Services on Demand
Journal
Article
Indicators
-
 Cited by SciELO
Cited by SciELO -
 Access statistics
Access statistics
Related links
-
 Cited by Google
Cited by Google -
 Similars in
SciELO
Similars in
SciELO -
 Similars in Google
Similars in Google
Share
Agronomía Colombiana
Print version ISSN 0120-9965
Agron. colomb. vol.31 no.3 Bogotá Sept./Dec. 2013
PLANT BREEDING, GENETIC RESOURCES & MOLECULAR BIOLOGY
Identification and chromosomal distribution of copia-like retrotransposon sequences in the coffee (Coffea L.) genome
Identificación y distribución cromosómica de secuencias relacionadas con retrotransposones del tipo copia en el genoma del café (Coffea L.)
Juan-Carlos Herrera1 Gloria Camayo1 Gloria De-La-Torre1,Narmer Galeano1,Edgar Salcedo1,Luis Fernando Rivera1,and Andrés Duran1
1Coffee Breeding Program, National Center of Coffee Research (Cenicafe-FNC ). Manizales (Colombia). juanc.herrera@cafedecolombia.comReceived for publication: 17 July, 2013. Accepted for publication: 1 November, 2013.
ABSTRACT
The presence of copia-like transposable elements in seven coffee (Coffea sp.) species, including the cultivated Coffea arabica, was investigated. The highly conserved domains of the reverse transcriptase (RT ) present in the copia retrotransposons were amplified by PCR using degenerated primers. Fragments of roughly 300 bp were obtained and the nucleotide sequence was determined for 36 clones, 19 of which showed good quality. The deduced amino acid sequences were compared by multiple alignment analysis. The data suggested two distinct coffee RT groups, designated as CRTG1 and CRTG2. The sequence identities among the groups ranged from 52 to 60% for CRTG1 and 74 to 85% for CRTG2. The multiple alignment analysis revealed that some of the clones in CRTG1 were closely related to the representative elements present in other plant species such as Brassica napus, Populus ciliata and Picea abis. Furthermore, the chromosomal localization of the RT domains in C. arabica and their putative ancestors was investigated by fluorescence in situ hybridization (FISH) analysis. FISH signals were observed throughout the chromosomes following a similar dispersed pattern with some localized regions exhibiting higher concentrations of those elements, providing new evidence of their relative conservation and stability in the coffee genome.Key words: LTR retrotransposons, reverse transcriptase, repeat sequences, fluorescent in situ hybridization (FISH).
RESUMEN
La presencia de retroelementos del tipo copia fue investigada en siete especies de café (Coffea sp.) incluida la especie cultivada Coffea arabica. El dominio conservado de la enzima transcriptasa reversa (RT ) propia de estos retrotransposones fue amplificada mediante PCR usando cebadores degenerados. Los fragmentos de aproximadamente 300 pb fueron secuenciados, obteniéndose 36 clones, 19 de los cuales fueron de buena calidad. Estos fueron traducidos a su secuencia proteica y comparados entre sí mediante alineamiento múltiple. Los resultados mostraron la presencia de dos grupos definidos: CRTG1 y CRTG2. La identidad de las secuencias al interior de los grupos varió entre 52 y 60% para CRTG1 y entre 74 y 85% para CRTG2. El alineamiento múltiple con secuencias de otras especies reveló alta homología del CRTG1 con secuencias RT presentes en las especies como Brassica napus, Populus ciliata y Picea abis. La hibridación in situ fluorescente (FISH) realizada con el fin de localizar estas secuencias en el genoma de C. arabica y sus dos especies ancestrales, reveló una distribución dispersa a lo largo de los cromosomas, con algunas zonas de mayor concentración. Esta información constituye una nueva evidencia sobre la presencia, conservación y estabilidad de este tipo de retroelementos en el genoma del café.
Palabras clave: retrotransposones LTR , transcriptasa reversa, secuencias repetidas, hibridación in situ fluorescente.
Introduction
A considerable proportion of genomic DNA in plant species is composed of repetitive elements that are consisted of sequence motifs ranging in size from dinucleotides to more than 10,000 base pairs (bp). Depending on their genomic organization and localization on the chromosomes, two major groups of repetitive DNA elements have been recognized. One of these groups includes sequences with tandem repeat units. In this group, we find satellite DNAs, telomeric repeats and rDNA, which are located preferentially at specific positions on the chromosomes, such as the pericentromeric, subtelomeric, telomeric or intercalary regions.The second group of repetitive DNA is dispersed along the chromosomes. These dispersed elements include mobile elements, such as DNA retroelements and transposable elements (TE s). TE s have been grouped by sequence or structural similarity and by the presence or absence of domains or motifs, as well as by transposition mechanisms (Finnegan, 1992; Kumar and Bennetzen, 1999). To date, two main classes have been identified in plants. Class I refers to the retrotransposons or genetic elements that transpose via an RN A intermediate converted into DNA by reverse transcription. The Class II elements are characterized by terminal inverted repeats that flank an open reading frame encoding a transposase enzyme (Bennetzen, 2000; Kidwell, 2002).
Retrotransposons either contain LTR s (long terminal repeats) or do not (non-LTR retrotransposons). The LTR s flank the genes encoding a core protein called gag and a polyprotein called pol. The polyprotein consists of four characteristic domains: protease, integrase, reverse transcriptase (RT ) and ribonuclease H (RN aseH). Because of their evolutive relationship, all RT -dependent mobile genetic elements have also been collectively termed retroelements (Schmidt, 1999; Kumar and Bennetzen, 1999). The differences in genes distinguish the copia and gypsy LTR retroelements types. Non-LTR elements, such as LINES (long interspersed nuclear elements) and SINES (short interspersed nuclear elements), are relatively rare and lack coding activity; therefore, they use the transposase and integration activity from other TE s (Schmidt, 1999). Thanks to similarities among TE s in different plant species, degenerate primers have been widely used to amplify the RT copia-like domains in many eukaryotic species, revealing the existence of multiple families of copia retrotransposons and demonstrating the universal nature of these primers (Matsuoka and Tsunewaki, 1999; Dixit et al., 2006).
The factors that govern TE richness and diversity in a genome are very complex and are likely a combination of properties that are intrinsic to the TE itself as well as extrinsic to the host (Pritham, 2009). It has been shown that TE s remain quiescent during normal growth and development and only become active and proliferate in response to stress adaptation (Melayah et al., 2001; Mirouze and Paszkowski, 2011). Thus, the characterization of the genomic organization and distribution of TE s in the plant kingdom will help further our understanding of the factors that have contributed to plant genome function and remodeling.
Coffee remains one of the principal commodities in the world with a total export value of 22.5 billion US dollars in 2012, equivalent to a volume of 144.6 million bags of 60 kg (ICO, 2012). All coffee species belong to the subgenus Coffea, which contains more than 100 described taxa. Major crop production relies on just two species, Coffea arabica and C. canephora, with the former accounting for over 70% of global coffee production. C. arabica is the only tetraploid (2n=4x=44) among the Coffea species and originated from the ancestral hybridization of two diploid relatives that are close to C. canephora and C. eugenioides (Lashermes et al., 1999).
Although TE s have been broadly studied in different economically important crops such as maize (SanMiguel et al., 1996), tomato (Rogers and Pauls, 2000), rice (Mao et al., 2000), and sunflower (Cavallini et al., 2010), their occurrence and chromosomal localization in coffee are not well characterized. The analysis of non-coding fractions of the C. arabica genome, focusing on repetitive sequences and TE s, would be of particular interest to understanding the global genome evolution of coffee since the formation of the ancestral hybrid between C. eugenioides and C. canephora. In addition, information on TE presence and distribution allows for gaining insight into the possible role of TE s in shaping the C. arabica genome as a result of breeding and selection. Despite their importance, only a few published studies have provided preliminary information on the presence of these elements in the coffee plant genome (Lopes et al., 2008; Hamon et al., 2011; Yuyama et al., 2012). Therefore, the goals of this study were to: (i) investigate the presence of copia-related sequences in the coffee genome and (ii) study their particular chromosomal distribution in the C. arabica genome with respect to its putative ancestral species C. eugenioides and C. canephora.
Materials and methods
Plant material
Seven coffee species, all originated from humid, evergreen, African forests, were selected from the germplasm collection of the Colombian coffee bank. Seed samples from the cultivated alotetraploid (2n=4x=44) C. arabica var. Caturra as well as from the diploid (2n=2x=22) species: C. canephora, C. liberica, C. congensis, C. kapakata, C. stenophylla (all from West and Central Africa) and C. eugenioides (from Central Africa) were investigated. All samples were collected from adult plants established in field conditions at the Naranjal Research Station from - the National Center of Coffee Research (Cenicafe) in Chinchina, Caldas, Colombia.
PCR amplification of RT-derived sequences
Genomic DNA was isolated from young leaves using the procedure described by Herrera et al. (2009). Partial sequences of the RT domain of the copia-like retroelements were amplified using different combinations of the following oligonucleotide primers, which were previously reported by Flavell et al. (1992) and Hirochika et al. (1992): TY1AF (5' ACNGCNTTYYTNCAYGG) encoding the TAFLHG domain, TY1AR (ARCATRTCRTCNCRTA) encoding the YVDDM L domain (in reverse), TY1BF (5'CARATGGARGTN AARAC) encoding the QMD VKT domain and TY1BR (5'CATRTCRTCNACRTA) encoding the YVDDM domain. PCR was performed in 25 mL reaction containing 50 ng of genomic DNA, 25 nM of each primer, 1.25 U Taq DNA polymerase per µL, 1x reaction buffer (Fermentas, Pittsburgh, PA), 200 µM dNTP s, and 2.5 mM MgCl2. The PCR amplification regime was as follows: 94°C for 3 min, 30 cycles at 39°C for 50 s, 72°C for 40 s, 94°C for 1 min and 72°C for 5 min. The obtained products were electrophoresed in 1% agarose gels. Band sizes were confirmed in 5% non-denaturing polyacrylamide gels and visualized by silver staining.
Sequencing and comparative analysis
The PCR products were gel purified and cloned in a pGEM ®-T vector system (Promega, Madison, WI). Colonies that resulted in a PCR -amplified band the size of the expected fragment (around 300 bp) were chosen and purified with thea QIAquick spin kit (Qiagen, Courtaboeuf, France). Three selected fragments per species were sequenced with the single extension method in an automatic 3730xl sequencer (Applied Biosystems, Foster City, CA).
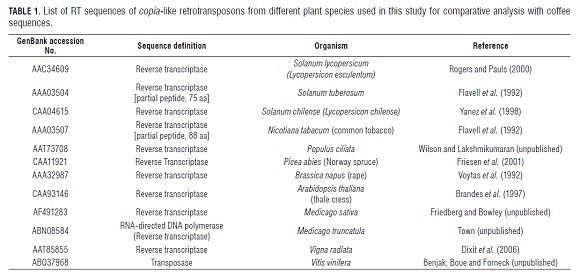
The RT sequenced clones were named with the letters TP followed by the clone number and species name (i.e., CAT for C. arabica var. Caturra, CAN for C. canephora, EUG for C. eugenioides, STE for C. stenophylla, KAP for C. kapakata, CN G for C. congensis, and EUG for C. eugenioides). Nucleotide sequences corresponding to PCR primer annealing sites were eliminated. The resultant RT sequences were translated to the corresponding amino acids with the help of a Perl script using a universal codon dictionary. Each putative amino acid coffee sequence was aligned using the program: ClustalW2. Furthermore, the TP sequences were aligned with the corresponding regions of known RT sequences from a select group of plant species of different taxonomic genera, including the model plants Arabidopsis, Nicotiana and Brassica, as well as more closely related genera, such as Solanum, Vitis and Populus. A multiple cross-analysis was carried out using a Constraint-Based Multiple Alignment Tool, COBALT (constraint-based multiple alignment tool) (Papadopoulos and Agarwala, 2007). Details for these sequences, along with their database accession numbers, organisms and sequence definitions, are provided in Tab.1
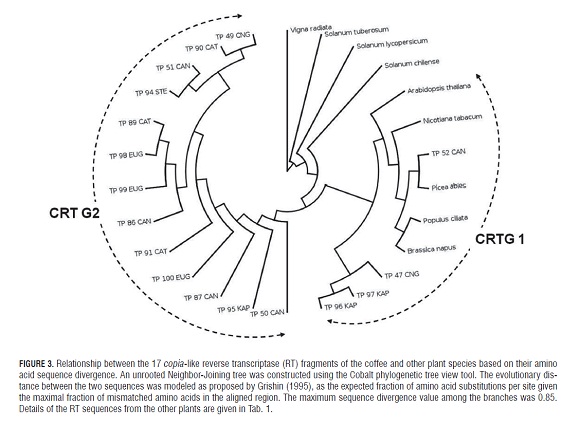
A comparative phylogenetic tree was constructed based on the amino acid sequences of the RT domains using the Neighbor-Joining (NJ) method. The evolutionary distance between two sequences was estimated following Grishin´s model (Grishin, 1995), with the expected fraction of amino acid substitutions per site and considering 0.85 as the maximal fraction of mismatched amino acids in the aligned region. The resulting tree was edited using the PhyloWidgeta tool (Jordan and Piel, 2008). All the nucleotide sequences of the coffee RT -clones reported in this study, corresponding to the copia-like retrotransposons, were published in the NCB I GenBank database under accession numbers JF974034 to JF974052.
Chromosome preparation and in situ hybridization analysis
To investigate the physical location of the copia-like sequences in the C. arabica genome and the related C. canephora and C. eugenioides species, we performed a fluorescence in situ hybridization (FISH, fluorescent in situ hybridization) assay using metaphase chromosomes. Root-tip chromosome preparations and FISH procedures were performed as previously published by Herrera et al. (2007), with a few modifications. Probes were prepared from amplification PCR -products of the RT domain of the copia-like retroelements using C. arabica genomic DNA as a template. In order to reduce the risk of unspecific probe hybridizations, the resulting amplicons were separated in 1% agarose and gel purified using a MinElute® gel extraction kit (Qiagen, Boston, MA). Probe labeling was performed with digoxigenin using a nick translation mix (Roche Molecular Biochemicals, Indianapolis, IN), following the manufacturer's recommendations. The hybridization mixture included 25 ng of labeled probe per slide. The hybridization sites were detected using anti-digoxigenin conjugated to fluorescein (fluorescent antibody enhancer set, Roche Molecular Biochemicals, Indianapolis, IN), which generated green signals. The final slide washes were as follows: 2x SSC , 0.5x SSC and 0.1x SSC at 42°C for 10 min, then 2x SCC at RT for 5 min and 1x PBS (150 mM NaCl, 10 mM NaQHPO4, 10 mM NaH2PO4, pH 7.4) in Tween® 20 (0.2%) at RT for 10 min.
The hybridization stringency and the use of high stringency washes allowed the probe-target combinations with more than 75% homology to remain stably hybridized. Chromosome preparations were counterstained with DAPI (4',6-diamidino-2-phenylindole). The preparations were analyzed using an Eclipse 90i digital microscope (Nikon Instruments, Melville, NY) equipped with a CCD camera. After background subtraction, the individual images from the DAPI and DIG channels were conveniently merged using the Lucia 3.1 software (Nikon, Düsseldorf, Germany).
Results and discussion
Sequence homology among the Copia-like sequences in the coffee
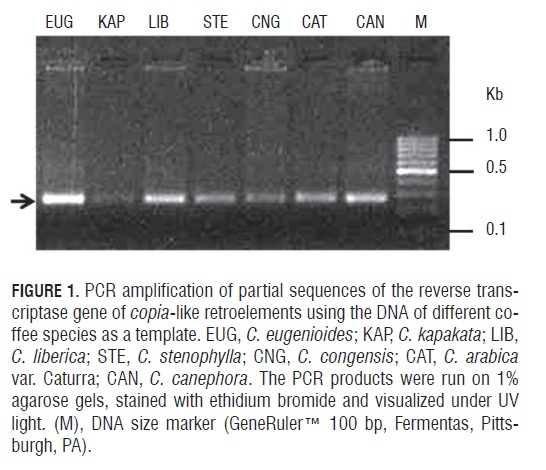
In the present study, we successfully amplified the RT domain of the copia-related elements, one of the principal families of retroelements, in seven coffee species, including the tetraploid C. arabica and six diploids. Using a PCR amplification strategy with degenerate oligonucleotide primers, single fragments of the expected size (300 bp) were obtained from genomic DNA samples (Fig.1). Although the PCR products could represent a mixture of sequences encompassing a heterogeneous pool of RT fragments, we observed that selected degenerated primer combinations allowed for the amplification of a unique band, indicating good specificity.
A total of 36 clones from seven coffee species and derived from clear bands amplified using TY1BF/TY1BR and TY1AF/TY1BR primer pairs combinations were eluted from the gel, purified, cloned and sequenced. After cleaning and quality selection (i.e., length > 250 bp and Phred value > 20), only 19 clones (52.8%) were retained for further comparison. The selected clones represented six of the seven analyzed coffee species (three from C. arabica var. Caturra, five from C. canephora, three from C. eugenioides, three from C. kapakata, three from C. stenophylla and two from C. congensis). Most of the clones (15) were obtained from amplification using TY1AF/TY1BR primer combination.
The multiple alignment analysis of the putative amino acid TP sequences isolated from the coffee is presented in Fig.2. In order to improve local alignment of the entire RT fragments, two TP clones (i.e. TP 92 and TP 93) were excluded from the analysis. The amino acid sequence comparisons allowed for the classification of the 17 remaining sequences into two distinct coffee RT groups, designated as CRTG1 and CRTG2. The amino acid identities between individual sequences belonging to the same group ranged from 52 to 60% for CRTG1 and 74 to 85% for CRTG2. Although not representative of the Coffea genus, such values of predicted amino acid similarities among the isolated RT sequences issued from the seven coffee species investigated seem to be highly heterogeneous.
The consensus sequence SLYGLKQA/SP /SR A/QW, characteristic of Ty1-copia plant elements, appears to be better conserved in CRTG1 rather than in CRTG2 (Fig.2). Also, the presence of properly translated primer sequences at both ends (5' QMD VKT and 3' YVDDM ) was revealed after manual alignments. Overall, the amino acid alignments showed a relative degree of sequence heterogeneity along the inter-domain regions (Fig.2).
Multiple comparisons between the putative amino acid TP sequences isolated from the coffee and RT fragments of other plant species identified from the GenBank database are listed in Tab.1, which allowed for the construction of a NJ-tree (Fig.3).
Most of the coffee RT sequences were placed in the CRTG2 cluster (Fig.3), while the remaining sequences of the Arabidopsis thaliana, Picea abies, Populus ciliata and Brassica napus species clustered into the same group CRTG1 along with the TP sequences from C. kapakata (TP 96 and TP 97), C. congensis (TP 47) and C. canephora (TP 52). As observed in Fig.3, these sequences were grouped separately from homologous sequences of Vigna radiata and Solanum sp.
The closeness among most of the coffee TP sequences is in agreement with the currently accepted hypothesis that all coffee diploids share a common base genome, which is also found in the tetraploid C. arabica (Berthaud and Charrier, 1988); and that coffee plants share an important part of their genome, including the repetitive fraction of LTR elements, with other species of an analogous evolutionary history. Indeed, the Coffea genus belongs to the Rubiaceae family, which is closely related to the tomato and potato (Solanaceae) in the Asterid clade. Other species, such as Arabidopsis, Carica, Medicago, Populus and Vitis, are all members of the Rosid clade that diverged from Asterids 114-125 million years ago (Wilkstrom et al., 2001). Emerging evidence based on colinearity comparisons between orthologous and paralogous regions of Asterids and Rosids has shown that these two clades, including all Coffea species, share the same hexaploid ancestor (Guyot et al., 2009; Cenci et al., 2010).
Previous studies on coffee have suggested that transposable elements, and particularly retrotransposons, are present with relative frequency in this genome. Lopes et al. (2008) for example, identified transposable elements in intergenic regions of expressed sequences from three coffee species (C. arabica, C. canephora and C. racemosa). Most of the TE -containing EST s found in these species (63.7%) were classified as LTR -retrotransposons, indicating the prevalence of this group of TE s in the coffee genome. In a comprehensive annotation analysis of a BAC sequence in C. canephora, Guyot et al. (2009) found that TE s accounted for 7.4 kb (i.e. 4.6%) of the total BAC sequence. Furthermore, these transposable elements appeared to be uniformly distributed along the BAC without a particular pattern of accumulation. Only one putative element was classified as LTR -retrotransposon while the majority belonged to the Class II transposon family, suggesting a strong bias for the presence of these elements as compared to Class I. Similarly, Dereeper et al. (2013) carried out a deep analysis of 131,412 BAC-end sequences (BES s) from two BAC libraries of C. canephora. They found that 11.9% of the total annotated sequences in the coffee genome seemed to correspond to known plant TE s. Interestingly, those reports highlight the extensive macro and micro-syntheny between the C. canephora genome and most of the reference dicotyledonous plants, such as the grapevine (V. vinifera), barrel medic (Medicago truncatula), black cottonwood (Populus trichocarpa) and Arabidopsis thaliana.
Distribution pattern of Copia-elements along the coffee genome
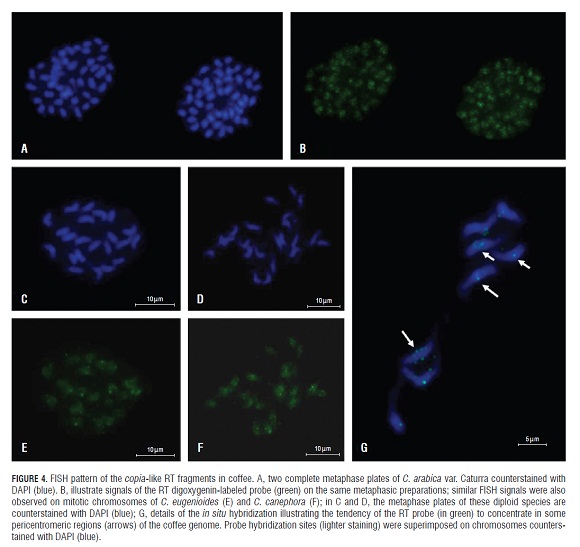
As mentioned in the introduction, the allotetraploid C. arabica genome derives from an ancestral hybridization event between two diploid species related to the current C. canephora and C. eugenioides species (Lashermes et al.,1999). Thus, it would be interesting to compare the distribution of Ty1-copia retroelements in the chromosomes of the tetraploid genome of the arabica relative of these two diploid species. The FISH cytological examination of the general pattern of the C. arabica copia-related fragments used as probes (TP -CAT fragments) revealed very faint signals of hybridization. However, it was evident that the genome regions recognized by the RT probe were scattered along the length of all the C. arabica chromosomes without any obvious preferential localization in specific chromosomes (Fig.4). Although a dispersed pattern was predominant, a detailed analysis revealed some chromosomal regions with higher concentrations of the elements (bright signals), most of them near, although not restricted, to centromeric or peri-centromeric regions (Fig.4G). A similar pattern was observed when a FISH analysis was performed on chromosome preparations from the two putative ancestral species: C. eugenioides and C. canephora (Fig.4E and Fig.4F). Although all chromosomes seem to be concerned, some bright signals were observed along the chromosomes despite the reduced size of the RT probe (around 300 bp). Such hybridization signals exhibiting different intensities along the chromosomes could be interpreted as possible differences in the number of copia-like element copies.
Similar results of FISH assays in the two diploid ancestors of C. arabica obtained in this study could be interpreted as the consequence of minimal changes in the enrichment mechanism of the copia-like sequences during retrotransposon evolution in the C. arabica genome. Therefore, it is conceivable that an overall distribution of copia-retroelements present in the C. arabica genome has remained constant since the ancestral hybridization event and a subsequent whole genome duplication, experienced by the allotetraploid C. arabica, presumably occurred less than 1 million years ago (Lashermes et al., 1999; Cenci et al., 2010).
This pattern of dispersed accumulation of copia-like elements in coffee seems to be in opposition to that observed for the LTR gypsy- elements. Indeed, in a similar study using FISH and a DOP-PCR approach, Yuyama et al. (2012) investigated the presence and chromosomal localization of a gag-like sequence in seven coffee species. Their results highlighted the generalized abundance of this type of retrotransposons in the coffee genome. Nevertheless, contrary to the present report, they observed two different patterns of FISH hybridization among the coffee genomes. In fact, when the gag-like sequences were used as probes, both dispersed and clustered signals were observed. While C. arabica var. typica showed these two hybridization profiles with approximately half of the chromosomes displaying clustered and/or dispersed hybridization signals in the diploid ancestors: C. canephora and C. eugenioides, most chromosomes showed only one pattern, that is, clustered and dispersed signals, respectively. A similar pattern of differential genome localization between Ty3-gypsy and Ty1-copia elements have been reported within the Helianthus genus (Natali et al., 2006).
Transposable elements in all organisms tend to exhibit biases for specific regions within the genome, particularly when they are embedded into repetitive DNA around the centromeres or at the chromosome ends. As observed by Heslop-Harrisson et al. (1997), different retrotransposon sub-families found in any species may behave differently with respect to their mechanisms of amplification and insertion, probably due to variations in the genes belonging to each retroelement family.
The dispersed pattern distribution of the copia-like retroelements on the coffee chromosomes as observed in this report seemed to be consistent with the current knowledge of the mode of replication and insertion of these plant retroelements (Heslop-Harrisson et al., 1997). Owing to several levels and degrees of specificity/bias for insertion or accumulation of plant retrotransposons, it is not surprising to observe local regions with an increased concentration of retroelements such as those reported in this study. These regions may be explained by either preferential selective insertion of retroelements, amplification through replicative mechanisms or an ancient presence in selective regions as a result of amplification and homogenization during evolution (Heslop-Harrisson et al., 1997; Kumar and Bennetzen, 1999).
Studies carried out in different plant species suggest that repeated DNA are mainly consisted of Class I elements, with the LTR elements as the most abundant and the non-LTR types making up a very small segment of plant genomes (Kumar and Bennetzen, 1999; Sidhu and Gill, 2004). In smaller genomes, such as Arabidopsis, retrotransposons make up a very small percentage (approximately 15%) of the genome and are mostly present around the centromeres. In contrast, in larger genomes, such as maize, the repetitive fraction ranges from 64-73% of the genome (Meyers et al., 2001) and the LTR type of retrotransposons seems to transpose preferentially in the gene-poor regions or the regions flanking the gene clusters (Fu et al., 2001). The chromosomal localization of the copia-like elements presented here strongly suggests that the distribution of these LTR transposons in the coffee genome appears to be similar to that of large genomes, where TE s are not concentrated in clusters or repetitive blocks.
Conclusions
Among the LTR elements in plant species, the Ty1-copia superfamily of retrotransposons has been recognized as an abundant component of the angiosperm and gymnosperm genomes (Brandes et al., 1997). In this study, we used a degenerated primer approach to reveal the presence of copialike elements in different coffee species and to investigate their chromosomal distribution in the coffee (C. arabica) genome as well as in the C. eugenioides and C. canephora diploids, considered its ancestral relatives.
The results showed that copia-related sequences were present in the genome of seven coffee species, which included six diploids, suggesting that all of them share this type of repetitive elements as part of their genomes. Furthermore, the amino acid sequence analysis supports the presence of two well differentiated groups of coffee RT -like sequences where the consensus domain SLYGLKQA/SP /SR A/QW, characteristic of Ty1-copia plant elements, was highly variable. The multiple comparison analysis also provided evidence of considerable residue conservation between the amino acid sequences from the coffee and the same RT domain in species such as Brassica napus, Populus ciliata, Picea abis, Nicotiana tabacum and Arabidopsis thaliana. Furthermore, the molecular cytological analysis carried out by FISH revealed no obvious differences between the hybridization patterns of the C. arabica-RT probe when hybridized on the C. arabica chromosomes as well as on their two ancestral relatives: C. eugenioides and C. canephora. Therefore, it could be supposed that copialike retroelements present in the C. arabica genome have remained moderately constant since the recent ancestral hybridization event, allowing for the formation of this cultivated allotetraploid.
Continued sequencing efforts such as those initiated by ICGN (International Coffee Genomic Network) as well as further cytological studies of C. arabica and its diploid relatives will allow for a better understanding of the role of repetitive elements in shaping coffee genomes. Future studies will focus on revealing the organization of retrotransposons and other repetitive elements with respect to the genome size and also on their implications on gene expression under stress conditions during cultivation.
Acknowledgements
The authors would like to thank Jonathan Nuñes, Luisa M. Vasquez and Laura F. Gonzales for their assistance with the molecular analyses. We also appreciate the constructive comments of Alberto Cenci and Romain Guyot during the manuscript preparation. This research was supported by grants from the National Coffee Federation and the Ministerio de Agricultura y Desarrollo Rural (Contract 074/2007). All of the authors contributed substantially to the document and approved the final submission.
Literature cited
Bennetzen, J.L. 2000. Transposable element contributors to plan gene and genome evolution. Plant Mol. Biol. 42, 251-269. [ Links ]
Berthaud, J. and A. Charrier. 1988. Genetic resources of Coffea. pp. 1-42. In: Clarke, R.J. and R. Macrae (eds.). Coffee. Vol. 4: Agronomy. Elsevier Applied Science, London. [ Links ]
Brandes, A., J.S. Heslop-Harrison, A. Kamm, S. Kubis, R.L. Doudrick, and T. Schmidt. 1997. Comparative analysis of the chromosomal and genomic organization of Ty1-copia-like retrotransposons in pteridophytes, gymnosperms and angiosperms. Plant Mol. Biol. 33, 11-21. [ Links ]
Cavallini, A., L. Natali, A. Zuccolo, T. Giordani, I. Jurman, V. Ferrillo, N. Vitacolonna, V. Sarri, F. Cattonaro, M. Ceccarelli, P.G. Cionini, and M. Morgante. 2010. Analysis of transposons and repeat composition of the sunflower (Helianthus annuus L.) genome. Theor. Appl. Genet. 120, 491-508. [ Links ]
Cenci, A., M.C. Combes, and P. Lashermes. 2010. Comparative sequence analyses indicate that Coffea (Asterids) and Vitis (Rosids) derive from the same paleo-hexaploid ancestral genome. Mol. Genet. Genomics 283, 493-501. [ Links ]
Dereeper, A., R. Guyot, C. Tranchant-Dubreuil, F. Anthony, X. Argout, F. de Bellis, M.C. Combes, F. Gavory, A. de Kochko, D. Kudrna, T. Leroy, J. Poulain, M. Rondeau, X. Song, R. Wing, and P. Lashermes. 2013. BAC-end sequences analysis provides first insights into coffee (Coffea canephora P.) genome composition and evolution. Plant Mol. Biol. 83, 177-189. [ Links ]
Dixit, A., K.-H. Ma, J.-W. Yu, E.-G. Cho, and Y.-J. Park. 2006. Reverse transcriptase domain sequences from Mungbean (Vigna radiata) LTR retrotransposons: Sequence characterization and phylogenetic analysis. Plant Cell Rep. 25, 100-111. [ Links ]
Finnegan, D.J. 1992. Transposable elements. Curr. Opin. Genet. Dev. 2, 861-867. [ Links ]
Flavell, A., E. Dunbar, R. Anderson, S.R. Pearce, R. Hartley, and A. Kumar. 1992. Ty1-copia group retrotransposons are ubiquitous and heterogeneous in higher plants. Nucleic Acids Res. 20, 3639-3644. [ Links ]
Friesen, N., A. Brandes, and J.S. Heslop-Harrison. 2001. Diversity, origin, and distribution of retrotransposons (gypsy and copia) in conifers. Mol. Biol. Evol. 18, 1176-1188. [ Links ]
Fu, H., W. Park, X. Yan, Z. Zheng, B. Shen, and H.K. Dooner. 2001. The highly recombinogenic bz locus lies in an unusually generich region of the maize genome. Proc. Natl. Acad. Sci. USA 98, 8903-8908. [ Links ]
Guyot, R., M. De la Mare, V. Viader, P. Hamon, O. Coriton, J. Bustamante, V. Poncet, C. Campa, S. Hamon, and A. De Kochko. 2009. Microcollinearity in an ethylene receptor coding gene region of the Coffea canephora genome is extensively conserved with Vitis vinifera and other distant dicotyledonous sequenced genomes. BMC Plant Biol. 9, 22. [ Links ]
Grishin, N.V. 1995. A more general evolutionary model: substitution rates vary for both amino acids and sites. J. Mol. Evol. 41, 675-79. [ Links ]
Hamon, P., P.-O. Duroy, C. Dubreuil-Tranchant, P.M.A. Costa, C. Duret, N.J. Razafinarivo, E. Couturon, S. Hamon, A. de Kochko, V. Poncet, and R. Guyot. 2011. Two novel Ty1-copia retrotransposons isolated from coffee trees can effectively reveal evolutionary relationships in the Coffea genus (Rubiaceae). Mol. Genet. Genomics 285, 447-460. [ Links ]
Herrera, J.C., A. D'Hont, and P. Lashermes. 2007. Use of fluorescent in situ hybridization as a tool for introgression analysis and chromosome identification in coffee (Coffea arabica L.). Genome 50, 619-626. [ Links ]
Herrera, J.C., G. Alvarado, H. Cortina, M.C. Combes, G. Romero, and P. Lashermes. 2009. Genetic analysis of partial resistance to coffee leaf rust (Hemileia vastatrix Berk and Br.) introgressed into the cultivated Coffea arabica L. from the diploid C. canephora species. Euphytica 167, 57-67. [ Links ]
Heslop-Harrisson, J.S., A. Brandes, S. Taketa, T. Schmidt, A.V. Vershinin, E.G. Alkhimova, A. Kamm, R.L. Doudrick, T. Schwarzacher, A. Katsiotis, S. Kubis, A. Kumar, S.R. Pearce, A.J. Flavell, and G.E. Harrison. 1997. The chromosomal distributions of Ty1-copia group retrotransposable elements in higher plants and their implications for genome evolution. Genetica 100, 197-204. [ Links ]
Hirochika, H., A. Fukuch, and F. Kikuchi. 1992. Retrotransposon families in rice. Mol. Gen. Genet. 233, 209-16. [ Links ]
Jordan, G.E. and W.H. Piel. 2008. PhyloWidget: web-based visualizations for the tree of life. Bioinformatics 24, 1641-1642. [ Links ]
Kidwell, M.G. 2002. Transposable elements and the evolution of genome size in eukaryotes. Genetica 115, 49-63. [ Links ]
Kumar, A. and J.L. Bennetzen. 1999. Plant retrotransposons. Ann. Rev. Genet. 33, 479-532. [ Links ]
Lashermes, P., M.C. Combes, J. Robert, P. Trouslot, A. D'Hont, F. Anthony, and A. Charrier. 1999. Molecular characterization and origin of the Coffea arabica L. genome. Mol. Gen. Genet. 261, 259-266. [ Links ]
Lopes, F.R., M.F. Carazzolle, G.A.G. Pereira, C.A. Colombo, and C.M.A. Carareto. 2008. Transposable elements in Coffea (Gentianales: Rubiacea) transcripts and their role in the origin of protein diversity in flowering plants. Mol. Genet. Gen. 279, 385-401. [ Links ]
Mao, L., T.C. Wood, Y. Yu, M.A. Budiman, J. Tomkins, S. Woo, M. Sasinowski, G. Presting, D. Frisch, S. Goff, R.A. Dean, and R.A. Wing. 2000. Rice transposable elements: A survey of 73,000 sequence tagged connectors. Genome Res. 10, 982-990. [ Links ]
Matsuoka, Y. and K. Tsunewaki. 1999. Evolutionary dynamics of Ty1-copia group retrotransposons in grass shown by reverse transcriptase domain analysis. Mol. Biol. Evol. 16, 208-217. [ Links ]
Melayah, D., E. Bonnivard, B. Chalhoub, C. Audeon, and M.A. Grandbastien. 2001. The mobility of the tobacco TnT1 retrotransposon correlates with its transcriptional activation by fungal factors. Plant J. 28, 159-168. [ Links ]
Meyers, B.C., S.V. Tingey, and M. Morgante. 2001. Abundance, distribution and transcriptional activity of repetitive elements in the maize genome. Genome Res. 11, 1660-1676. [ Links ]
Mirouze, M. and J. Paszkowski. 2011. Epigenetic contribution to stress adaptation in plants. Curr. Opin. Plant Biol. 14, 267-274. [ Links ]
Natali, L., S. Santini, T. Giordani, S. Minelli, P. Maestrini, P.G. Cionini, and A. Cavallini. 2006. Distribution of Ty3-gypsy- and Ty1-copia-like DNA sequences in the genus Helianthus and other Asteraceae. Genome 49, 64-72. [ Links ]
Papadopoulos, J.S. and R. Agarwala. 2007. COBALT: constraintbased alignment tool for multiple protein sequences. Bioinformatics 23, 1073-1079. [ Links ]
Pritham, E.J. 2009. Transposable elements and factors influencing their success in eukaryotes. J. Heredity 100, 648-655. [ Links ]
Rogers, S.A. and K.P. Pauls. 2000. Ty1-copia-like retrotransposons of tomato (Lycopersicon esculentum Mill.). Genome 43, 887-894. [ Links ]
SanMiguel, P., A. Tikhonov, Y.K. Jin, N. Motchoulskaia, D. Zakharov, A. Melake-Berhan, P.S. Springer, K.J. Edwards, M. Lee, Z. Avramova, and J.L. Bennetzen. 1996. Nested retrotransposons in the intergenic regions of the maize genome. Science 274, 765-767. [ Links ]
Schmidt, T. 1999. LINE s, SINE s and repetitive DNA: non-LTR retrotransposons in plant genomes. Plant Mol. Biol. 40, 903-910. [ Links ]
Sidhu, D. and K.S. Gill. 2004. Distribution of genes and recombination in wheat and other eukaryotes. Plant Cell Tiss. Org. Cult. 79, 257-270. [ Links ]
Voytas, D.F., M.P. Cummings, A. Konieczny, F.M. Ausubel, and S.R. Rodermel. 1992. Copia-like retrotransposons are ubiquitous among plants. Proc. Natl. Acad. Sci. USA 89, 7124-7128. [ Links ]
Wilkstrom, N., V. Savolainen, and M.W. Chase. 2001. Evolution of the angiosperms: calibrating the family tree. Proc. Biol. Sci. 268, 2211-2220. [ Links ]
Yanez, M., I. Verdugo, M. Rodriguez, S. Prat, and S. Ruiz-Lara. 1998. Highly heterogeneous families of Ty1-copia retrotransposons in the Lycopersicon chilense genome. Gene 222, 223-228. [ Links ]
Yuyama, P.M., L.F.P. Pereira, T.B. Dos-Santos, T. Sera, L.A. Vilas- Boas, F.R. Lopes, C.M.A. Carareto, and A.L.L. Vanzela. 2012. FISH using a gag-like fragment probe reveals a common Ty3-gypsy-like retrotransposon in genome of Coffea species. Genome 55, 825-833. [ Links ]