. La paloma y otras aves como reservorio de Cryptococcus spp. Rev Iberoam Micol. 2008;25:S13-8. [ Links ]
32. Kidd SE, Chow Y, Mak S, Bach PJ, Chen H, Hingston AO, et al. Characterization of environmental sources of the human and animal pathogen Cryptococcus gattii in British Columbia, Canada, and the Pacific Northwest of the United States. Appl Environ Microbiol. 2007;73:1433-43. [ Links ]
33. Granados D, Castañeda E. Isolation and characterization of Cryptococcus neoformans varieties recovered from natural sources in Bogotá, Colombia, and study of ecological conditions in the area. Microb Ecol. 2005;49:282-90. [ Links ]
34. Quintero E, Castañeda E, Ruiz A. Distribución ambiental de Cryptococcus neoformans en el departamento de Cundinamarca â Colombia. Rev Iberoam Micol. 2005; 22:93-8. [ Links ]














