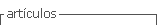Introduction
Tree Tomato (Solanum betaceum Cav.) is a tropical fruit, native of Bolivia; but it is also found in other South American countries such as Colombia and Ecuador (Bohs, 1989). Despite its origin, further breeding of the fruit was done in New Zealand, where it is known as "tamarillo" (Atkinson and Gardner, 1993). Lately, this fruit has lost importance in the latter country, while its commercial cultivation is increasing in Colombia and Ecuador, countries where current research is being generated. Tree tomato is an economically important crop in Ecuador, as it has become a production alternative for medium and small producers. This fruit is in great demand in both local and international markets. The planted area of this crop has increased substantially in Ecuador with around 5,000 ha, located mainly in the highlands of Tungurahua, Chimborazo, Cotopaxi, Pichincha, Carchi, Azuay and Napo Provinces (Viera et al., 2016).
Virosis is a problem which affects fruit quality and also reduces yield. Control and management of viral diseases have routinely been done either by applications of insecticide to control insect vectors like best strategy to avoid virus dispersion, or without any control at all. This disease is caused by viruses producing symptoms such as mosaic, necrotic spots, brown spots, leaf curling, mottling and stunting observed in tree tomato orchards from Ecuador (Sivaprasad et al., 2015; Insuasti et al., 2016; Sivaprasad et al., 2016; Yeturu et al., 2016). In other countries such as Colombia, virosis has been observed since 1968, the symptomatology observed has been concentric rings, violet leaves, spots with tan or oily appearance, chlorosis, leaf and fruit deformation caused mainly by Potyvirus (Gil et al., 2009), being Potato leaf roll virus (PLRV) the most prevalent (Ayala et al., 2010); however it has also been reported Cucumber mosaic virus (CMV) and Alfalfa mosaic virus (AMV), being the latter detected in lowest frequency (Jaramillo et al., 2011); these diagnosis have been done by serological and molecular techniques. In New Zealand, the symptoms observed correspond to mosaic, black spots and spotted fruit related to the occurrence of viruses such as AMV, CMV, Tomato spotted wilt virus (TSWV) and Tamarillo mosaic virus (TaMV), being the latter the most important in this country (Pearson et al., 2006).
Viruses affecting tree tomato orchards in Ecuador have been detected using serological methods such as Elisa test. However, in order to develop effective control programs, confirmation with more accurate methods (molecular techniques) are needed to avoid mistakes in the identification of the causal agent, and consequently in the control strategy to be implemented. For this reason, this study undertook the diagnosis of viruses present in tree tomato plantations from the provinces of Pichincha (Tumbaco) and Tungurahua (Pelileo), using techniques of high specificity and sensitivity such as the polymerase chain reaction (PCR), and the morphological analysis of viral particles by transmission electron microscopy (TEM).
Materials and methods
Sample collection
Leaf tissue with the most common symptoms such as vein yellowing, mosaic, deformation and blistering was collected and immersed in liquid nitrogen for transport and subsequent lyophilization. Sampling was conducted in Pichincha province (three samples), at the INIAP-Experimental Farm of Tumbaco, 00°12'56,5" S and 78°24'37,4" W; altitude of 2,348 m a.s.l., with an average relative humidity of 70.86% and an average temperature of 16°C. In addition, other samples were taken from Tungurahua province in Pelileo (9 samples), 01°17'37.5" S and 78°31'42.9" W; altitude of 2,560 m a.s.l., with an average relative humidity of 78.11% and an average temperature of 15°C. Those two locations represent the highest producing areas of this fruit in Ecuador. Samples were taken from individual plants showing different symptoms concentrated in severely affected orchards in each site.
Polymerase chain reaction with reverse transcription (RT-PCR)
Total RNA extraction was done from 0.05 g of lyophilized plant material by the method of Trisol LS Reagent (Invitrogen catalog number 10296028) and further cleaned with an extra phenol:chloroform and chloroform extraction. The clean RNA was precipitated, dried and re-suspended in 20µL DEPC water. The RNA concentration was measured with a spectrophotometer (NanoVue Plus; Biochrom US, Holliston, MA, USA) and stored at -80°C. Reverse transcription and PCR were performed by two methods: a) with SuperScriptR III Reverse Transcriptase (Invitrogen) followed by PCR with Taq DNA polymerase (Invitrogen), and b) with One-StepRT-PCR System with PlatinumTaq (Invitrogen). In the first method random hexamer primers (Invitrogen) were used to sinthesize cDNA in 20µL reaction including 7µL DEPC water, 1µL of 10 mM dNTPs, 1µL random primers and 4µL of total RNA. The mixture was incubated at 65°C for 5 min and immediately chilled on ice adding 4 µL of 5X buffer, 1µL DTT 0.1 µL M, 1µL DEPC water and 1 of SuperScript® III Reverse Transcriptase. The reaction was incubated in a thermocycler (C1000 TouchTM Thermal Cycler-BioRad; Bio-Rad, Hercules, CA, USA) with one cycle at 25°C/5 min, one cycle 50°C/45 min, and 70°C/15 min. The cDNA obtained was diluted in a 1:6 ratio, to perform a 25 µL PCR reaction that included: 8.39 µL DEPC water, 2.50µL 10x buffer, 1 MgCl2 50 mM, 0.38 µL dNTPs 10 mM, 1.50µL of each primer (10µM), 2.50 µL glycerol, 0.24µL de Taq DNA Polymerase 5U/µL (Invitrogen) and 7µL diluted cDNA. In the second method, specific primers for each virus were used and the SuperScriptIII One Step RT-PCR kit system in a 50µL reaction which included: 21µL DEPC water, 25µL 2X buffer, 1µL of each specific primer (Tab. 1), 1µL enzyme mixture (SuperScriptIII and PlatinumTaq) and 1µL total RNA. The amplification program consisted of an initial first strand cDNA synthesis at 50°C/45min, followed by a PCR program: one cycle 94°C/2 min, 35 cycles: 94°C/1 min, 54-59 (according to the primers)/30 s and 72°C/1 min; a final extension at 72/10 min.
Some primer sequences were obtained from the literature based on reports of virus in this fruit crop and Solanaceae family. Others primers were designed based on sequences obtained from the NCBI and using the software PRIMER3 (Tab. 1). The sequences recovered from NCBI were feed into the PRIMER3 program (http://bioinfo.ut.ee/primer3-0.4.0/primer3/) with the following settings: size between 20 and 30 base pairs, tm between 60 to 70°C, GC% between 40 to 80%. The other parameters were according to the program default. Once the program chose the primers, the best pair of the list were chosen based on their 3' end composition, length, and size of the amplicon.
Primer correspondence with each viral genome was carried out by alignment with the molecular database NCBI using the BLAST program. Positive controls for each virus except TaMV were obtained from infected tissue acquired from Agdia, Inc. The same protocol used in the RNA extraction of infected controls was applied for the collected samples. Primers GADPH Fw5'GGTTACCAGTTCCCGTGTGATTG-3' and GAPDH Rv5'AGACCACTCCAGGCCCACAAAA-3' were used as internal controls of ARN extraction. The PCR products were fractionated by electrophoresis in 2% agarose gels, with 2 µL of SYBR® Safe DNA Gel Stain (Invitrogen) and compared with 100 bp and a 1kb ladders (Invitrogen).
Electron microscopy - Transmission
Electron Microscopy (TEM)
The sap of symptomatic leaf samples was clarified prior to observation in the TEM. A clarification of 0.05 g of lyophilized infected tissue re-suspended in phosphate buffer pH 6.5 was done with chloroform (1:1) using continuous agitation for 10 min, and then centrifuged at 8000 rpm for 20 min at 4oC. The supernatant was transferred to a new tube, mixed with 1% Triton X-100 and kept at 4oC for one hour while stirring. A second clarification was carried out and further precipitated with polyethylene glycol (PEG) in a ratio of 40% w/v and 25µL 5M NaCl, centrifuged at 12,500 rpm for 20 min at 4oC. The pellet was re-suspended in 100µL of phosphate buffer at 4°C over night. The extract was again clarified with chloroform (1:2) and centrifuged for 10 min at 5,000 rpm. The supernatant was transferred to another clean tube and stored at 4°C until its observation in TEM.
TEM observation was performed on copper grids, previously covered with a layer of polivinilformol (formvar) plus a carbon cover. It was stained with a drop of phosphotungstic acid (PTA) (pH 6.8). Viruses were identified by morphological characteristics and sizes known from literature reports (Khan and Dijkstra, 2006), compared with the positive controls and subsequently with the infected samples.
Results
Symptoms in the field
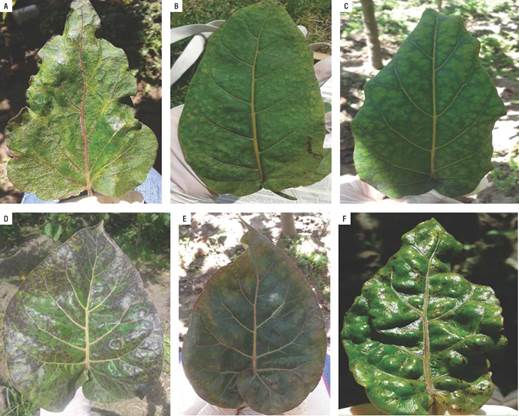
Common symptoms in foliar samples from Tumbaco (Pichincha) were mosaic, yellowing or blistering (Fig. 1).

FIGURE 1 Symptomatology (blistering) observed in samples collected from tree tomato plants in Tumbaco.
In Pelileo (Tungurahua) locality, the tree tomato leaves showed purple coloration symptoms in the upper, accompanied by tan color and a prominent midrib, enlarged and rolled leaves; others leaves presented oily spots symptoms (Fig. 2).
Transmission electron microscopy
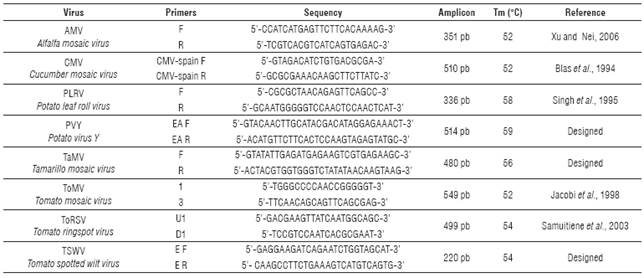
Virus particles were observed in all the positive control preparations and were used to validate the presence of particles in preparations from leaves with symptoms (Fig. 3).
Two of the three virus identified by molecular techniques were photographed with TEM, confirming the findings. Elongated flexible structures were observed at magnifications of 200 nm, with sizes between 650 to 750 nm long and 12 nm wide (Fig. 4 and 5). Symptoms related to these particles presence were foliar blistering, leaf deformation and spots with tan or oily appearance.

FIGURE 4 Viral structure (PVY) in leaf samples collected from tree tomato plants in Tumbaco locality.
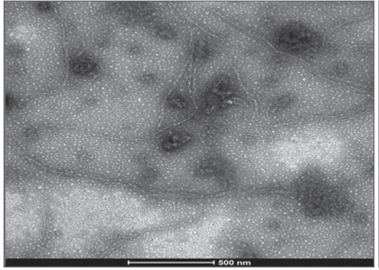
FIGURE 5 Viral structure (PVY) in leaf samples collected from tree tomato plants in Polileo locality.
Rigid rod structures were observed in leaf samples collected in Tumbaco locality. They were 300 nm length and 18 nm width (Fig. 6). Symptom related to this particle presence was foliar blistering.
Reverse transcription polymerase chain reaction (RT-PCR)
Validation of the RNA extraction and RT-PCR techniques using the positive controls of each virus (acquired from Agdia, Inc.) was very useful and efficient because in each case the size corresponding to the right amplicon was obtained.
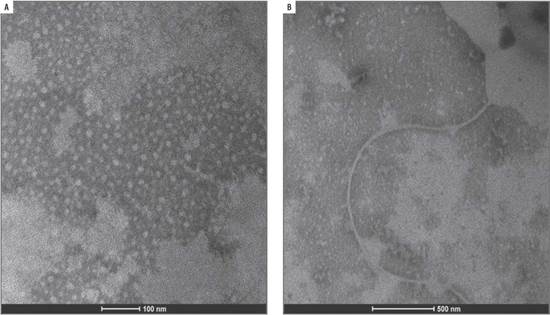
Amplification of positive controls and field samples from total RNA was done by two methods as detailed above. In both cases, the results obtained by the two methods were similar (Fig. 7).
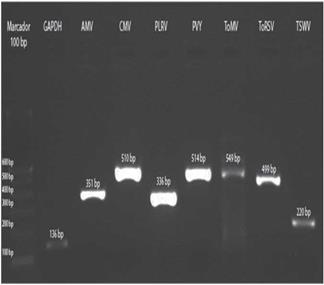
FIGURE 7 Amplification of control samples: GAPDH and the virus AMV, CMV, PLRV, PVY, ToMV, ToRSV and TSWV.
The three samples collected in Tumbaco, amplified with specific primers for PLRV, PVY and ToMV showed fragments of approximately 336 bp, 514 bp and 549 bp, respectively (Fig. 8). While the nine samples from Pelileo, gave similar products for PLRV and PVY but not for ToMV (Fig. 9).
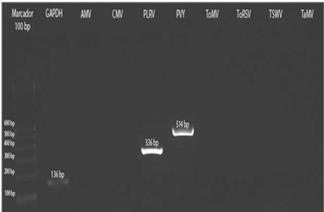
FIGURE 9 RT-PCR from preparations of symptomatic leaf samples of tree tomato from Pelileo localilty.
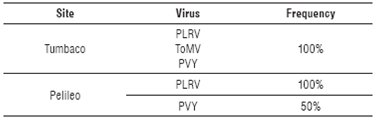
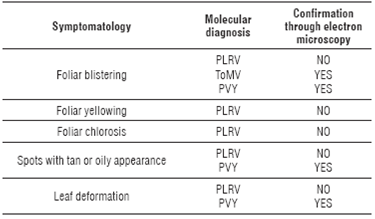
The relationship of the information between the molecular and microscopic diagnosis with the symptomatology observed in the field was confirmed (Tab. 2), with high frequencies of virus detected in each location (Tab. 3).
TABLE 2 Relationship between molecular and transmission electron microscopic diagnosis with symptoms observed in the field in tree tomato plants.
Discussion
There are various problems and diseases that attack the tree tomato crop in Ecuador. Virus infections have gotten strength in recent years and they are present in almost all fields where this fruit is grown in the country. Virus symptoms observed in tree tomato plantations in Tumbaco and Pelileo localities were similar to those reported by Sivaprasad et al. (2015; 2016). However, leaf blistering (common symptom of the three virus detected) with general plant decay was more frequent in Tumbaco samples. While in Pelileo, more varied symptoms such as deformation of leaves, yellowing and leaf chlorosis, spots with tan or oily appearance were observed (symptoms corresponding mainly to PLRV and PVY). In general, these symptoms have been previously reported by Sivaprasad et al. (2015; 2016) in Ecuador, whereas in Colombia, they have been described by Betancourth (2003), Gil et al. (2009), Ayala et al. (2010) and Jaramillo et al. (2011) (corresponding to PLRV, PVY, CMV and AMV). However, in this study, transmission electron microscopy techniques were used complementarily to the molecular analysis for each virus detected.
Transmission electron microscopy
PLRV detection in TEM was difficult because these viruses have an isometric shape and their size is between 25 to 30 nm in diameter (Khan and Dijkstra, 2006); for this reason, this method could be not useful for identification of all types of virus. In this study, the virus could not be photographed in the samples analyzed; however, the positive control showed a size of about 30 nm. According to Llácer et al. (1996), one of the reasons why a virus can-not be observed is due to the low concentrations in raw or semi-purified extracts. These low concentrations may be determined by the tropism of these viruses confined to the phloem of the infected plants. In such cases of vascular limited and low concentration viruses a step of sample concentration or inmuno-electron microscopy is a good method. Such difficulty was compensated by the sensitivity obtained with PCR reaction. Indeed the high percentage of this virus detected by PCR confirmed the importance of having more than one technique in the virus identification. The results obtained for PVY are consistent with the research done by Khan and Dijkstra (2006) who established that this virus has flexuous filaments of12 nm width and 680 to 900 nm length. For this virus, flexuous particles of a size of about 680-730 nm were photographed. In addition, it was determined that the viral concentration in the analyzed samples was low. ToMV was identified by TEM only in samples from Tumbaco (Pichincha), where photographs show structures in form of rigid rods with a length of about 300 nm and a width of 18 nm (Khan and Dijkstra, 2006). In the case of this virus, fragments were observed frequently. This fragmentation could be caused due to pH variations in the extraction buffers (Cann, 2011) and manipulation of the samples or the presence of another segmented helical virus, of a different genus, in addition to ToMV, thus more research will be needed to clarify these observations.
Identification by RT-PCR
Molecular analysis in this research indicated the presence of a viral complex formed by PLRV, PVY and ToMV (100% of samples) in the province of Pichincha (Tumbaco), the first two having been previously reported in this province (Sivaprasad et al., 2015; 2016). In this study, ToMV virus has been detected for the first time infecting tree tomato in Ecuador. Otherwise, in Tungurahua (Pelileo) the viral complex was formed only by PLRV (present in 100% of the samples) and PVY (present in 50% of samples).
In other countries such as Colombia where tree tomato is grown, PLRV virus has been reported by Jaramillo et al. (2011) at a frequency of 41% of the samples analyzed. In our study, this virus was the most frequent (100%), as it was detected in all twelve collected samples. PLRV is transmitted by aphids such as Myzus persicae and Macrosiphum euphorbiae in a persistent and non-propagative way, by seed or by grafting scions (Mowry and Ophus, 2002; Álvarez et al., 2007). Vector activity is crucial in the development of the spread of the virus because the aphid inoculates the virus in the phloem during salivation (Álvarez et al., 2007). Several studies have found that the percentages of virus transmission are variable depending on factors such as the virus genotype, vector, host, environmental conditions, time of infection, latency periods and the agricultural management of seed or seedlings (Johansen et al., 1994; Maule and Wang, 1996; Pasquini and Barba, 2006).
In the case of the Potyvirus PVY, an infection rate of 60% was found on the analyzed samples; while Ayala et al. (2010) reported the presence of this virus in 93% of samples analyzed in Colombian plantations of this fruit. In Ecuador, Insuasti et al. (2016) reported PVY as the most frequent virus; whereas according to the present study, the most frequent virus was PLRV, followed by PVY and less frequently ToMV. According to Ayala et al. (2010), transmission of the genus Potyvirus by seed is not the most common, but it is possible. Seed is considered to be the primary inoculum of the virus. The primary inoculum is the source for transmission by aphids, which are the direct vectors of these viruses by both persistent (survival of the virus in the vector) and non-persistent mechanisms (Johansen et al., 1994; Domier et al., 2007). Apparently, different regions of the HC-Pro and CP proteins are essential for seed and aphid virus transmission (Johansen et al., 1996).
In Tumbaco, the virus ToMV was detected in all analyzed samples. It is known that this virus is prevalent in other Solanaceae, being present in up to 94% of samples (ICTV, 2006). Several factors play an important role in the analysis by RT-PCR of tree tomato tissues. Ayala et al. (2010) and Jaramillo et al. (2011) report the presence of various secondary metabolites that rapidly oxidize tree tomato plant tissue, thus becoming a major problem for this methodology. Due to this drawback, the lyophilization process was included within the RNA extraction method. Lyophilization partially prevented oxidation but the final total RNA was dirty requiring extra washes with phenol, phenol-chloroform and chloroform. Furthermore, the above authors suggest using antioxidants in the extraction method or commercial purification columns of nucleic acid in order to reduce or inactivate enzyme inhibitors.
In the case of the virus AMV, CMV, ToRSV, TSWV and TaMV that did not amplify using the specific primers, it can be stated that the absence of amplification or generation of expected fragments is not a common situation in studies using the PCR technique. This lack of amplification may result from the presence of inhibitors of enzymatic reactions. However, this option is rejected because both the technique and the primers were tested on amplifications of positive controls, thus leading to the conclusion that none of these viruses were present in the samples analyzed from the two locations.
Conclusion
Symptoms such as foliar blistering, yellowing, chlorosis, leaf deformation and spots with tan or oily appearance can be associated with the presence of virus in tree tomato orchards; however, they cannot be related to a specific virus agent without other diagnosis methods like RT-PCR or TEM. The viral disease observed in the tree tomato plantations in the study areas was constituted by a viral complex that includes different taxonomic types of virus such as Potyvirus, Polerovirus and Tobamovirus. A complex of three viruses (PLRV, PVY and ToMV) was identified in leaf samples from Tumbaco locality and two viruses (PLRV and PVY) in the leaf samples of Pelileo locality. The frequency in order of prevalence of the virus detected was PLRV, PVY and ToMV. In the viral complex obtained from the molecular analysis, the presence was observed of at least two of the three viruses identified in the samples analyzed in both localities. TEM observation confirmed the results obtained by RT-PCR; however only two (PVY and ToMV) of the three viruses identified were able to be photographed by TEM. In this study, ToMV virus has been detected for the first time by both RT-PCR and TEM infecting tree tomato in Ecuador.