Services on Demand
Journal
Article
Indicators
-
 Cited by SciELO
Cited by SciELO -
 Access statistics
Access statistics
Related links
-
 Cited by Google
Cited by Google -
 Similars in
SciELO
Similars in
SciELO -
 Similars in Google
Similars in Google
Share
Acta Biológica Colombiana
Print version ISSN 0120-548X
Acta biol.Colomb. vol.15 no.3 Bogotá Sept./Dec. 2010
A SYSTEM FOR ACCESSING A COLLECTION OF HISTOLOGY IMAGES USING CONTENT-BASED STRATEGIES
Sistema para acceder una colección de imágenes histológicas mediante estrategias basadas en el contenido
GONZÁLEZ F.1, Ph. D.; CAICEDO JC.1, Est-Ph. D.; CRUZ-ROA A.1, Est-M.Sc.; CAMARGO J.1, Est-Ph. D.; SPINEL C.2,3, Ph. D.
1Departamento de Ingeniería de Sistemas e Industrial, Facultad de Ingeniería, Universidad Nacional de Colombia. Bogotá, D.C., Colombia. 2Laboratorio de Biofísica, Centro Internacional de Física, Universidad Nacional de Colombia. Bogotá, D.C., Colombia. 3Departamento de Biología, Facultad de Ciencias, Universidad Nacional de Colombia. Bogotá, D.C., Colombia. Correspondencia: Profesor Fabio Augusto González, Departamento de Ingeniería de Sistemas e Industrial, Universidad Nacional de Colombia. Carrera 30 # 45 – 03, Edificio 453. Bogotá, D.C., Colombia. fagonzalezo@unal.edu.co.
Presentado 15 de marzo de 2010, aceptado 4 de junio de 2010, correcciones 22 de junio de 2010.
ABSTRACT
Histology images are an important resource for research, education and medical practice. The availability of image collections with reference purposes is limited to printed formats such as books and specialized journals. When histology image sets are published in digital formats, they are composed of some tens of images that do not represent the wide diversity of biological structures that can be found in fundamental tissues. Making a complete histology image collection available to the general public having a great impact on research and education in different areas such as medicine, biology and natural sciences. This work presents the acquisition process of a histology image collection with 20,000 samples in digital format, from tissue processing to digital image capturing. The main purpose of collecting these images is to make them available as reference material to the academic comunity. In addition, this paper presents the design and architecture of a system to query and explore the image collection, using content-based image retrieval tools and text-based search on the annotations provided by experts. The system also offers novel image visualization methods to allow easy identification of interesting images among hundreds of possible pictures. The system has been developed using a service-oriented architecture and allows web-based access in http://www.informed.unal.edu.co
Key words: histological images, virtual atlas, content-based image retrieval
RESUMEN
Las imágenes histológicas son un importante recurso para la investigación, la educación y la práctica médica. La disponibilidad de imágenes individuales o colecciones de imágenes de referencia está limitada a formatos impresos como libros y revistas científicas. En aquellos casos en donde se publican conjuntos de imágenes digitales, éstos están compuestos por algunas cuantas decenas de imágenes que no representan la gran diversidad de estructuras biológicas que pueden encontrarse en los tejidos fundamentales. Contar con una completa colección de imágenes histológicas es de gran apoyo para los procesos de investigación y educación en diferentes áreas de la medicina, biología y ciencias. En este trabajo se presenta el proceso de adquisición de una colección de 20.000 imágenes histológicas en formato digital, desde la preparación y fijación de los tejidos hasta su digitalización bajo el microscopio, con el propósito de publicarlas como material de referencia para la comunidad académica en general. Además, se presenta el diseño y la arquitectura de un sistema para consultar y explorar la colección de imágenes, utilizando herramientas de búsqueda basadas en el contenido de las imágenes y en las anotaciones provistas por los expertos. El sistema también ofrece novedosos mecanismos de visualización de las imágenes, para facilitar la tarea de identificar las imágenes interesantes entre otros cientos posibles en la colección. El sistema fue desarrollado usando una arquitectura orientada a servicios y ofrece acceso a través de la Web en http://www.informed.unal.edu.co
Palabras clave: imágenes histológicas, atlas virtual, recuperación de imágenes basada en contenido.
INTRODUCTION
Biomedical images are an important asset for health research and medical practice. There are a multitude of devices for biomedical image acquisition that range from simple, e.g. a digital camera coupled with a conventional optical microscope, to complex, e.g. specialized equipment for Positron Emission Tomography (PET). These devices are routinely used in the daily medical practice and biomedical research, generating a continuous stream of images. The great majority of these images are digital and a good amount of them are permanently stored in digital image repositories. These image collections are a potential source of information and knowledge. However, the realization of this potential requires effective mechanisms to access, to explore and to visualize large image collections.
In particular, histology images are of great importance for studying the composition of cells, glands, tissues and organs, and the possible pathologies that may affect them. Research in histology heavily relies on high quality images to characterize biological structures using different stain procedures and microscopy techniques. Also, the education and training of students in medicine and natural sciences require the constant evaluation of histology images to understand the composition and behaviour of living beings, and to learn how to distinguish between normal and abnormal conditions of biological structures. In that direction, we want to build an accessible collection of histology images that can support different tasks in research and training processes (Aijón Noguera et al. 2008).
A collection of histology images has to meet some properties to be considered as reference for such taks. First, it has to have enough material to satisfy different information needs in histology, including samples from different tissue varieties, acquired from distinct organs and systems. Also, these images should be acquired using various staining techniques and microscopy modalities, that reveal complementary aspects of the underlying biological structure. Second, the collection has to be indexed and organized in such a way that accessing it and finding useful images can be a straightforward task.
The construction of a publicly-available histology image collection that meets those requirements has not been previously addressed by the research comunity. Several projects to make biology images accessible have been proposed, such as the Human Visible project (Ackerman 1999), in which the internal organs of a complete human body can be visualized and studied. However, this project is not specifically related to histology and does not have indexing methods to allow flexible image search. Some projects to build histology image retrieval systems have been addresed as well (Tang et al. 2003, Zheng et al. 2003), but they lack of a complete model for accessing images and concentrate only on one aspect of search functionalities.
The work presented in this paper is the first effort to simultaneously build a large histology image collection and to develop a fully operational system to retrieve and explore these images. The image collection has been acquied from scratch and the process included organ obtention, tissue impregnation and slicing, microscopy analysis and digital image capturing. The developed system is able to automatically index the image collection using visual features and text annotations, allowing users to query for specific images, to browse similar pictures and to explore particular image sets.
As a result, a collection of 20,000 histology images has been collected, which comprises samples from the four fundamental tissues (epithelial, connective, muscle and nervous) and three main systems (digestive, respiratory and nervous). This collection has been indexed using the proposed computerized methods that allow image storage, content-based retrieval and image collection exploration, involving state-of-the-art technologies for image indexing. The system also provides an annotation tool that allows experts to attach global annotations and to define regions of interest (ROI). Around 19,348 images were annotated with global descriptions and more than 2,164 ROIs were labeled with specific tags. The system is now available online in http://www.informed.unal.edu.co, which provides simplified search tools allowing full access to the complete image collection, including available descriptions and ROIs.
MATERIALS AND METHODS
IMAGE ACQUISITION
Organ Obtention. Twelve ICR mice (young adults, 30 g), from Bioterio Experimental de la Facultad de Veterinaria y Zootecnia, Universidad Nacional de Colombia, sede Bogotá (UN), were used. Animals were handled according to the requirements of the National Research Council (AVMA, 2001), fulfilling ethical requirements of Colombian legislation, Res. 8430/1993, Title IV and Law 84/1996 which refers to biomedical research with animals. Each mouse is anesthetized with an overdose i.p. of pentobarbital 250 µg/g, it is attached to a dissecting table, abdominal skin is cleaned with alcohol 70%. We performed a incision from the abdomen to the level of the clavicles and lateral incisures of skin are done at the diaphragm. The thoracic cavity is opened and the perfusion system is placed in the left ventricle of heart. Before pumping (~260 mm Hg) the fixative (4% paraformaldehyde in phosphate buffered saline (PBS) 0.1 M, pH 7.4), a cut is done in the vena cava where it connects with the atrium, in order to use the animal circulation in such a way that the fixative is irrigated through the body assuring a correct preservation of all the tissues. The micro-dissection of the different organs of interest is performed under the stereoscope leaving small tissue fragments in the fixative. For the nervous system and tissue, the animal is turned around and the micro-dissection is performed on the dorsal side of the animal extracting ganglia, spinal cord and brain (Luna 1968).
Tissue Impregnation. The fragments are left between 6 and 12 hours in the same fixative. Subsequently, they are dehydrated in increasing concentrations of ethyl alcohol, each of 30 min 75%, 85%, 90% and 5X absolute alcohol. The process continues with xylene for 1 h, and it continues in xylene/paraffin 1/1 for one more hour. Finally fragments are passed through paraffin for 1h at 60 °C. For blocks mounting, a cold plate is used with paraffin at 60 °C, each block is marked with a code for subsequent identification. These blocks are allowed to cool and removed from the metal mold, and thus gives the paraffin block of tissue or organ for histological sections. These are saved in the block storage of Laboratorio de Biofísica for further processing (Luna 1968, Bernal 2006).
Histological Slices. 5 mm-thick cuts are made with a microtome. Then, they are stained with hematoxylin and eosin (purple and pink respectively) for the controls and to verify optimal histological preparation. To better determine the different tissues, trichrome and pentachrome (five colors ranging from light blue and dark orange, pink and shades of red depending on cell type) were used.
For special tissues, such as keratinized epithelial stratified squamous was obtained from human skin and urinary epithelial tissue of cat, which were donated by Laboratorio de Patología, Universidad Nacional de Colombia.
Digital Image Acquisition. The acquisition of 20,000 digital images (or photomicrographs) was performed in a photomicroscope (Zeiss) using a video camera (Sony).The images were stored in color format JPG at 640x480 resolution.
All images were labeled, using the annotation tool to be described in Section 3.4, with a general text that describes the tissue or/and organ which represent them. On the other hand, we made a marking of regions in the images delineating a polygon of presence in the images of different tissues: epithelial, connective, muscular and nervous (Fawcett and Bloom 1997). Also some of these images have descriptions of the different tissues that characterize the organs of different systems (digestive, respiratory and nervous).
SYSTEM ARCHITECTURE AND FUNCTIONALITY
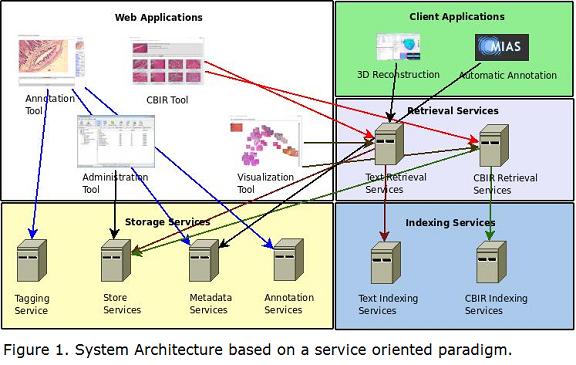
The architecture of the system is based on a service-oriented paradigm, where each component exposes its functionality as a web service. Figure 1 shows an overview of the architecture. The storage component manages image collection operations including image upload, image download, image meta-data, collection information, user and control access. The annotation component offers a web interface to the expert for annotation purposes. The indexing component manages the index structure, which brings typical information retrieval functionalities like stop-words removal, stemming, similarity measures, recommendations, among others. The image retrieval component manages the content-based image retrieval tasks (visual and textual). The user interface component exposes the system functionality to the user using a web interface. Finally, the visualization component offers a 2D visualization that allows the user to interactively explore the image collection.
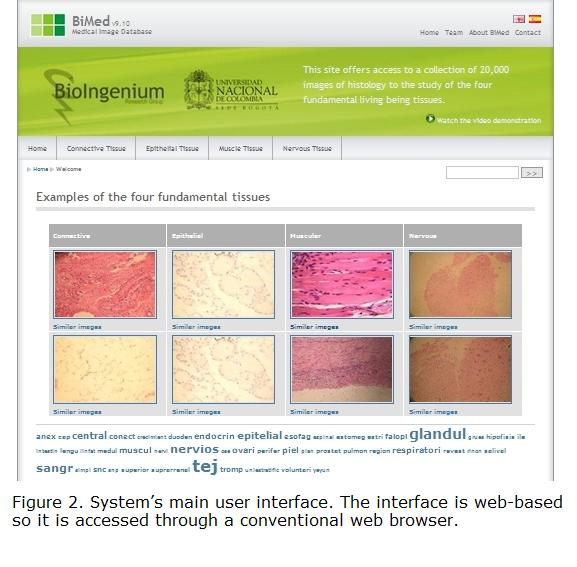
User Interface. The system provides content-based image retrieval functionalities that allow the user to find images of interest querying the system with textual keywords or visual examples. The look-and-feel of the end-user interface is shown in Figure 2. The retrieval interface of the application has the following interacting tools: (1) a main menu with the most common keywords in the collection; (2) a text box where user enters textual keywords; (3) check boxes for selecting the low-level features to be used in the visual retrieval process; (4) a similar-images link for retrieving images by visual content; (5) a cloud of concepts automatically generated from textual annotations; and (6) a 2D visualization of the results obtained in visual and textual searches (Gonzalez and Romero 2009).
Image Representation and Storage. The image collection is composed of 20,000 images in JPG format. We generated a set of thumbnails, one per image, which are visualized when the user query the system. JPG files are stored in a web server, which is part of the storage component. An image is referenced by a unique identifier and is accessed through a web service exposed in the storage component.
Images also have text annotations that describe their content. This text is captured when the user annotate the images using the annotation tool. This unstructured text is processed to create a structured version in XML format that is used by the indexing component.
Image Indexing and Retrieval. Due to the amount of images stored in the system, it was necessary to implement an efficient mechanism to access the image repository. In order to query the system, the user can use a set of keywords in the case of text-based search, and select an image by example in the case of visual-based search. Figure 3 and Figure 4 show the results for textual search and visual similarity search respectively.
Visual Indexing. The goal of a content-based image retrieval system is to offer the possibility of finding similar images based on their visual content. Therefore, it is important to define a representation of the image content. To build this representation, we used image processing methods to objectively quantify image properties. The similarity between two images can be obtained by comparing their corresponding visual feature representations.
Three main visual features were extracted, each one related to some visual characteristic. All features are represented as histograms that indicate the frequency of visual patterns within the image. The three features are the following:
-Color histogram. A histogram with the distribution of colors in the image is constructed according to Siggelkow (2002). Color is an important feature in this histology image collection, since different staining procedures were used to highlight biological structures, including hematoxilin-eosin and inmunohistochemistry.
-Local binary patterns. This is a texture measure, which indicates regular patterns along the image content (Berman and Shapiro 1999). Textures define a meaningful characteristic of histology images, since tissues are defined by regular cell structures.
-Sobel histogram. This histogram is used to characterize visual changes in the image, commonly known as edges (Sobel 1970). Several biological structures can be recognized by the contrast they produced with respect to tissues or other structures. Then, we considered this feature as an important one to represent histology images.
Each image is represented by these three histograms, comprising three different visual image indexes, that is, we can find images with certain visual properties due to the computation of these features. To efficiently retrieve images using that visual information, an appropriate similarity function has to be defined. The similarity function used in this work is based on histogram intersection simmilarity, which evaluates the common area between two histograms (Barla et al. 2003). The maximum value is reached when both histograms have exactly the same shape and the minimum value when they are completely different.
A user can combine these three features by picking two or three features in the user interface, to indicate to the system which of them are used to compute image similarity. In other words, a user might be interested in finding images with similar colors, so only this feature would be taken into account. But, if a user wants to retrieve similar images according to a combination of colors and textures, it can be defined just by selecting them before querying the system. This provides a flexible search process, in terms of the visual features that catch the user’s attention.
Text Indexing. In the case of text-based search it is possible to find images by keywords. Each image has associated annotations that are provided by the expert. All annotations are used to construct a unique document, one per image, which is then represented using the vector space model (VSM) (Baeza et al. 1999). In VSM each document is represented by using an algebraic model in which the document is modeled as a vector. Each component of this vector is weighted according to the number of occurrences of the term in the document and normalized according to the number of occurrences in the collection (Salton et al. 1975). When the user expresses a query by means of keywords, this query is also represented by the system as a vector. In order to find similar documents, the system compares this query to the documents stored in the data base. Due to compare the query with all documents sequentially is not efficient, the system uses a computational structure, that is known as index, which allows to the system to solve the query in an efficient way. This text indexing functionality is provided by the indexing component, which was built using Solr. Solr is an open source implementation of the described indexing strategy and offers basic functionalities to support indexing and text retrieval. Solr is available at http://lucene.apache.org/solr/.
IMAGE ANNOTATION AND LABELING
The annotation tool allows the expert to annotate an image by both, drawing the region of interest with its respective text description (or with a concept previously entered in other image), and entering a global description of the image. This text is used to support textual search in the indexing component. The annotation tool offers visual controls to mark regions of interest with its respective text. Figure 5 shows a screenshot of the tool. The user marks with the mouse the region of interest. The coordinates of the annotated region are stored along with the associated text.
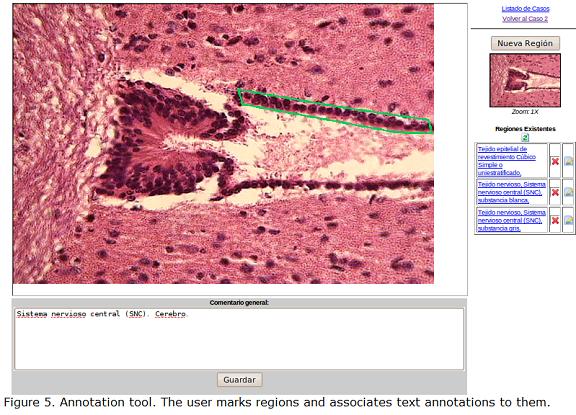
IMAGE COLLECTION VISUALIZATION
The results of both textual and visual search are displayed by default in a grid layout. The user has the alternative of visualizing these results in a more graphical and intuitive way. In this visualization, the images are organized in the screen by visual similarity. That is, images that share visual similarity are projected into the same neighborhood.
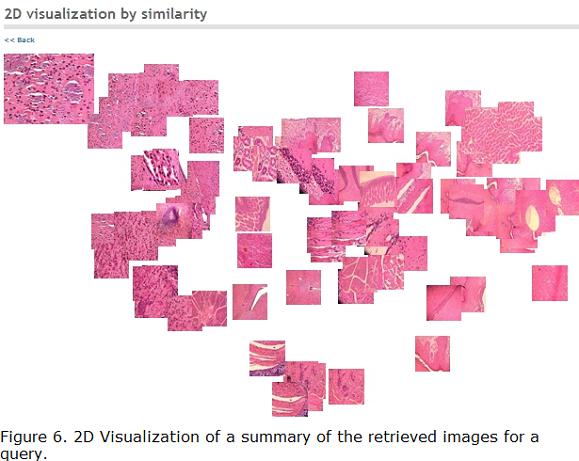
We generate a 2D visualization of the image collection according to the similarity function defined in this Section. We used a dimensionality reduction method, which finds a low-dimensional representation to project each image into 2D coordinates system. The method used in this system was kernel principal component analysis (KPCA) (Schölkopf et al. 1998). Images are represented based on their projections to the two principal components, which produces a 2-dimensional representation that is expected to preserve, to some extent, the similarity relationship, i.e., two very similar images are expected to be projected to the same region of the 2-dimensional space. See (Camargo and González. 2009) for more details. Figure 6 shows an example of this visualization.

RESULTS AND DISCUSSION
HISTOLOGY IMAGE COLLECTION
The digital image collection was obtained from a total of 20,000 histology images for study of the four fundamental tissues. Among this dataset a total of 19,348 images were annotated with a global description of these tissues, from which 762 images are of connective tissue, 1280 of epithelial tissue, 850 of muscle tissue and 1626 of nervous tissue. The acquired images show these tissues in different stains (hematoxylin-eosin, trichrome, PAS, immunocytochemistry, etc.) and at different magnifications and cuts of slides. Some details of the result of digitization and the collection of histological slides are presented in Table 1.

SYSTEM ARCHITECTURE AND PERFORMANCE
The system has been developed using the Java programming language under a serviceoriented architecture. Five main modules, each with a specific responsibility, are interconnected using web services. These modules are: Visual indexing, text indexing, image storage, main user interface, and image collection visualization. This organization allows flexible development and rapid scalability, since each functionality is encapsulated in a separate module. Having implemented the system, we wanted to evaluate two different aspects related to performance. The first aspect is search precision, that is, when a user requests for some images, we want to know whether the system is delivering the correct images to properly satisfy user’s information needs. The second apect is efficiency in the use of computational resources, i.e. when the system has to find images, how long does the user have to wait to get the requested images. We explored these two issues using visual information only, since the underlying methods may require extra computing power. We assume text operations being both, precise and efficient, since the implemented methods are standard information retrieval operations that have been widely explored by the research community, and are now well-established for different applications.
For visual search precision, we evaluated a set of queries assuming that a user wants to retrieve similar images, expecting that the resulting images are the same fundamental tissue as the query. So for instance, if the user wants similar images to a particular nervous sample, we assume all the results should be nervous samples as well, order by visual similarity. Since the visual search algorithm does not have semantic information to filter out non-nervous images, the proposed evaluation aims to count how many images in the results delivered by the system share the same fundamental tissue as the query. Experimental results showed that using color information, the system gets up to 57% of precision along the first 10 images in the screen. That means that almost 6 out of 10 images are semantically related to the query, in terms of the fundamental tissue. These experiments were conducted by biologists that marked the results as relevant or not according to the particular test.
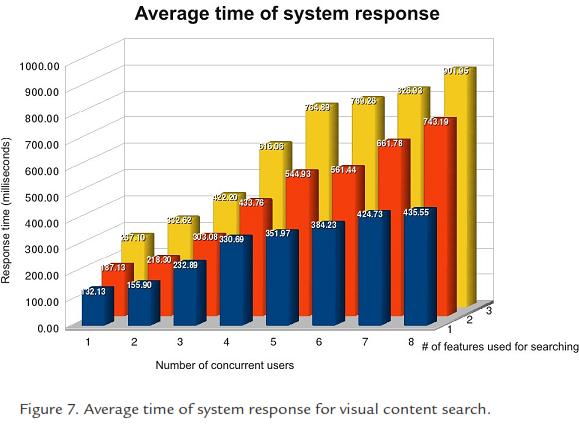
To evaluate system response times, we simulated concurrent users querying the system for similar images. We simulated up to 8 concurrent users, that is, 8 search requests that arrive exactly at the same time, so the computer system has to process all of them simultaneously, and deliver the appropriate results to each of the users as fast as possible. The simulation also included an additional variable related to the number of features the user wants to combine in the search process to rank similar images. Real users might want to search using color, texture or edges only, the three of them simultaneously, or any possible combination.
Figure 7 shows a plot that shows how the delay grows as long as more users and more features are introduced in the search process. The time is consistently longer as the number of concurrent users grows as well as the number of features each user wants to include. For instance, the situation in which 8 users are demanding similar images using 3 features each, forces the system to delay each response up to 901 milliseconds in average. A system response is considered to be interactive if it delivers results in a time shorter than 1 second. Then, this experimental evaluation shows that the implemented system is able to provide an interactive response even under the worst evaluated conditions.
CONCLUSIONS
The main conclusion of this work is that there is a great research potential in the intersection of biomedicine and image retrieval technologies. Images are an important asset for biomedical practice and research, however, this potential is usually underexploited because of the lack of efficient mechanisms to access the huge amount of visual and non-visual information present in image collections. The system presented in this paper is a success case that will serve as basis for future developments in this exciting area.
ACKNOWLEDGMENTS
This work was partially funded by the project SISTEMA DISTRIBUIDO DE ANOTACIÓN AUTOMÁTICA Y RECUPERACIÓN SEMÁNTICA DE IMÁGENES DE HISTOLOGÍA (110148725779) of Ministerio de Educación Nacional de Colombia by Convocatoria COLCIENCIAS 487 de 2009: Apoyo a proyectos de investigación, desarrollo tecnológico e innovación que hagan uso de la infraestructura y servicios de la Red Nacional Académica de Tecnología Avanzada (RENATA). We also want to thank COLCIENCIAS (110140520161), to Research Headquarters Bogotá Unit of Colombia National University and to program ECOS-Nord/Colciencias/ICFES/ICETEX.
BIBLIOGRAPHY
AIJÓN NOGUERA A, ALONSO PEÑA J, RAMÓN J, ARÉVALO ARÉVALO R, GARROSA GARCÍA M, GAYOSO RODRÍGUEZ MJ. Atlas virtual de preparaciones citológicas e histológicas. Departamento de Biología Celular y Patología, Universidad de Salamanca. Nov 6/2008. Localización URL: http://biblioteca.universia.net/html_bura/ficha/params/title/atlas-virtual-preparaciones-citologicas-histologicas/id/38120421.html.
[ Links ]ACKERMAN, M. The visible human project: a resource for education. Acad Med; 1999;74(6):667-670.
[ Links ]AVMA. Report of the AVMA panel on euthanasia. JAVMA. 2001;218:669-692.
[ Links ]BAEZA R, RIBEIRO B. Modern Information Retrieval. New York: ACM Press, A Division of the Association for Computing Machinery, Inc.; 1999.
[ Links ]BARLA A, ODONE F, VERRI A. Histogram intersection kernel for image classification. Proc Int Conf Image Proc. 2003(3);513-516.
[ Links ]BERMAN AP, SHAPIRO LG. A Flexible Image Database System for Content-Based Retrieval. Comput Vis Image Underst. 1999;75:175-195.
[ Links ]BERNAL E, SPINEL C. Estudio inmunhistoquímico de dos co-transportadores de la familia SLC5 em el sistema digestivo. acta Biol Colomb. 2006;11(2):102.
[ Links ]FAWCETT DW, BLOOM W. A textboock of histology. 9th ed. Philadelpphia: W.B. Saunders Co.; 1997.
[ Links ]LUNA LG. Manual of histological staining methods of armed forces institute of pathology. New York: McGraw-Hill Book Co.;1968.
[ Links ]CAMARGO J, GONZÁLEZ F. A multi-class kernel alignment method for image collection summarization. Lect Notes Comput Sci. 2009;5856:545-552.
[ Links ]GONZALEZ F, ROMERO E, editors. Biomedical image analysis and machine learning technologies: Applications and techniques. Hershey: Idea Group Inc.; 2009.
[ Links ]SALTON G, WONG A, YANG CS. A vector space model for automatic indexing. Commun ACM. 1975:18(11);613-620.
[ Links ]SCHÖLKOPF B, SMOLA A, MÜLLER KR. Nonlinear component analysis as a kernel eigenvalue problem. Neural Comput. 1998;10:1299-1319.
[ Links ]SOBEL I. Camera models and perception (Ph. D. thesis). California: Artificial Intelligence Lab, Stanford University Stanford; 1970.
[ Links ]SIGGELKOW S. Feature histograms for content-based image retrieval (Ph. D. thesis). Lüneburg: Fakultät für Angewandte Wissenschaften, Albert-Ludwigs-Universität Freiburg; 2002.
[ Links ]TANG H, HANKA R, IP H. Histological image retrieval based on semantic content analysis. IEEE Trans Inf Technol Biomed. 2003;7(1):26-36.
[ Links ]ZHENG L, WETZEL A, GILBERTSON J, BECICH M. Design and analysis of a content-based pathology image retrieval system. IEEE Trans Inf Technol Biomed 2003;7(4):249-255.
[ Links ]













