Services on Demand
Journal
Article
Indicators
-
 Cited by SciELO
Cited by SciELO -
 Access statistics
Access statistics
Related links
-
 Cited by Google
Cited by Google -
 Similars in
SciELO
Similars in
SciELO -
 Similars in Google
Similars in Google
Share
Agronomía Colombiana
Print version ISSN 0120-9965
Agron. colomb. vol.30 no.2 Bogotá May/Aug. 2012
PLANT BREEDING, GENETIC RESOURCES & MOLECULAR BIOLOGY
Genetic similarity among commercial oil palm materials based on microsatellite markers
Similaridad genética entre materiales comerciales de palma de aceite basado en marcadores microsatélites
Diana Arias1,Carmenza Montoya1, Leonardo Rey1 and Hernán Romero.1,2
1Biology Program and Breeding Palm, Corporación Centro de Investigación en Palma de Aceite (Cenipalma). Barrancamermeja (Colombia).2Department of Biology, Faculty of Science, Universidad Nacional de Colombia. Bogota (Colombia). hmromeroa@unal.edu.co Received for publication: 29 April, 2012. Accepted for publication: 29 June, 2012.
ABSTRACT Microsatellite markers are used to determine genetic similarities among individuals and might be used in various applications in breeding programs. For example, knowing the genetic similarity relationships of commercial planting materials helps to better understand their responses to environmental, agronomic and plant health factors. This study assessed 17 microsatellite markers in 9 crosses (D x P) of Elaeis guineensis Jacq. from various commercial companies in Malaysia, France, Costa Rica and Colombia, in order to find possible genetic differences and/or similarities. Seventy-seven alleles were obtained, with an average of 4.5 alleles per primer and a range of 2-8 amplified alleles. The results show a significant reduction of alleles, compared to the number of alleles reported for wild oil palm populations. The obtained dendrogram shows the formation of two groups based on their genetic similarity. Group A, with ~76% similarity, contains the commercial material of 3 codes of Deli x La Mé crosses produced in France and Colombia, and group B, with ~66% genetic similarity, includes all the materials produced by commercial companies in Malaysia, France, Costa Rica and Colombia.
Key words: Elaeis guineensis Jacq, breeding programs, alleles, genetic differences.
RESUMEN
Los marcadores moleculares son utilizados para determinar la similaridad genética entre individuos y podrían ser utilizados en diversas aplicaciones en los programas de mejoramiento. El conocer las relaciones de similitud genética de los materiales comerciales ayuda a entender mejor sus respuestas a los factores ambientales, bióticos y abióticos. Este estudio evaluó 17 marcadores de microsatélites en 9 cruces (D x P) de Elaeis guineensis Jacq. de varias empresas comerciales en Malasia, Francia, Costa Rica y Colombia, con el fin de encontrar posibles diferencias genéticas y/o similitudes. Se obtuvieron 77 alelos, con un promedio de 4,5 alelos por cebador y una gama de 2-8 alelos amplificados. Los resultados muestran una reducción significativa de los alelos, en comparación con el número de alelos reportados para las poblaciones silvestres de palma de aceite. El dendrograma obtenido muestra la formación de dos grupos con base a su similitud genética. El grupo A con similitud ~76% contiene material comercial de 3 códigos de cruzamientos Deli x La Mé producido en Francia y Colombia, y el grupo B con ~66% de similitud genética que incluye todos los materiales producidos por empresas comerciales en Malasia, Francia, Costa Rica y Colombia.
Palabras clave: Elaeis guineensis Jacq, programa de mejoramiento, alelos, diferencias genéticas.
Introduction
The African oil palm (Elaeis guineensis Jacq.) is a species native to Africa. It is very important because of the oil extracted from the endosperm of its fruits, which is refined to obtain products of great commercial value, which have multiple uses in the food industry and for biofuels. The oil palm has the highest yields per hectare of all the oil crops. The expansion of oil palm cultivation began to develop in Africa and South-East Asia around the colonial period because of its multiple industrial and food uses (Corley and Tinker, 2003). According to population growth estimates, global consumption of palm oil is expected to reach 37.9 million t by 2020. This represents an average annual growth rate of 3.3% (Fedepalma, 2008). To meet these targets, it is necessary to produce improved, high-yielding genetic material with better oil quality, slower growth and high tolerance to pests and diseases; characteristics that are crucial to commercial plantations for ensuring adequate returns (Bakoumé et al., 2007).
The oil palm is normally monoecious, with male or female flowers, but sometimes both; with inflorescences developing in the axils of the leaves. It is an allogamous species and its diploid genome is comprised of 16 pairs of homologous chromosomes (Singh et al., 2008). In oil palm, there are three types that produce different fruit, depending on the thickness of the shell and its ability to produce oil. The Dura type has fruits with thick shells (2 to 8 mm), occasionally less; accounting for 25-55% of the fruit weight, with 35- 70% pulp and 1-20% kernel. The Pisifera type has shell less fruit and the plants generally exhibit female sterility, the total fruit pulp accounts for 82-99% of the total weight and the kernels 1-18%. The Tenera type is a product of the intra-specific cross between the Dura and pisifera types. Its fruit has a thin shell, 0.5 to 4.0 mm, representing 5-30% of the fruit weight with 3-15 and 60-91% kernel pulp; and constitutes the material grown at the commercial level (Corley and Tinker, 2003).
Oil palm breeding programs are characterized by using the reciprocal recurrent selection scheme which uses two Dura and Pisifera-type starting populations to make the crossings and progeny testing of which the best parents are chosen; and Tenera-type seeds are produced. Subsequently, new starting populations are generated and the cycle is repeated (Corley and Tinker, 2003). History indicates that the oil palm material grown commercially in the world comes from four plants planted in the Bogor Botanical Garden in Java (Indonesia) in 1884. This very narrow genetic base has led to uniformity in the production of the material that is distributed worldwide. Therefore, planted materials share the same female parental stock, Deli dura, with some introgressions, and are combined with three male parental stocks (Pisiferas), the Avros, La Mé and Yagambi (Bakoumé, 2007; Cochard, 2009). This narrow genetic base has driven oil palm breeders to plave greater importance ongenetic resources of the species to increase the genetic variability in breeding programs.
Due to the perennial nature and long generative cycles of oil palm, conventional breeding can take several years, which greatly hampers rapid and efficient progress in the selection of individuals. Various molecular biology techniques are available today for detection of genetic variability and for establishing genetic similarity relationships among individuals and can be used in various applications in breeding programs. For example, microsatellite markers or SS R (Simple Sequence Repeats) have been developed in oil palm since 1999 to carry out studies on genetic diversity (Billotte et al., 2001; Billotte et al., 2005; Bakoumé et al., 2007; Singh et al., 2008), varietal identification (Norziha et al., 2008), pedigree analysis, genome mapping and QTL detection for molecular markerassisted selection (Billotte et al., 2010). SS Rs have great advantages over other markers such as isozymes (Hayati et al., 2004), RAPD (Shah et al., 1994), restriction fragment length polymorphism (RFLP; Maizura et al., 2006) and amplified fragment length polymorphism (AFLP; Kularatne, 2001). SS Rs have the highest Polymorphism Information Content (PIC) and high distribution of loci within the genome (Ferreira and Grattapaglia, 1998). Additionally, Mohammadi and Prasanna (2003) underpin the importance of using microsatellite markers for the following reasons: i) each locus is defined as codominant, so they are ideal for marker-assisted breeding, genetic mapping and diversity measurements; ii) they are highly variable and can distinguish between closely related cultivars and iii) they are highly reproducible through PCR.
Additionally, it is a tool that would allow us to keep track of parents and their offspring. The aim of this study is the molecular characterization of 9 crosses (D x P) of Elaeis guineensis Jacq. from various commercial companies in Malaysia, France, Costa Rica and Colombia; planted in the Palmar de la Vizcaína Research Center (Barrancabermeja, Colombia), to find possible genetic differences and/or similarities among these commercial materials.
Materials and methods
Plant material and DNA isolation
Plants of 189 commercial materials of 9 crosses of oil palm produced by commercial companies in Malaysia, France, Costa Rica and Colombia were used. The origins of the materials used are: Deli Dumpy × Avros; Deli × La Mè; Deli × Avros; Deli × Ghana; Deli × Nigeria; Deli × Yagambi; Deli (Avros × Yagambi); Compact × Ekona and Compact × Ghana. The materials were coded from 1 to 10 within the same cross and a letter of the alphabet was assigned to differentiate between commercial companies. The label of the materials is given by the name of their parents. Thus, the individual Deli × Yagambi A10 represents the individual numbered ten from a Deli × Yagambi cross of commercial company A. These materials are planted in a Cenipalma agronomic trial at the Palmar de la Vizcaína Research Center, Barrancabermeja, Colombia.
Total genomic DNA was obtained from young leaflets. Each leaf sample was macerated with liquid nitrogen and the DNA was isolated using a Qiagen Extraction Kit (Ref: 69106, Gentech, Medellin, Colombia)), following the manufacturers instructions. The DNA was quantified using agarose gel and a spectrophotometer, and finally the samples were diluted to a concentration of 5 ng µL-1.
PCR amplification
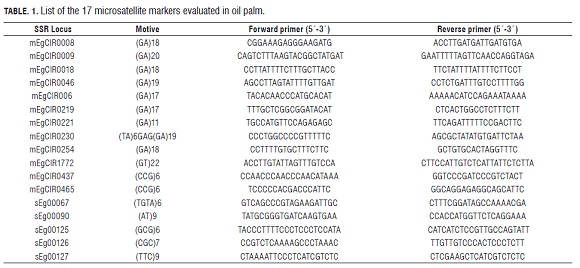
The 189 DNA samples were evaluated using Polymerase Chain Reaction (PCR) with 17 microsatellite markers (Tab.1) according to the amplification conditions reported by Billotte et al. (2001) and Singh et al. (2008). To visualize the product of the amplification, a denaturing polyacrylamide gel stained with silver nitrate was used.
Statistical analysis
The number of amplified alleles was determined for each locus using the amplification patterns reported by Billotte et al. (2001) and Singh et al. (2008) as a reference. The total number of alleles and average polymorphic information content (PIC) were calculated for each locus from the algorithms included in the FSTAT software (Goudet, 2002).
The data were analyzed using the NTSYS pc 2.11L software (Rohlf, 2000) to generate a similarity matrix according to Nei and Li's (1979) definition as follows:
Sij: 2a/ (2a + b + c) (1)
Where Sij is the similarity between individuals i and j; a is the number of bands present in both i and j; b is the number of bands present in i but absent in j; and c is the number of bands present in j but absent in i. From the similarity matrix, a dendrogram was generated using the Unweighted Pair-Group Mean Arithmetic Average method (UPGMA ). Additionally, a principal coordinate analysis was performed, based on which a diagram was generated so that the geometric distances among individuals might reflect genetic distances with minimal distortion. The aggregation patterns in the diagram reveal genetic similarity among individuals (Mohammadi and Prasanna, 2003). To this end, the statistical analysis program NTSYS pc 2.11L (Rohlf, 2000) was used.
To determine allele frequencies, number of alleles per locus and the list of single alleles per cross, the GenAlex ver. 6.0 program (Genetic Analysis in Excel®) was used (Peakall and Smouse, 2006).
Results and discussion
Polymorphism
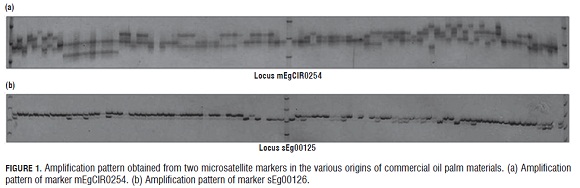
The amplification patterns obtained from the 17 microsatellite markers showed that some markers had more polymorphism within and between genetic origins (Fig.1a), while other primers were polymorphic, but had fewer differences within and between genetic origins (Fig.1b).
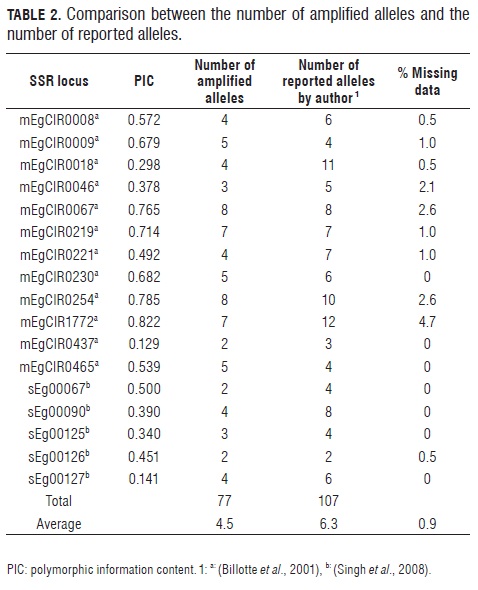
The 17 amplified microsatellite markers were polymorphic, the percentage of missing data was 0.9. A total of 77 alleles were obtained for the 189 assessed samples of oil palm E. guineensis, with an average of 4.5 alleles per primer and a range of 2-8 amplified alleles. The PIC showed a maximum value of 0.822 for locus mEgCIR1772, which suggests that this locus is highly informative and locus mEgCIR0437 was the least informative (PIC = 0.141) (Tab.2). However, only nine markers had a PIC value = 0.5, which means they could be selected for studies on breeding oil palm material.
Results show a significant reduction of alleles as compared to the number of alleles reported by Billotte et al. (2001) in 18 accessions of E. guineensis from germplasm collections of different origins and the number of alleles reported by Singh et al. (2008) in 66 wild E. guineensis from seven countries (Tab.2). The low number of alleles found could be explained by the fact that the materials used as breeding material and the crosses are generally derived from a few individuals, which affects the population and decreases allele variability.
Indeed, different studies show a general tendency of loss of diversity after several cycles of selection. Bakoumè et al. (2007) determined the allelic diversity of 3 breeding materials, using 16 microsatellite markers and found that alleles that were rare in wild oil palm populations showed a reduction due to many years of selection in these materials. Condón et al. (2008) studied the genetic diversity in improved barley varieties with 70 microsatellite markers and found that allelic diversity indices decreased from an average of 5.89 alleles per locus in the most ancestral group to 2.43 alleles per locus after four decades of breeding. Fu et al. (2006) analyzed wheat cultivars released from 1845 to 2004 using 37 EST -derived SS Rs and found a significant reduction in the number of alleles (16 alleles in 14 loci found in pre-cultivars in 1910 were missing in cultivars released after 1990) and a reduction in variation in four ancestral families from 14.7 to 12.8% after six periods of breeding. Therefore, the results of this study show a general trend of loss of genetic diversity caused by the genetic improvement of the oil palm.
Dendrogram and principal coordinates analysis
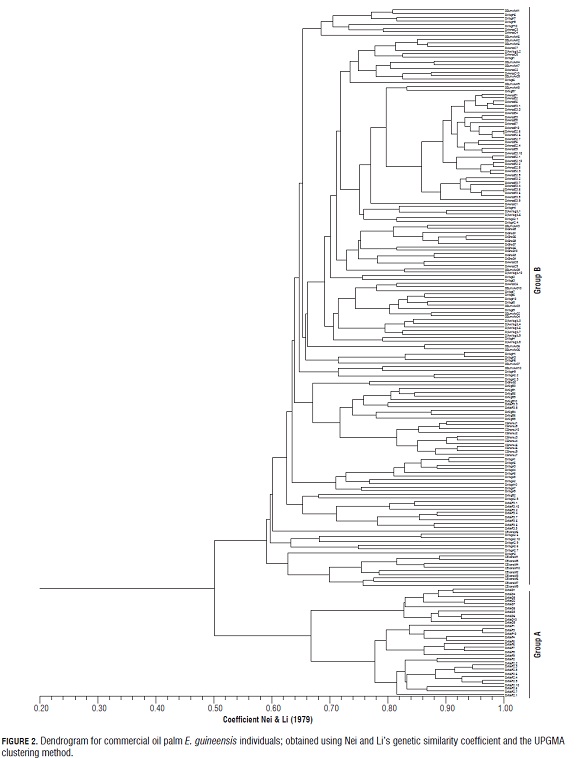
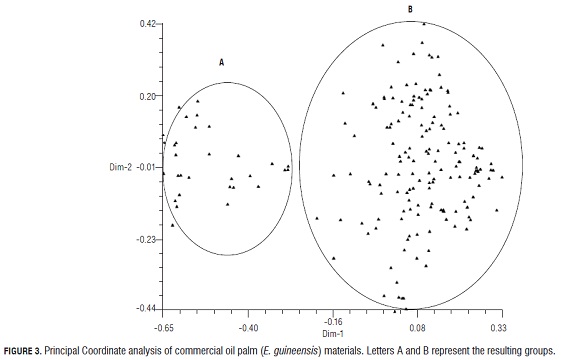
The dendrogram (Fig.2) shows the formation of two groups based on their genetic similarity. Group A, with ~76% similarity, contains commercial materials Deli x La Mé (3 different codes) produced by commercial companies in France and Colombia. These materials have a high genetic similarity between themselves but low similarity with other commercial materials (~50%), reflecting a genetic gain in terms of heterozygosity. It is important to note that there were 7 private alleles in the cross: a single 225-bp allele at locus mEgCIR0008 two single 175-pb and 168-pb alleles at locus mEgCIR0067; a single 200-pb allele at locus mEgCIR0221; a single 137-pb allele at locus mEgCIR0465; a single 152-pb allele at locus sEg00090 and a single 167-pb allele at locus sEg00127. Single alleles are important because they are the raw material of the evolutionary process and are likely to allow a population to respond to a change in the environment. Group B, with ~ 66% of genetic similarity, is the largest one and comprises materials produced by commercial companies in Malaysia, France, Costa Rica and Colombia. This reflects the narrow genetic base of Deli and Avros populations used for these crosses. The high value of genetic similarity between all of the palms analyzed could be explained by different factors such as: a) the fact that they have the same female Deli parent (Deli dura Deli Dumpy and Compact), and different male parents (AVROS, Yagamby, La Mé, Ghana, Nigeria and Ekona), b) the global exchange of plant resources that breeding programs of the various commercial companies have done for many years (Richardson, 1995; Sterling et al., 2002) and c) the limited origin of these populations, which has been described by several authors (Bakoumé, 2007; Dumortier, 2003; Cochard, 2009). The analysis of principal coordinates (Fig.3) also shows the composition of these two groups (A and B). Dimension 1 shows the separation of the genotypes Deli × La Mé (Group A) from the other commercial genotypes: Deli Dumpy × Avros; Deli × Avros; Deli × Ghana; Deli × Nigeria; Deli × Yagambi; Deli (Avros × Yagambi); Compacta × Ekona and Compact × Ghana (Group B). Dimension 2 shows the distribution of each individual within each group. The high degree of observed genetic similarity is consistent with the results reported by different genetic diversity studies of wild E. guineensis, which include Deli dura-type materials as a reference; and it could be concluded that genetically improved populations have less genetic diversity (expressed as heterozygosity values and the percentage of polymorphic loci). According to an analysis using RFLP (Restriction Fragment Length Polymorphism) markers, improved materials are grouped with 80% of genetic similarity (Mayes et al., 2000) and 17% of polymorphic loci, versus 65% in wild populations from Cameroon (Maizura et al., 2001).
The study found a loss of genetic variability in terms of average number of alleles per locus and proportion of polymorphic alleles, but concludes that gene frequency differences among the populations is one of the main factors contributing to the expression of heterozygosity and increased genetic distance among the varieties. The differences between the two groups could be based on the origin and type of progenies. Due to the selection process, different types of alleles have been fixed according to the objectives of each breeding program. For group A, the majority of seed production is based on the selection of Dura-type genotypes of the Deli base as almost their sole dura source, and selection of Pisifera has been performed mainly from La Mè and Yangambi (Corley and Tinker, 2003). This selection is done through the subsequent evaluation of progenies for high oil yield, following the Reciprocal Recurrent Selection (RRS) method (Gascon and Berchoux, 1964). For group B, the selection of Dura-type genotypes has been based on the original Bogor palms, but after four generations of breeding, the best progeny was established, the so called "Ulu Remis" populations from which many breeding programs in the world take the Deli dura breeding population (Corley and Tinker, 2003). The method adopted for seed production was the Family and Individual Selection (FIS ) method using phenotypic selection (Hardon, 1970).
In the seed production of palm oil, the choice of parental lines is fundamental, representing a greater opportunity to recombine genes so that they can maintain and/or maximize the heterozygosity of the new materials, to develop in each cycle. This ensures a way of genetic variability and therefore new allelic combinations within a species. However, several generations of selection and selfing can reduce the number of alleles and increase the coefficient of inbreeding. This circumstance has prompted oil palm breeders to place greater importance on the genetic resources of the species, in order to increase the genetic variability of the base populations through crosses between populations with a wide range of origins, which allows for a rapid selection progress, exploiting the additive genetic variance to the fullest extent and therefore, significant genetic progress in their productive potential. Wild populations of oil palm are found in central and western tropical Africa and on the island of Madagascar or kept in ex situ conditions in various germplasm banks (Corley and Tinker, 2003). Two of the world's most important ex situ collections are found in the MPOB (Malaysian Palm Oil Board) and the NI FOR (Nigeria Institute for Oil Palm Research) with natural populations of oil palm from more than 12 African countries, including Tanzania and Madagascar (Rajanaidu, 1986; Corley and Tinker, 2003).
Conclusion
The results obtained in this research show the conformation of two groups, which reflects the fixation of alleles depending on selection and breeding methods and the base breeding populations used.
While it is true that the breeding programs carried out by commercial seed companies have introduced E. guineensis germplasm with excellent results, there is no doubt that this process must continue, because failure to do so could have negative effects on the required rapid development of new varieties with low vulnerability to adverse environmental factors. A successful breeding program depends on the available genetic variability and on how it is used during the selection process to find high quality palms. The importance of genetic diversity in breeding programs and the knowledge of plant genetic resources are of great social and scientific value for the economic development and food security of humankind. Therefore, it is imperative to provide the greatest genetic diversity possible to breeders and farmers to obtain new cultivars that respond to different soil and weather conditions with enhanced resistance to biotic and abiotic stresses.
Literature cited
Bakoumé, C., R. Wickneswari, N. Raijanaidu, A. Kushairi, P. Amblard, and N. Billotte. 2007. Allelic diversity of natural oil palm (Elaeis guineensis Jacq.) populations detected by microsatellite markers. Implication in conservation. Rev. Palmas 28(1), 149-158. [ Links ]
Billotte, N., A. Rusterucci, E. Barcelos, J. Noyer, P. Amblard, and F. Baurens. 2001. Development, characterization, and acrosstaxa utility of oil palm (Elaeis guineensis Jacq.) microsatellite markers. Genome 44(3), 413-425. [ Links ]
Billotte, N., N. Marseillac, A. Risterucci, B. Adon, P. Brottier, F. Baurens, R. Singh, A. Herrán, A. BillotC, P. Amblard, T. Durand- Gasselin, B. Courtois, D. Asmono, S. Cheah, W. Rohde, E. Ritter, and A. Charrier. 2005. Microsatellite-based high density linkage map in oil palm (Elaeis guineensis Jacq.). Theor. Appl. Genet. 110, 754-765. [ Links ]
Billotte, N., M. Jourjon, N. Marseillac, A. Berger, A. Flori, H. Asmady, B. Adon, R. Singh, B. Nouy, F. Potier, S. Cheah, W. Rohde, E. Ritter, B. Courtois, A. Charrier, and B. Mangin. 2010. QTL detection by multi-parent linkage mapping in oil palm (Elaeis guineensis Jacq.). Theor. Appl. Genet. 120, 1673-1687. [ Links ]
Cochard, B., B. Adon, S. Rekima, N. Billotte, R. Desmier, A. Koutou, B. Nouy, A. Omoré, A. Razak, J. Glazsmann, and J. Noyer. 2009. Geographic and genetic structure of African oil palm diversity suggests new approaches to breeding. Tree Genet. Genomes 5(3), 493-504. [ Links ]
Condón, F., C. Gustus, D. Rasmusson, and K. Smith. 2008. Effect of advanced cycle breeding on genetic diversity in barley breeding germplasm. Crop Sci. 48, 1027-1036. [ Links ]
Corley, R.H.V. and P.B. Tinker. 2003. The oil palm. 4th ed. World Agricultural Series. Blackwell Publishers, Oxford, UK. [ Links ]
Dumortier, F. 2003. Breeding for high yielding progenies at Dami OPRS. pp. 51-74. In: Proc. Agric. Conf. Palm oil: The Power- House for the Global Oils & Fats Economy. Malaysian Palm Oil Board, Kuala Lumpur. [ Links ]
Fedepalma. 2008. Statistical yearbook. Kimpres, Bogota. [ Links ]
Ferreira, M. and D. Grattapaglia. 1998. Introduction to the use of molecular markers in genetic analysis. Embrapa; Cenargen, Brasilia. [ Links ]
Fu, Y.-B., G. Peterson, W. Yu, J. Gao, J. Linfeng, and K. Richards. 2006. Impact of plant breeding on genetic diversity of the Canadian hard red spring wheat germplasm as revealed by EST -derived SS R markers. Theor. Appl. Genet. 112, 1239-1247. [ Links ]
Goudet, J. 2002. Institute of ecology. Biology building, UNI L software (FSTAT ), version 2.9.3.2. In: www.unil.ch/popgen/softwares/fstat.htm consulted: July, 2012. [ Links ]
Hardon, J.J. 1970. Inbreeding in population of the oil palm (Elaeis guineensis Jacq.) and its effects on selection. Oléagineux 25, 449-456. [ Links ]
Hayati, A., R. Wickneswari, I. Maizura, and N. Rajanaidu. 2004. Genetic diversity of oil palm (Elaeis guineensis Jacq.) germplasm collections from Africa: implications from improvement and conservation of genetic resources. Theor. Appl. Genet. 108, 1274-1284. [ Links ]
Kularatne, R.S., F.H. Shah, and N. Rajanaidu, 2001. The evaluation of genetic diversity of Deli dura and African oil palm germplasm collection by AFLP technique. Trop. Agric. Res. 13, 1-12. [ Links ]
Maizura, I., S. C. Cheah, and N. Rajanaidu. 2001. Genetic diversity of oil palm germplasm collections using RFLPs. pp. 526-535. Proc 2001 PIPOC Intl Palm Oil Congr–Cutting-edge technologies for sustained competitiveness (Agriculture). Buenos Aires. [ Links ]
Maizura, I., N. Rajanaidu, A. Zakri, and S. Cheah. 2006. Assessment of Genetic diversity in oil palm (Elaeis guineensis Jacq) using restriction fragment length polymorphism (RFLP). Genet. Resourc. Crop Evol. 53(1), 187-195. [ Links ]
Mayes, S., P. Jack, and H. Corley. 2000. The use of molecular markers to investigate the genetic structure of an oil palm breeding programme. Genet. Soc. Great Brit. Her. 85, 288-293. [ Links ]
Mohammadi, A. and B. Prasanna. 2003. Analysis of genetic diversity in crop plants-salient statistical tools and considerations. Crop Sci. 43(4), 1235-1248. [ Links ]
Nei, M. and W.H. Li. 1979. Mathematical model for studying genetic variation in terms of restriction endonucleases. Proc. Natl. Acad. Sci. 76, 5269-5273. [ Links ]
Norziha, A., M. Rafii, I. Maizura, and S. Ghizan. 2008. Genetic variation among oil palm parent genotypes and their progenies based on microsatellite markers. J. Oil Palm Res. 20, 533-541. [ Links ]
Peakall, R. and P. Smouse. 2006. GenAlEx version 6.0: Genetic Analysis in Excel. Population genetic software for teaching and research. Mol. Ecol. 6, 288-295. [ Links ]
Pinto, R., C. Souza, L. Carlini-Garcia, A. Garcia, and P. Souza. 2003. Comparison between molecular markers and diallel crosses in the assignment of maize lines to heterotic groups. Maydica 48, 63-73. [ Links ]
Rajanaidu, N. 1986. Collection of oil palm (Elaeis guineensis) genetic materials in Tanzania and Madagascar. IS OPB Newsletter 3(4), 2-6. [ Links ]
Rajanaidu, N.A. Kushairi, M. Rafii, M. Din, I. Maizura, and B.S. Jalani. 2000. Oil palm breeding and genetic resources. pp. 171- 227. In: Basiron, Y., B.S. Jalani, K.W. Chan (eds.) Advances in Oil Palm Research. Malaysian Palm Oil Board, Kuala Lumpur. [ Links ]
Richardson, D. 1995. La historia del mejoramiento genético de la palma aceitera en la compañía United Fruit en América. AS D Oil Plam Papers 11, 1-22. [ Links ]
Rohlf, F. 2000. NTSYS pc: Numerical taxonomy and multivariate system, ver. 2.1, computer program, Exeter Publishing, Setauket, New York, NY . [ Links ]
Shah, F.H., O. Rashid, A.J. Simons, and A. Dunsdon, 1994. The utility of RAPD markers for the determination of genetic variation in oil palm (Elaeis guineensis). Theor. Appl. Genet. 89, 713-718. [ Links ]
Singh, R., N. Mohd, N. Ting, R. Rosli, S. Tan, E. Leslie, M. Ithnin, and S. Cheah. 2008. Exploiting an oil palm EST database for the development of gene-derived SS R markers and their exploitation for assessment of genetic diversity Biol Cel. Mol. Biol. 63(2), 227-235. [ Links ]
Sterling, F. and A. Alvarado. 2002. Historia de las colecciones de germoplasma de palma aceitera de AS D de Costa Rica. AS D Oil Palm Papers 24, 17-23. [ Links ]