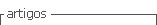Serviços Personalizados
Journal
Artigo
Indicadores
-
 Citado por SciELO
Citado por SciELO -
 Acessos
Acessos
Links relacionados
-
 Citado por Google
Citado por Google -
 Similares em
SciELO
Similares em
SciELO -
 Similares em Google
Similares em Google
Compartilhar
Agronomía Colombiana
versão impressa ISSN 0120-9965
Agron. colomb. v.29 n.3 Bogotá set./dez. 2011
PLANT BREEDING, GENETIC RESOURCES & MOLECULAR BIOLOGY
Identification in silico of SSR markers for genotyping Hevea sp. clone gardens in Colombia
Identificación in silico de marcadores SSR para genotipificar jardines clonales de Hevea sp. de Colombia
Ibonne Aydee García R.1,3,Sandra Milena González S.1, Dolly Montoya C.1 and Fabio Aristizabal.2
1Institute of Biotechnology, Universidad Nacional de Colombia. Bogota (Colombia).2Institute of Biotechnology and Faculty of Science, Universidad Nacional de Colombia. Bogotá (Colombia).
3Corresponding author. ibonne@gmail.com Received for publication: 6 November, 2009. Accepted for publication: 2 November, 2011.
ABSTRACT The rubber crops profitability depends largely on genotypes established in plantations, meaning that clone identity must be ascertained. This work was aimed at identifying commercial clones Hevea sp. by microsatellites. Primers were designed from sequences reported in Genbank using Primer3, PrimerQuest and OlgoPerfect software for PCR amplification of microsatellites. The primers so obtained were thermodynamically analysed by Oligo Analyzer 3.1 software and experimentally evaluated on 12 Hevea sp. clones. The 15 of the 561 microsatellite markers were selected; they had 2- and 3-bp repeat motifs and 11- to 23-bp repeat extension ranges. The most informative ones were microsatellites amplified with SSRH103, SSRH134, SSRH510 and SSRH516 primers with seven alleles and SSRH403 primers with eight alleles. Four microsatellite markers were sufficient for discriminating 10 of the 12 clones. Clustering analysis involved all the markers on the clones evaluated here, showing Brazilian clones' narrow genetic base compared to Asiatic ones. The current work provides new markers and joins work published by other authors for identifying and diversity studies of natural rubber clone.
Key words: polymorphism, molecular markers, Hevea brasiliensis, SSR, clone, PCR, natural rubber.
RESUMEN
La rentabilidad del cultivo de caucho depende en gran medida de los genotipos establecidos en plantación, por lo tanto es necesario asegurar la identidad de los clones. Este trabajo tuvo como objetivo identificar clones comerciales de Hevea sp. mediante microsatélites. Se diseñaron primers a partir de secuencias reportadas en el Genbank con los programas Primer3, PrimerQuestSM y OlgoPerfectSM para amplificación por PCR de microsatélites. Los primers obtenidos se analizaron termodinámicamente mediante el programa Oligo Analizer 3.1 y se evaluaron experimentalmente sobre 12 clones de Hevea sp. Se seleccionaron 15 de 561 marcadores microsatélites con motivos de repetición de 2 y 3 pb y rangos de extensión entre 11 y 23 repeticiones. Los más informativos fueron: con siete alelos los microsatélites amplificados con lo primers SSRH103, SSRH134, SSRH510 y SSRH516, con ocho alelos los primers SSRH403. Cuatro marcadores microsatélites fueron suficientes para discriminar diez de los 12 clones. El análisis de agrupamiento realizado con la totalidad de los marcadores sobre los clones evaluados, evidencia la estrecha base genética de los clones brasileños respecto a los asiáticos. Este trabajo aporta nuevos marcadores y se suma a los publicados por otros autores, para la identificación y estudios de diversidad de clones de caucho natural.
Palabras clave: polimorfismo, marcadores moleculares, Hevea brasiliensis, SSR, clon, PCR, caucho natural.
Introduction
Natural rubber (Hevea sp.) crops have its origin in Amazon region; it belongs to the Euphorbiáceae family grouping several characteristic genera for latex production. Hevea brasiliensis is the most important specie from this genus due to high natural rubber production and the excellent physical-chemical properties which synthetic rubber cannot provide (Polhamus, 1962; Mooibroek and Cornish, 2000; Cornish, 2001).
Asia has the greatest planted area of natural rubber; this continent pioneered worldwide latex marketing. Around 7,000,000 ha have been established in Indonesia, Thailand and Malaysia, such production representing 70% of the world market for this biopolymer. Latin-America represents just 3% of such production (365,000 ha planted) despite being the center of origin of this species (IRSG, 2010). Colombia currently has 29,917 ha planted with rubber (MADR, 2010); it is hoped to broaden this to supply natural rubber needs by promoting new seeding programs, 889,674 ha being available which have favourable climatic conditions for the crop (Candelo and Motta, 1996).
Establishing new plantations has show that existing clone gardens usually do not have clear historical records concerning the origin of their vegetal material. Sanitary control and certification techniques must be implemented to overcome such limitation within the legal framework provided by ICA resolution 001478/2006 stating the need for follow-up, sanitary surveillance and genetic identification in natural rubber propagation nurseries to guarantee the vegetal material's source and quality, preventing low yields in adult plantations and the introduction and dissemination of diseases.
Vegetal material destined for this specie's propagation is currently identified by the isoenzyme technique (Chevallier, 1988) which has limitations related to specificity and costs. The limited number of alleles which can be detected this type of marker is one of its main disadvantages (i.e. poor discrimination power) (Vera et al., 1999; Belleti et al., 1992). This technique is linked to gene expression phenotypes which could be influenced by environmental factors and the crop's development stages, meaning that it is hardly reproducible.
Recently DNA-based molecular markers have been widely used due to their sensitivity and rapidity in varietal identification in different vegetal species (Raina et al., 2001; Vera et al., 1999; Prince et al., 1995), as well as leading to discriminating individuals having closely-related genotypes (Ilbi, 2003). Such markers are classified into two large groups: hybridisation-based molecular markers and the PCR-based molecular markers. The latter are more used due to the low DNA concentration required for their development, their ability to amplify genome sequences from preserved tissue and being methodologically accessible by small laboratories regarding equipment, ease of use and cost. RAPD, SSR or microsatellites, AP-PCR and AFLPs are some of the currently available PCR-based markers (Semagn, 2006).
Microsatellites or simple sequence repeats (SSR) are amongst the most efficient molecular markers due to their co-dominant multi-allele nature and their broad random distribution throughout the whole genome, thereby making them a highly polymorphic marker. SSR have been successfully used in plants in genetic diversity studies, constructing genetic maps (Lespinasse et al., 2000) assisted improvement by molecular markers and genotype identification studies (Okogbenin et al., 2006; Rajeev et al., 2005).
RAPD molecular markers have been used for genotyping natural rubber clones (Varghese et al., 1997; Hernández et al., 2006; Nakkanong et al., 2008); however, this is not an efficient methodology for mass evaluation of clones due to the technique's inherent characteristics such as low reproducibility, low discrimination power and difficulty with profile analysis. Studies have been developed using microsatellite markers. Lekawipat et al. (2003) reported the SSR M574 marker as being highly polymorphic after evaluating 108 Hevea brasiliensis accessions and obtaining 21 alleles. Saha et al. (2005) reported four markers (HMAC4, HMAC5, HMCT1 and HMCT5) for discriminating 27 Hevea sp. clones. Studies by Saha et al. (2007) reported that the HMGR marker was useful for studying genetic variability in wild Hevea brasiliensis material. Feng et al. (2009) have recently published a set of markers designed from EST sequences stored in GenBank for evaluating genetic diversity and SSR transferability between Hevea species.
Four SSR markers proposed by Saha et al. (2005) were used in preliminary assays in six Hevea sp. clones having commercial interest for Colombia; these assays showed that all of them could only be discriminated by the HMCT5 marker. However, a clear and reproducible banding pattern could not be obtained, even when modifying annealing temperatures (Tm) and using PCR adjuvants. The above work also reports an evaluation of the EST-SSR markers proposed by Feng et al. (2009) on 67 rubber clones and by Saha et al., (2007) on six clones respectively, obtained low polymorphism. The present work was aimed at identifying new microsatellite markers having greater discrimination power for genotyping Hevea sp. clones from genome sequences reported in GenBank databases for experimentally designing and evaluating primers leading to easy amplification and reading.
Methodology
Prior evaluation. The 12 clones evaluated (Tab.1) in this study had been previously analysed with the HMCT1, HMCT5, HMAC1 and HMAC4 microsatellites described for identifying Asian natural rubber genotypes (Saha et al., 2005). These PCR and allele visualisation conditions were performed as proposed by the authors.
Oligo design. Sequences found in Genbank were manually analysed using the search word "Hevea brasiliensis microsatellite sequence." Sequences were initially chosen which presented two and three repeat motifs (minimum 11 and 4 repeats from the motif, respectively). It was also was verified whether there had been any redundancy in a reduced genetic base had to be discriminated, such as the American clones from the IAN and FX series.
The information found in GenBank for some of the selected sequences included some primer sets, together with their amplification conditions, which were not included due to them not fitting the selection criteria for the primers used in this work (lower Tm and shorter length) since using primers having low annealing Tm and short size increases the probability of non-specific amplimers (Abd-Elsalam et al., 2003).
The 560 pairs of primers were obtained from the selected sequences (i.e. four or five per sequence); 15 pairs were selected after analysis with OligoAnalyzer 3.1 software was they fulfilled the thermodynamic criteria descried in the methodology and were sent to be synthesised. Two of these primer sets led to amplifying microsatellites having three bp repeat domains in spite of not fulfilling all previously established criteria (Tab.1); this parameter allowed better differentiation of the fragments obtained.
Prior knowledge of Tm obtained with the design programmes led to a better approach to establishing the PCR cycle, specific amplimers being thereby obtained for all the primers. These parameters avoided having to apply formulae for finding Tm and these values' discrepancy with PCR components, as well as using empirical thermocycling assays with touchdown (Don et al., 1991; Roux, 2009). PCR was optimised regarding MgCl2, dNTP, primer and template DNA concentrations; hybridisation temperature based on the lower limit of the range selected for designing primers was also standardised (54 to 59°C experimental hybridisation Tm) (Tab.1). As specific, easily interpreted amplifications were obtained with 10 of the 13 primer sets, this showed that the selection criteria defined for their design were sound.
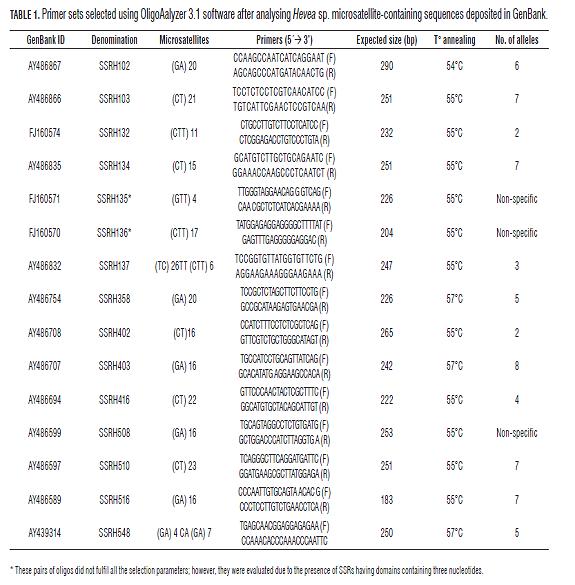
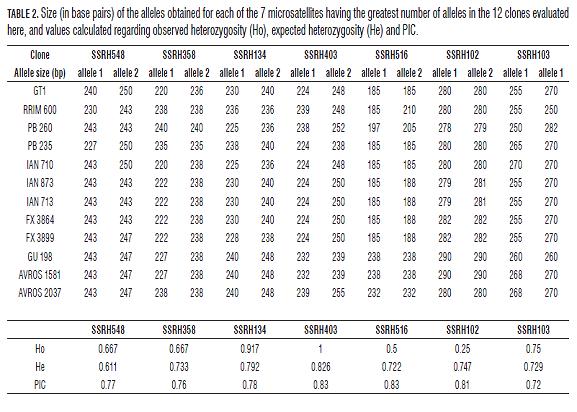
It was found that 3 of the 15 primer sets evaluated in the 12 Hevea clones had non-specific band patterns (SSRH136, SSRH135 and SSRH508). Two of the markers only generated two alleles (SSRH132 and SSRH 402); the SSRH137 and SSRH416 markers generated three and four alleles, respectively and seven had 5 to 8 alleles (Tab.2). The sizes obtained ranged from 222 to 290 bp, these being congruent with size predicted by primer design programmes (Fig.1, Tab.2). These diversity and polymorphism measurements calculated for microsatellites which generated more than 5 alleles ranged from 0.66 to 0.82 for He, 0.25 to 1.00 for Ho and 0.72 to 0.83 for PIC (Tab.2).
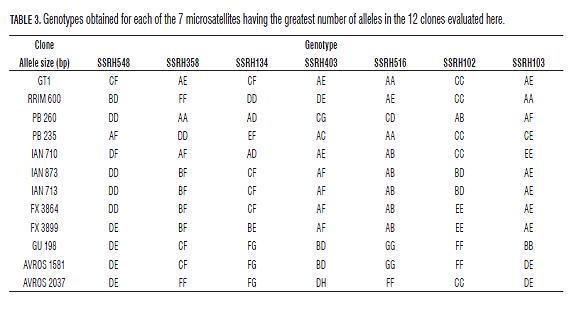
The SSRH403 microsatellite (He=1.000, Ho=0.826 and PIC=0.830) amplified the greatest number of alleles (8) and could discriminate the 12 clones into eight different groups. The PB260 clone was differentiated from the other Asian clones with 7 of the 11 markers evaluated, this being the most divergent clone (both Asian and American). The SSRH548, SSRH358, SSRH134 and SSRH403 microsatellites discriminated Asian clones PB 235, RRIM 600 and GT-1 (Ho=0.667-1.000, He=0.611-0.826 and PIC 0.76-0.87). The GU-191 (from Guatemala) and AVROS-1981 (Asian) clones were only differentiated with microsatellite SSRH103 (Ho=0.750, He=0.729 and PIC=0.720). The American clones obtained in Brazilian genetic improvement programmes (FX 3899, FX 3864, IAN 873, IAN 710 and IAN 713) were discriminated by fewer microsatellites grouped together with most markers. The FX clones could only be differentiated with the SSRH134 microsatellite. IAN 713 and IAN 873 clones were not differentiated with any of the 10 microsatellites. The minimum number of microsatellites allowing 10 of the 12 clones to be differentiated was four (i.e. SSRH403, SSRH548, SSRH358 and SSRH103). Tab.3 summarises the foregoing, showing the different genotypes obtained for all the clones.
Similarity analysis with the 10 most polymorphic microsatellites differentiated 10 of the 12 clones evaluated; a group clustering the five American clones (Brazilian) included in the study (0.48 similarity) was also observed, the selected sequences by using alignment with the bl2seq algorithm (Altschul et al., 1990). These sequences were used for designing primers with OligoPerfect (http://tools.invitrogen.com/content.cfm?pageid=9716), Primer3 (Rozen and Skaletsky, 2000) and PrimerQuest (http://www.idtdna.com/Scitools/Applications/Primerquest/Default. aspx) software, taking the following criteria into account: 18 to 24 base pair (bp) primer size, 180 to 300 bp expected fragment size, 55 to 62°C fusion Tm and 45 to 55% GC. IDT OligoAnalyzer 3.1 software (http://www.idtdna.com/Scitools/Scitools.aspx) was used for evaluating the primers' thermodynamics. Maximum 3 bp complementariety was taken into account to avoid branch formation; homodimer and heterodimer formation was then determined (maximum 6 bp complementariety) (Montoya, 2009). NCBI Blast (Altschul et al., 1990) was then used for alignments to ensure specificity and homology between primers and target sequence.
Vegetal material: 12 Hevea brasiliensis clones or their Hevea benthamiana hybrids were analysed to ascertain designed primer polymorphism (Tab.2) using three individuals per clone. Foliar material were collected from Mavalle (Remolinos, Meta department), Vorágine (Medellín, Antioquia department) and Rincón Llanero (Puerto Gaitán, Meta department) clone gardens which had clear vegetal material source records. Four leaflets from each clone were stored in sealed hermetic bags with silica gel as preservative and then taken to the laboratory for total DNA extraction.
Nucleic acid extraction. Genomic DNA was extracted from each sample using the method proposed by Varghese et al. (1997). The DNA so obtained was visualised on 0.8% agarose gel and quantified by fluorometry (Quant-iT ds BR assay kit, Invitrogen).
PCR amplification and microsatellite analysis. The PCR reaction was carried out for 25 µL final volume containing 25 ng genomic DNA, 0.4 µM of each oligo, 1.5 mM MgCl2, 200 mM dNTPs and 1 Taq polymerase unit. The master mix was placed in a MyCycler (BioRad) thermocycler using the following amplification programme: 94°C initial denaturing for 3 min, followed by 35 denaturing cycles at 94°C for 15 s, annealing for 1 min (temperatures reported in Tab.1 for each set of primers), extension at 72°C for 15 s and a final 7 min extension step at 72°C.
The amplimers obtained were run on 7% polyacrylamide gels and visualised by silver staining (Bassam et al., 1991). The sizes of the alleles obtained by microsatellite were determined using 10 bp molecular weight ladder (Invitrogen) as reference.
Data analysis: The number of alleles detected for each microsatellite so amplified was estimated in the 12 clones evaluated. The degree of polymorphism was calculated from expected heterozygosity (He) by applying the formula He= 1- S pi 2 where p was the frequency of the ith allele in the genotypes examined. Observed heterozygosity (Ho) was calculated using the ratio between the number of heterozygotes regarding the amount of total genotypes analysed using GENALEX 6 software (Peakall and Smouse, 2000). Polymorphism information content (PIC) was calculated using the PIC= 1- S fi 2 i = 1 equation where fi 2 was the frequency of the ith allele (Anderson et al., 1993). Similarity was analysed using NTSYS-pc 2.1 software (Rohlf, 2006); the Dice index was used for generating a similarity matrix, followed by a dendrogram using the UPGMA algorithm.
Results and discussion
Only two of the 12 clones evaluated in this study (Asian clones GT 1 and PB 235) had been analysed by Saha et al. (2005) whose results were confirmed for 3 of the microsatellites (HMCT1, HMAC4 and HMCT5), unlike the results obtained with microsatellite HMAC5. The authors reported two alleles for GT 1 (270/272 bp) and PB 235 (272/274 bp) with this microsatellite; a single 265 bp allele was observed in this work for both clones. It was also found that the clones being studied (FX 3899, FX 3864, IAN 873, IAN 713, GU 191 and AVROS 1981) were not discriminated using the four reported microsatellites as a set (data not shown). It is thus recommended using these microsatellites for identifying some Asian clones present in Colombian clone gardens; however, additional markers must be included for American clones to allow them to be differentiated.
The 120 of the 604 sequences found in the search for microsatellite regions contained in the H. brasiliensis genome in GenBank using the search word "H. brasiliensis microsatellite sequence" were selected for designing primers; bearing in mind repeat motifs' greater sizes and the greater number of repeats, work on other vegetal species such as Capsicum annuun have reported increased polymorphism in SSR sequences having the aforementioned characteristics (Yi et al., 2006).
The search did not include microsatellites present in H. brasiliensis expressed sequence tag (EST) libraries (Okogbenin et al., 2006), due to them usually being more conserved between individuals from the same specie and related species in contrast to genomic SSRs, thereby implying less polymorphism (Eujay et al., 2002), such characteristic not being useful in this work given that vegetal material having a reduced genetic base had to be discriminated, such as the American clones from the IAN and FX series.
The information found in GenBank for some of the selected sequences included some primer sets, together with their amplification conditions, which were not included due to them not fitting the selection criteria for the primers used in this work (lower Tm and shorter length) since using primers having low annealing Tm and short size increases the probability of non-specific amplimers (Abd-Elsalam et al., 2003).
The 560 pairs of primers were obtained from the selected sequences (i.e. four or five per sequence); 15 pairs were selected after analysis with OligoAnalyzer 3.1 software was they fulfilled the thermodynamic criteria descried in the methodology and were sent to be synthesised. Two of these primer sets led to amplifying microsatellites having three bp repeat domains in spite of not fulfilling all previously established criteria (Tab.1); this parameter allowed better differentiation of the fragments obtained.
Prior knowledge of Tm obtained with the design programmes led to a better approach to establishing the PCR cycle, specific amplimers being thereby obtained for all the primers. These parameters avoided having to apply formulae for finding Tm and these values' discrepancy with PCR components, as well as using empirical thermocycling assays with touchdown (Don et al., 1991; Roux, 2009). PCR was optimised regarding MgCl2, dNTP, primer and template DNA concentrations; hybridisation temperature based on the lower limit of the range selected for designing primers was also standardised (54 to 59°C experimental hybridisation Tm) (Tab.1). As specific, easily interpreted amplifications were obtained with 10 of the 13 primer sets, this showed that the selection criteria defined for their design were sound.
It was found that 3 of the 15 primer sets evaluated in the 12 Hevea clones had non-specific band patterns (SSRH136, SSRH135 and SSRH508). Two of the markers only generated two alleles (SSRH132 and SSRH 402); the SSRH137 and SSRH416 markers generated three and four alleles, respectively and seven had 5 to 8 alleles (Tab.2). The sizes obtained ranged from 222 to 290 bp, these being congruent with size predicted by primer design programmes (Fig.1, Tab.2). These diversity and polymorphism measurements calculated for microsatellites which generated more than 5 alleles ranged from 0.66 to 0.82 for He, 0.25 to 1.00 for Ho and 0.72 to 0.83 for PIC (Tab.2).
The SSRH403 microsatellite (He=1.000, Ho=0.826 and PIC=0.830) amplified the greatest number of alleles (8) and could discriminate the 12 clones into eight different groups. The PB260 clone was differentiated from the other Asian clones with 7 of the 11 markers evaluated, this being the most divergent clone (both Asian and American). The SSRH548, SSRH358, SSRH134 and SSRH403 microsatellites discriminated Asian clones PB 235, RRIM 600 and GT-1 (Ho=0.667-1.000, He=0.611-0.826 and PIC 0.76-0.87). The GU-191 (from Guatemala) and AVROS-1981 (Asian) clones were only differentiated with microsatellite SSRH103 (Ho=0.750, He=0.729 and PIC=0.720). The American clones obtained in Brazilian genetic improvement programmes (FX 3899, FX 3864, IAN 873, IAN 710 and IAN 713) were discriminated by fewer microsatellites grouped together with most markers. The FX clones could only be differentiated with the SSRH134 microsatellite. IAN 713 and IAN 873 clones were not differentiated with any of the 10 microsatellites. The minimum number of microsatellites allowing 10 of the 12 clones to be differentiated was four (i.e. SSRH403, SSRH548, SSRH358 and SSRH103). Tab.3 summarises the foregoing, showing the different genotypes obtained for all the clones.
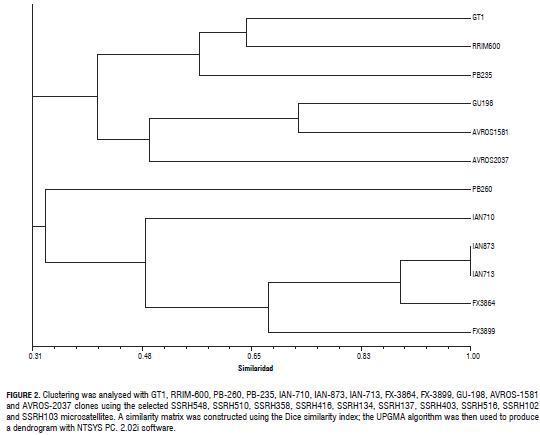
Similarity analysis with the 10 most polymorphic microsatellites differentiated 10 of the 12 clones evaluated; a group clustering the five American clones (Brazilian) included in the study (0.48 similarity) was also observed thus confirming this group's reduced gene base (Fig.2). Similar results were obtained by Hernández et al. (2006) who evaluated Asian and Brazilian clones using RAPD markers. The foregoing could explain the three commercial clones from the IAN series which, together with the FX 3864 clone, share the maternal parent (PB 86). The GU 198 clone was grouped with Asian AVROS clones (0.71 similarity), AVROS 1581 (0.48) and AVROS 2087 (0.34), thereby agreeing with the results obtained by Hernández et al. (2006) who concluded that this clone was produced by crossing Asian parental clones; however, it is known that one of its parental clones corresponded to the American FX 16 clone. The RRIM 600, GT 1 and PB 235 clones were grouped at a 0.49 genetic distance and the PB 260 clone was grouped at 0.31 genetic distance.
The IAN 873 and IAN 713 clones could not be differentiated with any of the 10 microsatellites. It is thus considered that evaluation of microsatellite markers should be considered as this leads to discriminating genotypes of clones present in Colombian clone gardens. Recent work has offered other useful markers in this field, such as that reported by Souza et al. (2009) who produced 30 microsatellites which were then applied to studying the genetic diversity of 30 different H. brasiliensis genotypes, including genotypes from other Hevea species; the loci analysed presented observed and expected heterozygosity ranging from 0.13 to 0.88 and 0 to 0.89, respectively. Work by Le Guen et al. (2009) reported 15 microsatellites applied to studying the genetic diversity of natural rubber germplasm collections established in French Guyana, Peru and Brazil, all having high polymorphism.
It is worth stressing that the study presented here was not aimed at analysing the genetic diversity of the clones included in it; however, the results have led to us suggesting using the microsatellites evaluated here for this type of study. This work has contributed towards identifying Hevea sp. clones for organising clone gardens which are the commercial source of vegetal material distribution for establishing plantations, since suitable selection of clones to be planted in a determined region will greatly ensure latex performance and thus increase producers' economic benefits. Prior identification of clones is needed as they perform differently according to a particular region's characteristics. It is clear that when a plantation's genetic base can be controlled then better profitability can be expected, bearing in mind that natural rubber is a late performance crop thereby involving high management costs.
Conclusions
The preliminary evaluation of the 12 clones being studied with microsatellites reported in the pertinent literature for identifying some Asian rubber genotypes showed that they were unable to differentiate American clones present in Colombian clone gardens. This study made advances in characterising four clones which could not be differentiated by the microsatellites reported by Saha et al. (2005). Implementing thermodynamic analysis programmes has led to establishing parameters for selecting and designing oligos, thereby simplifying standardising PCR amplification the conditions in the laboratory. Sets of oligos have thus been obtained which have allowed microsatellite amplification, in turn, managing to differentiate 10 of the 12 Hevea sp. clones evaluated in this study with a minimum number of four of these markers, thereby abbreviating identification of the 12 clones.
Analysis of clustering the clones characterised in this work has revealed the need for continuing to apply molecular tools leading to the discrimination of all the clones present in Colombia so that they can be used in varietal identification and genetic variability studies.
Acknowledgements
We would like to acknowledge the project entitled, Towards certifying Hevea brasiliensis commercial material of interest for Colombia by using molecular techniques (Contribución a la certificación por técnicas moleculares de material comercial de Hevea brasiliensis de interés para Colombia) co-financed by MADR and carried out in the Instituto de Biotecnología de la Universidad Nacional covered by an agreement with the Instituto Amazónico de Investigaciones Científicas - Sinchi.
Literature cited
Abd-Elsalam, K.A. 2003. Bioinformatic tools and guideline for PCR primer design. Afr. J. Biotechnol. 2(5), 91-95. [ Links ]
Altschul, S.F., W. Gish, W. Miller, E.W. Myers, and D.J. Lipman. 1990. Basic local alignment search tool. J. Mol. Biol. 215, 403-410. [ Links ]
Anderson, J., A.G.A. Churchill, J.E. Sutrique, S.D. Tanksley, and M.E. Sorrthes. 1993. Optimizing parental selection for genetic linkage maps. Genome 36, 181-186. [ Links ]
Bassam, B.J., G. Caetano-Anolles, and P.M. Gresshoff. 1991. Fast and sensitive silver staining of DNA in polyacryelmide geles. Anal. Biochem. 196, 80-83. [ Links ]
Belletti, P., S. Lanteri, F. and Saracco. 1992. Allozyme variability in Capsicum. pp. 221-226. In: Proceedings of the Eighth Meeting on Genetics and Breeding on Capsicum and Eggplant. Rome, Italy. [ Links ]
Candelo, C.R. and T.M. Motta. 1996. Perspectivas económicas para el cultivo de caucho. Technical Series No. 37. Corporación Nacional de Investigación and Fomento Forestal (Conif), Bogota. [ Links ]
Chevallier, M.H. 1988. Genetic variability of Hevea brasiliensis germplasm using isozyme markers. J. Nat. Rubber Res. 3, 42-53. [ Links ]
Cornish, K. 2001. Similarities and differences in rubber biochemistry among plant species. Phytochemistry 57, 1123-1134. [ Links ]
Don R., H., P.T. Cox, B.J. Wainwright, K. Baker, and J.S. Mattick. 1991. 'Touchdown' PCR to circumvent spurious priming during gene amplification. Nucleic Acids Res. 19(14), 4008. [ Links ]
Eujayl, I., M.E. Sorrells, M. Baum, P. Wolters, and W. Powell. 2002. Isolation of EST-derived microsatellite markers for genotyping the A and B genomes of wheat. Theor. Appl. Genet. 104, 399-407. [ Links ]
Feng, S.P., W.G. Li, H.S. Huang, Y. Wang, and Y.T. Wu. 2009. Development, characterization and cross-species/genera transferability of EST-SSR markers for rubber tree (Hevea brasiliensis). Mol. Breeding 23, 85-97. [ Links ]
Hernández, C. R., L. Afanador, R.A. Isaza, and A.M. Lobo. 2006. Analysis of genetic variation in clones of rubber (Hevea brasiliensis) from Asian, south and central American origin using RAPD markers. Rev. Colomb. Biotecnol. 8(2), 29-34. [ Links ]
Ilbi, H. 2003. RAPD markers assisted varietal identification and genetic purity test in pepper, Capsicum annuum. Scientia Hort. 97, 211-21 [ Links ]
IRSG, International Rubber Study Group. 2010. Rubber statistical bulletin. Report 125. Singapore, Republic of Singapore. [ Links ]
Le Guen, V., F. Doaré, C. Weber, and M. Seguin. 2009. Genetic structure of Amazonian populations of Hevea brasiliensis is shaped by hydrographical network and isolation by distance. Tree Genet. Genomes 5, 673-683. [ Links ]
Lekawipat, N., M. Teerawatanasuk, S.N. Rodier-Goud, A. Vanavichit, T. Toojinda, and S. Tragoonrung. 2003. Genetic diversity analysis of wild germplasm and cultivated clones of Hevea brasiliensis Muell. Arg. by using microsatellite markers. J. Rubber Res. 6, 36-47. [ Links ]
Lespinasse, D., M. Rodier-Goud, L. Grivet, A. Leconte, H. Legnate, and M. Seguin. 2000. A saturated genetic linkage map of rubber tree (Hevea sp.) based on RFLP, AFLP, microsatellite, and isozyme markers. Theor. Appl. Genet. 100, 127-138 [ Links ]
Lieberei, R. 2007. South-American leaf blight of the rubber tree (Hevea sp.): new steps in plant domestication using physiological features and molecular markers. Ann. Bot. 1, 1-18. [ Links ]
Montoya, J.D. 2009. Determinación de la secuencia de genes putativamente involucrados en la producción de 1, 3 propanodiol en la cepa nativa colombiana Clostridium sp. IBUN 158B. M.Sc. thesis. Faculty of Science, Universidad Nacional de Colombia, Bogota. [ Links ]
Mooibroek, H. and K. Cornish. 2000. Alternative sources of natural rubber. Appl. Microbiol. Biotechnol. 53, 355-365. [ Links ]
Nakkanong, K., C. Nualsri, and S. Sdoodee. 2008. Analysis of genetic diversity in early introduced clones of rubber tree (Hevea brasiliensis) using RAPD and microsatellite markers. Songklanakarin J. Sci. Technol. 30(5), 553-560. [ Links ]
Okogbenin, E., J. Marin, and M. Fregene. 2006. An SSR-based molecular genetic map of cassava. Euphytica 147, 433-440. [ Links ]
Peakall, R. and P.E. Smouse 2006. GENALEX 6: genetic analysis in Excel. Population genetic software for teaching and research. Mol. Ecol. Notes 6, 288-295. [ Links ]
Polhamus, L.G. 1962. Rubber, botany, production, and utilization. Leonard Hill, London. [ Links ]
Prince, J.P., V.K. Lackney, C. Angeles, J.R. Blauth, and M.M. Kyle. 1995. A survey of DNA polymorphism within the genus Capsicum and the fingerprinting of pepper cultivars. Genome 38, 224-231. [ Links ]
Raina, S.N., V. Rani, T. Kojima, Y. Ogihara, K.P. Singh, and R.M. Devarumath. 2001. RAPD and ISSR fingerprints as useful genetic markers for analysis of genetic diversity, varietal identification, and phylogenetic relationships in peanut (Arachis hypogaea) cultivars and wild species. Genome 44, 763-772. [ Links ]
Rajeev, K.V., A. Graner, and M. Sorrells. 2005. Genic microsatellite markers in plants: features and applications. Trends Biotechnol. 23(1), 48-54. [ Links ]
Rohlf, F.J. 2000. NTSYS-pc: numerical taxonomy and multivariate analysis system. Exeter Software, Setauket, NY. [ Links ]
Roux, K.H. 2009. Optimization and troubleshooting in PCR. In: Chen, B.-Y. and H.W. Janes (eds.). Methods in molecular biology. Vol. 192, PCR cloning protocols. 2nd. Edition Humana Press, Totowa, NJ [ Links ]
Rozen, S. and H.J. Skaletsky. 2000. Primer3 on the WWW for general users and for biologist programmers. pp. 365-386. In: Krawetz S. and S. Misener (eds.). Bioinformatics methods and protocols: methods in molecular biology. Humana Press, Totowa, NJ. [ Links ]
Saha, T., B. Roy, M. Ravindran, K. Bini, and M.A. Nacer. 2007. Alltheic diversity revealed through SSR polymorphisms at the locus encoding HMG-coa reductase in rubber (Hevea brasiliensis). Silvae Genetica 56, 58-65. [ Links ]
Saha, T., C.R. Bindu, and M.A. Nazeer. 2005. Microsatellite variability and its use in the characterization of cultivated clones of Hevea brasiliensis. Plant Breeding 124, 86-92. [ Links ]
Semagn, K., Å. Bjørnstad, and M.N. Ndjiondjop. 2006. An overview of molecular marker methods for plants. Afr. J. Biotechnol. 5(25), 2540-2568. [ Links ]
Souza, L. M., C.C. Mantello, M.O. Santos, P.S. Gonçalves, and A.P. Souza. 2009. Microsatellites from rubber tree (Hevea brasiliensis) for genetic diversity analysis and cross-amplification in six Hevea wild species. Conservation Genet. Resources 1, 75-79. [ Links ]
Varghese, Y., A. Knaak, and C. Sethuraj R. 1997. Evaluation of random amplified polymorphic DNA (RAPD) markers in Hevea brasiliensis. Plant Breeding 116, 47-52. [ Links ]
Vera, C.M., M.C. Paredes, and V. Becerra. 1999. Estudio comparativo de diversidad morfológica, isoenzimática y RAPDs dentro y entre clases comerciales de frijol chileno (Phaseolus vulgaris L.). Agric. Téc. 59(4), 247-259. [ Links ]
Yi, G., J. Lee, S. Lee, D. Choi, and B.D. Kim. 2006. Exploitation of pepper EST-SSRs and an SSR-based linkage map. Theor. Appl. Genet. 114, 113-130. [ Links ]