Serviços Personalizados
Journal
Artigo
Indicadores
-
 Citado por SciELO
Citado por SciELO -
 Acessos
Acessos
Links relacionados
-
 Citado por Google
Citado por Google -
 Similares em
SciELO
Similares em
SciELO -
 Similares em Google
Similares em Google
Compartilhar
Agronomía Colombiana
versão impressa ISSN 0120-9965
Agron. colomb. v.29 n.3 Bogotá set./dez. 2011
PLANT BREEDING, GENETIC RESOURCES & MOLECULAR BIOLOGY
Isolation of high-quality DNA in 16 aromatic and medicinal Colombian species using silica-based extraction columns
Aislamiento de DNA de alta calidad en 16 especies aromáticas y medicinales Colombianas utilizando columnas de extracción de sílica
Nelson Enrique Vega-Vela1and María Isabel Chacón Sánchez1,2
1Department of Agronomy, Faculty of Agronomy, Universidad Nacional de Colombia. Bogota (Colombia).2Corresponding author. michacons@unal.edu.co Received for publication: 11 November, 2010. Accepted for publication: 2 November, 2011.
ABSTRACT
Aromatic and medicinal plant species are a valuable resource for research and development of pharmaceutical, cosmetic, crop protection and nutritional agents, due to the high amount of bioactive phytochemicals that they contain. However, these compounds are a major obstacle in the isolation of high-quality DNA suitable for genetic analyses. In this paper, we report a protocol that optimizes the use of the cationic detergent CTAB and the reductant β-mercaptoethanol in cell lysis. The elimination of plant secondary metabolites such as polysaccharides and polyphenols, that typically co-isolate with DNA, was achieved using the chemical denaturing properties of the guanidinium cation, which together with the adsorbent chemical specificity of the silica, resulted in the purification of high-quality DNA suitable for digestion with restriction enzymes and optimal for PCR amplification of AFLP-type molecular markers. This protocol was evaluated on 16 Colombian aromatic and medicinal plant species promising for their essential oils. The results allow suggesting that this procedure might be appropriate for other species, tissues and sample types recalcitrant to DNA extraction.
Key words: chaotropic agent, guanidinium cation, silica-based extraction columns, nucleic acid extraction kit, guanidinium salts, molecular markers.
RESUMEN
Las especies vegetales aromáticas y medicinales son un recurso valioso para la investigación y el desarrollo de agentes farmacéuticos, cosméticos, agrícolas y de alimentos, debido a la alta cantidad de fitoquímicos bioactivos que contienen. Sin embargo, estos compuestos son un obstáculo en el aislamiento de ADN de alta calidad adecuado para análisis genéticos. En el presente artículo reportamos un protocolo que optimiza el uso del detergente catiónico CTAB y el agente reductor ß-mercaptoetanol en la lisis celular. La eliminación de metabolitos secundarios como polisacáridos y polifenoles que típicamente co-aíslan con el ADN, se logró utilizando las propiedades químicas desnaturalizantes del catión guanidinio que junto con la especificidad química adsorbente de la sílica, resultó en la purificación de ADN de alta calidad adecuado para la digestión por enzimas de restricción y óptimo para la amplificación por PCR de marcadores moleculares tipo AFLPs. Este protocolo se evaluó en 16 especies vegetales aromáticas y medicinales colombianas promisorias por sus aceites esenciales. Los resultados obtenidos permiten sugerir que este procedimiento podría ser apropiado para su aplicación en otras especies, tejidos y tipos de muestras recalcitrantes a la extracción de ADN.
Palabras clave: agente caotrópico, catión guanidinio, columnas de extracción de sílica, kit de extracción de ácidos nucleicos; sales de guanidinio, marcadores moleculares.
Introduction
Worldwide, aromatic and medicinal plant species are used as sources of new molecules for the synthesis of chemopreventive and anti-cancer drugs (Balunas and Kinghorn, 2005), medicinal, cosmetic, antibacterial, antiparasitic (Bakkali et al., 2008), antimicrobial, antioxidant and anti-inflammatory agents (Cowan, 1999; Svoboda and Hampson, 1999). Their importance lies in that they contain many bioactive phytochemicals, plant secondary metabolites and essential oils (Balunas and Kinghorn, 2005; Cowan, 1999), significant for the pharmaceutical, crop protection, cosmetic and food industries (Bakkali et al., 2008), and certain research areas such as pharmacognosy (Balunas and Kinghorn, 2005), biotechnology, plant breeding, genetics, molecular and evolutionary biology (Canter et al., 2005; Gómez-Galera et al., 2007; Kumar and Gupta, 2008).
In Colombia, there is an increasing interest in studying the composition and biochemical properties of essential oils from different aromatic and medicinal native plants, and their potential uses (Meneses et al., 2009; Nerio et al., 2009). The research in this kind of promising species must be accompanied by: i) the development of cultivable lines in conventional plant breeding programs and/or assisted by molecular markers from wild materials to ensure the proper agronomic exploitation and the conservation of genetic resources (Gómez-Galera et al., 2007; Kumar and Gupta, 2008), ii) the application of molecular and genomic tools appropriate for handling the modulation in the production of these oils and plant secondary metabolites (Gómez-Galera et al., 2007), and iii) the development of quality control testing to prevent piracy or adulteration of commercial products (Breton et al., 2004; Marieschi et al., 2009).
The use of molecular markers such as AFLPs (Vos et al., 1995), to assist the selection process in plant breeding (Bernardo, 2008; Collard et al., 2005) and quality control procedures (Breton et al., 2004; Marieschi et al., 2009), requires the development of protocols for isolation of high-quality DNA that allows enzymatic digestion with restriction endonucleases and amplification of regions or fragments with high reproducibility (Mace et al., 2003; Matasyoh et al., 2008; Sarwat et al., 2006). However, the isolation of DNA in aromatic and medicinal plant species is a difficult, tedious and time-consuming procedure (Khanuja et al., 1999; Michiels et al., 2003; Padmalatha and Prasad, 2006; Pirttilä et al., 2001). Basically, the problem is the formation of complexes between DNA and plant secondary metabolites such as polyphenols and polysaccharides. Among these metabolites are tannins, flavonoids, quinones, terpenes and alkaloids, that make not only attractive this group of plants but also the procedure of isolation difficult, because they degrade the extracted DNA and directly or indirectly inhibit the action of enzymes used in downstream molecular applications such as PCR amplification (Ivanova et al., 2008; Khanuja et al., 1999; Michiels et al., 2003; Padmalatha and Prasad, 2006; Pirttilä et al., 2001; Porebski et al., 1997).
The commonly reported methods for the extraction of DNA in medicinal and aromatic plants are modifications of classical procedures such as the CTAB of Doyle & Doyle and SDS of Dellaporta (Dellaporta et al., 1983; Doyle and Doyle, 1990), which must be adjusted for each sample type, tissue and species. Variable results are obtained when these procedures are applied in other species, even those closely related (Marva et al., 2007; Pirttilä et al., 2001; Sangwan et al., 2000; Sarwat et al., 2006). Because of this, the aim of this work was to develop a high quality DNA extraction protocol for recalcitrant or difficult samples, that it is chemically simple, fast, reproducible and optimal for aromatic and medicinal plant species, based on the use of chaotropic agents and membrane-silica-based extraction columns.
The use of chaotropic agents for protein denaturation dates back to the work of Svedberg in 1937, and Greenstein in 1938 (Greenstein, 1938; Greenstein, 1939a; Greenstein, 1939b; Greenstein and Edsall, 1940; Svedberg, 1937). Among the chaotropes more commonly used in chemistry and molecular biology for the structural study of proteins and for nucleic acid isolation are the guanidinium cation (Gdm+) -salts (Bowtell, 1987; Castellino and Barker, 1968; Cox, 1968; Greenstein and Edsall, 1940; Mason et al., 2003). These salts, made up of Gdm+ or Gdm+-substituted and an anion such as Cl-I-,Br- or thiocyanate (SCN-), constitute the strongest denaturing agents known in nature, and its denaturation strength depends on the accompanying anion, being Cl? the weakest and the SCN- the most powerful reported (Castellino and Barker, 1968; Greenstein, 1939b; Greenstein and Edsall, 1940; Mason et al., 2003). Thus, the denaturation or dissociation and solvent properties of these salts on proteins are in function of the type of cation, the accompanying anion and the concentration in solution (Castellino and Barker, 1968; Greenstein, 1939a; Greenstein, 1939b). On the other hand, silica is known for its ability to adsorb specifically nucleic acids (Alexander et al., 2007; Boom et al., 1990; Rogstad, 2003; Sarwat et al., 2006). Boom et al. (1990) reported for the first time the use of Gdm+ and silica particles jointly in samples of human serum and urine for the isolation of DNA and RNA. This work demonstrated the potentiality of the method, simple chemistry and a basic foundation: in presence of any chaotropic agent and solvent as Gdm+-salts, nucleic acids are specifically adsorbed by silica particles (Boom et al., 1990; Boom et al., 1999).
The procedure reported in this paper is reproducible, simple, fast, with minimal requirements of specialized equipment and chemistry of easy handling, which proved to be suitable for obtaining high-quality DNA in aromatic and medicinal plant species. This protocol allows recovering on average 4.35 µg of DNA per 150 mg of tissue, with spectral quality of 1.85 for the ratio A260/A280 and 2.08 for the ratio A260/A230. The extracted DNA was appropriate for digestion with restriction enzymes, PCR amplification with primers of importance in population genetic studies and amplification of AFLP-type molecular markers.
Materials and methods
Plant material
Young leaves were harvested from 16 medicinal and aromatic Colombian species of importance for its content of essential oils (represented by 20 accessions of interest, see Tab.2 below), growing under greenhouse conditions at the Agronomy Faculty at the Universidad Nacional de Colombia, Bogotá. The leaves were placed in an aluminum bag and kept in ice to reduce tissue oxidation until processing.
Buffers and reagents
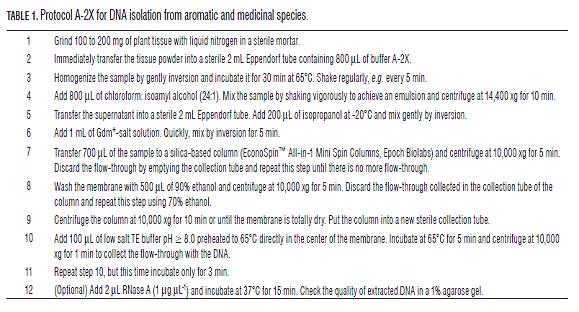
The protocol of DNA extraction for aromatic species called A-2X is described in detail in Tab.1. The buffers and reagents used in the procedure are as follows: i) Buffer A-2X: 2% CTAB w/v, 1.5 M NaCl, 20 mM EDTA, 100 mM Tris-HCl pH 8.0, and 1% β-mercaptoethanol (add just before use), ii) Chloroform : isoamyl alcohol (24:1, v/v), iii) Isopropanol at -20°C, iv) Gdm+-salt solution: a) Guanidinium Hydrochloride (GdmHCl) Solution: 2 M or 4 M GdmHCl dissolved in sterile distilled water or b) Guanidinium Thiocyanate (GdmSCN) Solution: 2 M or 4 M GdmSCN, v) Ethanol at 90 and 70%, vi) Low salt TE buffer pH = 8.0, and vii) Stock RNase A in TE buffer at a concentration of 1 µg µL-1.
Chemistry of the protocol A-2X
The A-2X extraction buffer contains the reagents CTAB, EDTA, NaCl and β-mercaptoethanol. The CTAB is a cationic surfactant used as a detergent for both selective precipitation of nucleic acids and reduction of contamination by polysaccharides and denatured proteins (Cheng et al., 1997; Kriman et al., 2006; Michiels et al., 2003; Pirttilä et al., 2001; Sarwat et al., 2006). The EDTA is used as a chelate to inhibit DNases (Matasyoh et al., 2008). The NaCl in high concentrations prevents co-precipitation of DNA together with polysaccharides (Jobes et al., 1995; Kriman et al., 2006; Porebski et al., 1997). The β-mercaptoethanol is a reducing agent used to break di-sulfide bridges, inhibit different types of enzymes by destabilization and prevent oxidation of polyphenols (Cheng et al., 1997). The buffer typically has pH between 7.0 and 8.0 for reducing contaminant RNA (Chomczynski and Sacchi, 1987).
The DNA purification is achieved using Gdm+-salts and silica-based extraction columns. The salts denature and break any inter- and intra-molecular interaction between DNA and proteins. We suggest the use of GdmHCl and GdmSCN, two of the most commonly used salts and with different denaturation powers, the first being the weakest and the other about 2.5 times more powerful (Castellino and Barker, 1968). The silica in the extraction columns efficiently adsorbs high molecular weight DNA after treatment with Gdm+-salts, which has been used successfully (Alexander et al., 2007; Ding et al., 2008; Ivanova et al., 2008; Sassa, 2007).
Quantification of extracted DNA
Two microliters of each extracted sample were used in order to determine its concentration, quantity and purity in a NanoDrop™ 2000 spectrophotometer (Thermo Fisher Scientific Inc.). Additionally, five microliters of each sample were loaded in a 1% agarose gel to determine the integrity of DNA. The staining of the products were carried out with EZVision™ Three (Amresco®) and each gel was run at 90 V for 30 min. The gels were documented using a gel documentation system Gel Doc XR (Bio-Rad Laboratories Inc.).
Enzymatic digestion with MseI
The restriction analysis was performed in nine of the 20 accessions under study. For this, 200 ng of extracted DNA were incubated for each of the samples with 1 U of MseI enzyme (Invitrogen™) for 2.5 h, following the manufacturer's instructions. The samples were run at 90 V for 50 min and visualized in a 1% agarose gel.
PCR using primers of importance in genetic population studies
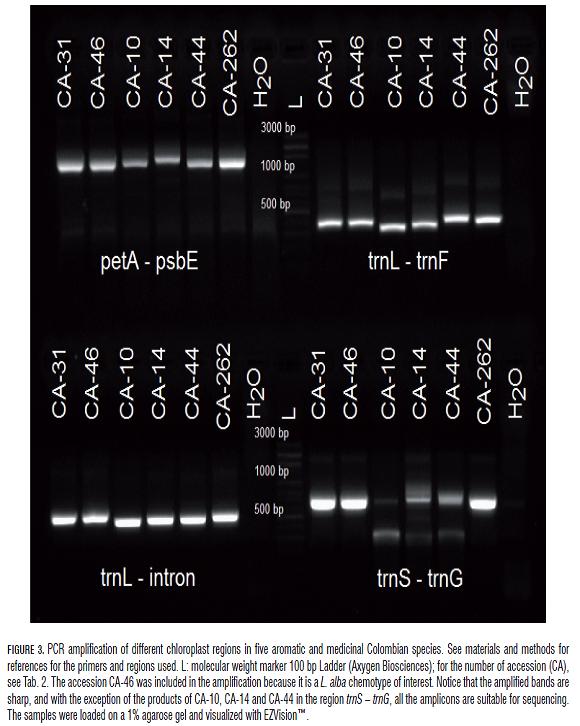
Four pairs of universal primers were used to amplify different non-coding chloroplast regions, trnL (UAA) – trnF (GAA), trnL (UAA) intron (Taberlet et al., 1991), petA – psbE (Fofana et al., 1997), and trnS (GCU) – trnG (UCC) (Hamilton, 1999), on six of the 20 accessions under study. Each PCR reaction contained 1 U of recombinant Taq DNA polymerase (Fermentas Inc.), 1X PCR Buffer with KCl, 4 mM MgCl2, 200 µM of each dNTP, 0.2 µM of each primer, for a final reaction volume of 25 µL. The reaction was performed on a C1000 Thermal Cycler (Bio-Rad Laboratories Inc.), using the following amplification protocol: 1 cycle at 94°C for 4 min, 35 cycles at 94°C for 30 s, 53°C for 30 s and 72°C for 60 s, and a final extension step at 72°C for 5 min. The products were run at 90 V for 45 min and analyzed by 1% agarose gel electrophoresis.
Amplification of AFLP-type molecular markers The kit AFLP® Analysis System I (Invitrogen™) was used to amplify three selective primer combinations (E-ACC – M-CAA, E-ACC – M-CAC, and E-ACC – M-CAG), on the same six accessions used previously to amplify noncoding regions of the chloroplast DNA. Two genotypes per accession were taken in order to reveal the presence of polymorphisms. The AFLP amplification was carried out following the manufacturer's instructions. The visualization of the products was performed as recommended by the manufacturer, although the staining of 6% PAGE was performed in silver nitrate. The gel was documented with a digital camera Olympus® SP-350.
Results
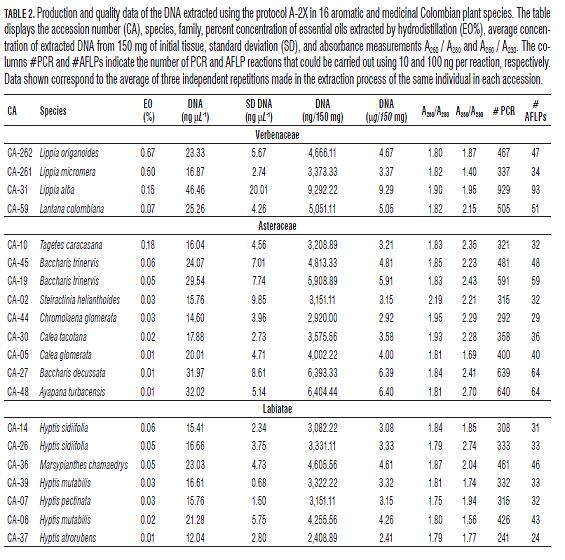
The concentration, quantity, quality and usefulness of the DNA extracted with the protocol A-2X were evaluated. The whole extraction process, from the harvesting of tissue until DNA quantification in NanoDrop™ 2000 and visualization in agarose gel on the 16 species (represented by 20 accessions of interest), was repeated three times independently for statistical support (Tab.2). Similar results in quality and quantity were obtained in all the repetitions performed. An average of 4.35 µg of DNA with an average concentration of 21.73 ng µL-1 was extracted for all species evaluated using 150 mg of fresh tissue, being L. alba the species with the highest amount of DNA (9.29 µg / 150 mg of tissue at a concentration of 46.46 ng µL-1, SD = 20.01 ng µL-1), and H. atrorubens the species with the lowest amount (2.41 µg / 150 mg of tissue at a concentration of 12.04 ng µL-1, SD = 2.8 ng µL-1). The average absorbance measurements to qualify the purity of the extracted samples were: A260/A280 = 1.85 and A260 / A230 = 2.08. For each accession the number of PCR and AFLP reactions that could be performed using 10 ng and 100 ng, respectively, were calculated from the total amount of extracted DNA per accession. On average, 434 PCR reactions and 43 AFLP reactions could be carried out from 150 mg of tissue sample using the extraction protocol A-2X.
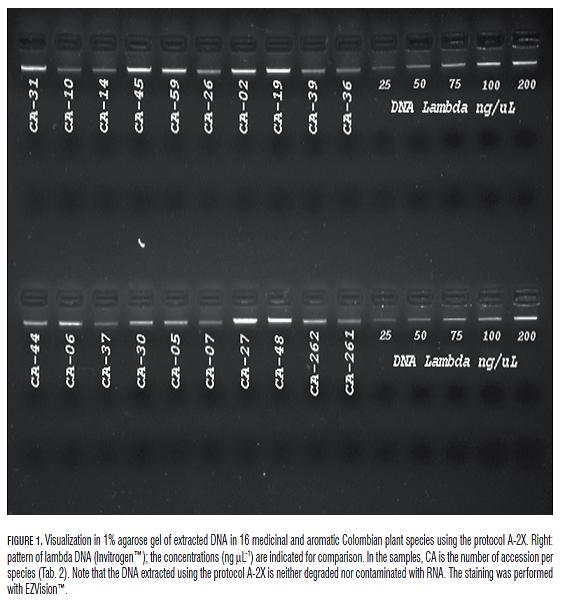
The quality and the molecular weight of the extracted DNA can be seen in Fig.1. The DNA is clean in all evaluated species and does not present degradation or residual RNA. Furthermore, if we consider that the lambda DNA sequence is 48.5Kb, the molecular weight of the DNA extracted by the protocol A-2X would be close to the molecular weight of lambda DNA.
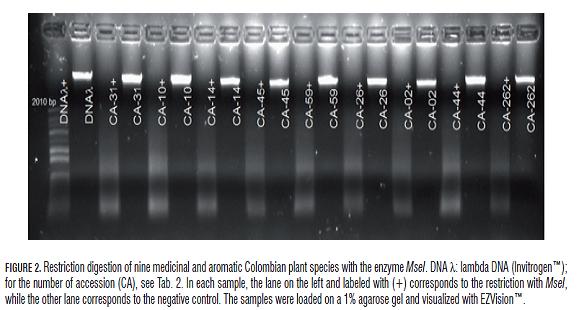
On the other hand, the full restriction of DNA with the frequent-cutter enzyme MseI resulted in a high production of fragments lower than 2 Kb (Fig.2). Different noncoding regions of the chloroplast DNA were amplified (Fig.3), as well as three combinations of selective primers using the kit AFLP® Analysis System I (Invitrogen™) (Fig.4).
Discussion
The isolation of high quality DNA for genetic analyses in aromatic and medicinal plant species is a difficult and time-consuming procedure (Ivanova et al., 2008; Khanuja et al., 1999; Matasyoh et al., 2008; Michiels et al., 2003; Pirttilä et al., 2001), which is strengthened by the use of conventional protocols that are not easily adaptable to other systems (Marva et al., 2007; Pirttilä et al., 2001; Sangwan et al., 2000; Sarwat et al., 2006). For this reason, we used the potentiality of the chemistry of nucleic acid extraction with Gdm+-salts and silica (Boom et al., 1990).
Using the procedure described in Tab.1, we were able to extract high quality and high molecular weight DNA in 16 aromatic and medicinal Colombian plant species with variable yields of essential oils (Tab.2). The DNA extracted by the protocol A-2X is of high molecular weight and high quality which is probably explained by the chemistry used both in the extraction buffer and during the DNA purification (Fig.1). Furthermore, the use of solutions with Gdm+ and the nature of the silica-based extraction columns allowed efficient elimination of contaminants and denatured proteins and recovering of large amounts of high molecular weight DNA.
The absorbance measurements of DNA at 280 nm, 260 nm and 230 nm, allowed to calculate two types of ratios widely adopted and qualify the DNA as "pure" or of high-quality (in average, A260/A280 1.8 and A260/A230 2.0). Nevertheless, some accessions exhibited ratios slightly away from these values. Digestion with the restriction enzyme MseI carried out in nine accessions with variable yields of essential oils and different ratios of purity (Tab.2), showed that the potential contaminants detected by these ratios did not affect the use of the extracted DNA in restriction digestion and suggest that these ratios per se are not reliable or accurate indicators of the usefulness of the extracted DNA in this study. Amplification of four non-coding chloroplast regions in the six evaluated accessions resulted to be successful (Fig.3) and suitable for sequencing and for subsequent population genetic studies. Likewise, PCR amplification with three selective AFLP primer combinations resulted to be optimal and showed several polymorphisms for each of the accessions evaluated (Fig.4), which is desirable for the study of genetic variation and the development of molecular markers.
In summary, the DNA extracted using the protocol A-2X met the requirements of quality and quantity to apply different molecular techniques such as PCR, SSRs, RAPDs, ISSRs, AFLPs, sequencing, construction of genomic libraries, among other applications (Mace et al., 2003; Marva et al., 2007; Sarwat et al., 2006). As a more efficient and less expensive alternative to the use of commercial kits for nucleic acid extraction, the protocols that use Gdm+-salts in conjunction with membranes or particles of silica and report the chemistry of the extraction may be easily implemented in the isolation of DNA from recalcitrant samples (Ivanova et al., 2008).
Acknowledgments
This work was funded by the Ministerio de Agricultura y Desarrollo Rural (MADR) of Colombia, contract No. 064- 2007V7163-50-07, the Universidad Nacional de Colombia and Colciencias through a research fellowship under the "Programa de Jóvenes Investigadores e Innovadores" contract No. 155-2008.
Literature cited
Alexander, P.J., G. Rajanikanth, C.D. Bacon, and C.D. Bailey. 2007. Recovery of plant DNA using a reciprocating saw and silicabased columns. Mol. Ecol. Notes 7, 5-9. [ Links ]
Bakkali, F., S. Averbeck, D. Averbeck, and M. Idaomar. 2008. Biological effects of essential oils - A review. Food Chem. Toxicol. 46(2), 446-475. [ Links ]
Balunas, M.J. and A.D. Kinghorn. 2005. Drug discovery from medicinal plants. Life Sci. 78, 431-441. [ Links ]
Bernardo, R. 2008. Molecular markers and selection for complex traits in plants: learning from the last 20 years. Crop Sci. 48, 1649-1664. [ Links ]
Boom, R., C.J.A. Sol, M.M.M. Salimans, C.L. Jansen, P.M.E. Wertheim-Van Dillen, and J. Van der Noordaa. 1990. Rapid and simple method for purification of nucleic acids. J. Clin. Microbiol. 28(3), 495-503. [ Links ]
Boom, R., C.J.A. Sol, M. Beld, J. Weel, J. Goudsmit, and P.M. Wertheim-Van Dillen. 1999. Improved silica-guanidinium thiocyanate DNA isolation procedure based on selective binding of bovine alpha-casein to silica particles. J. Clin. Microbiol. 37(3), 615-619. [ Links ]
Bowtell, D.D.L. 1987. Rapid isolation of eukaryotic DNA. Anal. Biochem. 162(2), 463-465. [ Links ]
Breton, C., D. Claux, I. Metton, G. Skorski, and A. Bervillé. 2004. Comparative study of methods for DNA preparation from olive oil samples to identify cultivar SSR alleles in commercial oil samples: possible forensic applications. J. Agric. Food Chem. 52, 531-537. [ Links ]
Canter, P.H., H. Thomas, and E. Ernst. 2005. Bringing medicinal plants into cultivation: opportunities and challenges for biotechnology. Trends Biotechnol. 23(4), 180-185. [ Links ]
Castellino, F.J. and R. Barker. 1968. The denaturing effectiveness of guanidinium, carbamoylguanidinium and guanylguanidinium salts. Biochemistry 7(11), 4135-4138. [ Links ]
Collard, B.C.Y., M.Z.Z. Jahufer, J.B. Brouwer, and E.C.K. Pang. 2005. An introduction to markers, quantitative trait loci (QTL) mapping and marker-assisted selection for crop improvement: the basic concepts. Euphytica 142, 169-196. [ Links ]
Cowan, M.M. 1999. Plant products as antimicrobial agents. Clin. Microbiol. Rev. 12(4), 564-582. [ Links ]
Cox, R.A. 1968. The use of guanidinium chloride in the isolation of nucleic acids. pp. 120-129. In: Grossman L. and K. Moldave (eds.). Methods in enzymology. Vol. 12. Academic Press, Orlando, FL. [ Links ]
Cheng, F.S., S.K. Brown, and N.F. Weeden. 1997. A DNA extraction protocol from various tissues in wood species. HortScience 32(5), 921-922. [ Links ]
Chomczynski, P. and N. Sacchi. 1987. Single-step method of RNA isolation by acid guanidinium thiocyanate-phenol-chloroform extraction. Anal. Biochem. 162, 156-159. [ Links ]
Dellaporta, S.L., J. Wood, and J.B. Hicks. 1983. A plant DNA minipreparation: version II. Plant Mol. Biol. Rep. 1(4), 19-21. [ Links ]
Ding, L.-W., Q.-Y. Sun, Z.-Y. Wang, Y.-B. Sun, and Z.-F. Xu. 2008. Using silica particles to isolate total RNA from plant tissues recalcitrant to extraction in guanidine thiocyanate. Anal. Biochem. 374, 426-428. [ Links ]
Doyle, J.J. and J.L. Doyle. 1990. A rapid total DNA preparation procedure for fresh plant tissue. Focus 12, 13-15. [ Links ]
Fofana, B., L. Harvengt, J.P. Baudoin, and P. du Jardin. 1997. New primers for polymerase chain amplification of cpDNA intergenic spacers in Phaseolus phylogeny. Belg. J. Bot. 129, 118-122. [ Links ]
Gómez-Galera, S., A.M. Pelacho, A. Gené, T. Capell, and P. Christou. 2007. The genetic manipulation of medicinal and aromatic plants. Plant Cell Rep. 26, 1689-1715. [ Links ]
Greenstein, J.P. 1938. Sulfhydryl groups in proteins: I. Egg albumin in solutions of urea, guanidine and their derivates. J. Biol. Chem. 125, 501-513. [ Links ]
Greenstein, J.P. 1939a. Sulfhydryl groups in proteins: II. Edestin, excelsin and globin in solutions of guanidine hydrochloride, urea and their derivates. J. Biol. Chem. 128, 233-240 [ Links ]
Greenstein, J.P. 1939b. Sulfhydryl groups in proteins: III. The effect on egg albumin of various salts of guanidine. J. Biol. Chem. 130, 519-526. [ Links ]
Greenstein, J.P. and J.T. Edsall. 1940. The effect of denaturing agents on myosin: I. Sulfhydryl groups as estimated by porphyrindin titration. J. Biol. Chem. 133, 397-408. [ Links ]
Hamilton, H.B. 1999. Four primer pairs for the amplification of chloroplast intergenic regions with intraespecific variation. Mol. Ecol. 8, 521-523. [ Links ]
Ivanova, N.V., A.J. Fazekas, and P.D.N. Hebert. 2008. Semiautomated, membrane-based protocol for DNA isolation from plants. Plant Mol. Biol. Rep. 26, 186-198. [ Links ]
Jobes, D.V., D.L. Hurley, and L.B. Thien. 1995. Plant DNA isolation: a method to efficiently remove polyphenolics, polysaccharides and RNA. Taxon 44, 379-386. [ Links ]
Khanuja, S.P.S., A.K. Shasany, M.P. Darokar, and S. Kumar. 1999. Rapid isolation of DNA from dry and fresh samples of plants producing large amounts of secondary metabolites and essential oils. Plant Mol. Biol. Rep. 17, 1-7. [ Links ]
Krizman, M., J. Jakse, D. Baricevic, B. Javornik, and M. Prosek. 2006. Robust CTAB-activated charcoal protocol for plant DNA extraction. Acta Agric. Slov. 87(2), 427-433. [ Links ]
Kumar, J. and P.K. Gupta. 2008. Molecular approaches for improvement of medicinal and aromatic plants. Plant Biotechnol. Rep. 2, 93-112. [ Links ]
Mace, E.S., H.K. Buhariwalla, and J.H. Crouch. 2003. A highthroughput DNA extraction protocol for tropical molecular breeding programs. Plant Mol. Biol. Rep. 21, 459a-459h. [ Links ]
Marieschi, M., A. Torelli, F. Poli, G. Sacchetti, and R. Bruni. 2009. RAPD-based method for the quality control of mediterranean oregano and its contribution to pharmacognostic techniques. J. Agric. Food Chem. 57, 1835-1840. [ Links ]
Marva, A., H. Padh, and N. Shrivastava. 2007. Plant genomic DNA isolation: an art or a science. Biotechnol. J. 2, 386-392. [ Links ]
Mason, P.E., G.W. Neilson, C.E. Dempsey, A.C. Barnes, and J.M. Cruickshank. 2003. The hydration structure of guanidinium and thiocyanate ions: implications for protein stability in aqueous solution. PNAS 100(8), 4557-4561. [ Links ]
Matasyoh, L.G., F.N. Wachira, M.G. Kinyua, A.W.T. Muigai, and T.K. Mukiama. 2008. Leaf storage conditions and genomic DNA isolation efficiency in Ocimum gratissimum L. from Kenya. Afr. J. Biotechnol. 7(5), 557-564. [ Links ]
Meneses, R., R.E. Ocazionez, J.R. Martínez, and E.E. Stashenko. 2009. Inhibitory effect of essential oils obtained from plants grown in Colombia on yellow fever virus replication in vitro (online). Ann. Clin. Microbiol. Antimicrob. 8, 8, http://www.ann-clinmicrob.com/content/8/1/8 ; consulted: November, 2011. [ Links ]
Michiels, A., W. Van den Ende, M. Tucker, L. Van Riet, and A. Van Laere. 2003. Extraction of high-quality genomic DNA from latex-containing plants. Anal. Biochem. 315, 85-89. [ Links ]
Nerio, L.S., J. Olivero-Verbeland, and E.E. Stashenko. 2009. Repellent activity of essential oils from seven aromatic plants grown in Colombia against Sitophilus zeamais Motschulsky (Coleoptera). J. Stored Prod. Res. 45, 212-214. [ Links ]
Padmalatha, K. and M.N.V. Prasad. 2006. Optimization of DNA isolation and PCR protocol for RAPD analysis of selected medicinal and aromatic plants of conservation concern from peninsular India. Afr. J. Biotechnol. 5(3), 230-234. [ Links ]
Pirttilä, A.M., M. Hirsikorpi, T. Kämäräinen, L. Jaakola, and A. Hohtola. 2001. DNA isolation methods for medicinal and aromatic plants. Plant Mol. Biol. Rep. 19, 273-273. [ Links ]
Porebski, S., L.G. Bailey, and B.R. Baum. 1997. Modification of a CTAB DNA extraction protocol for plants containing high polysaccharide and polyphenol components. Plant Mol. Biol. Rep. 15(1), 8-15. [ Links ]
Rogstad, S.H. 2003. Plant DNA extraction using silica. Plant Mol. Biol. Rep. 21, 463a-463g. [ Links ]
Sangwan, R.S., U. Yadav, and N.S. Sangwan. 2000. Isolation of genomic DNA from defatted oil seed residue of opium poppy (Papaver sominiferum). Plant Mol. Biol. Rep. 18, 265-270. [ Links ]
Sarwat, M., M.S. Negi, M. Lakshmikumaran, A.K. Tyagi, S. Das, and P.S. Srivastava. 2006. A standardized protocol for genomic DNA isolation from Terminalia arjuna for genetic diversity analysis. Electron. J. Biotechnol. 9(1), 86-91. [ Links ]
Sassa, H. 2007. A technique to isolate DNA from woody and herbaceous plants by using a silica-based plasmid extraction column. Anal.Biochem. 363, 166-167. [ Links ]
Svedberg, T. 1937. The ultra-centrifuge and the study of highmolecular compounds. Nature 139, 1051-1062. [ Links ]
Svoboda, K.P. and J.B. Hampson. 1999. Bioactivity of essential oils of selected temperate aromatic plants: antibacterial, antioxidant, antiinflammatory and other related pharmacological activities. pp. 43-49. In: Special Chemicals for the 21st Century. Ademe, Paris. [ Links ]
Taberlet, P., L. Gielly, G. Pautou, and J. Bouvet. 1991. Universal primers for amplification of three non-coding regions of chloroplast DNA. Plant Mol. Biol. 17, 1105-1109. [ Links ]
Vos, P., R. Hogers, M. Bleeker, M. Reijans, T. Van de Lee, M. Hornes, A. Frijters, J. Pot, J. Peleman, M. Kuiper, and M. Zabeau. 1995. AFLP: a new technique for DNA fingerprinting. Nucleic Acids Res. 23(21), 4407-4414. [ Links ]