INTRODUCTION
Coal bed methane (CBM) refers to methane generated by either thermogenic or biogenic processes in coal beds (Moore, 2012). This gas trapped in the coal bed is recovered by using production wells that cut coal beds, allowing the migration of gas from the coal beds to the wells, as is illustrated by Figure 1. The stable carbon (δ13C) and deuterium (δ D) isotopic signatures and gas composition analyses in numerous basins worldwide have shown important microbial CBM occurrence (Strapoc et al., 2011), generating much interest in CBM technology. CBM generation through bio-stimulation and bio-augmentation have been documented as a potential technology for methane production (Jones et al., 2010). Currently, CBM is supplying 6% of the total natural gas consumed in the United States of America (U.S. Energy Information Agency, 2018).
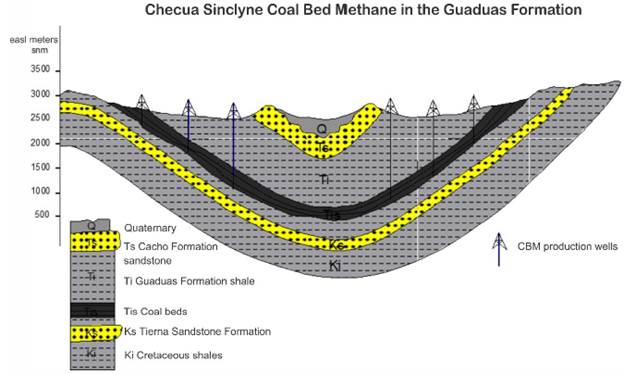
Figure 1 Cross section of the Checua Synclyne in the Bogota Basin Colombia, illustrating the presence of a coal-bearing sequence in the Guaduas formation. This coal sequence presents a potential CBM resource that can be recovered using gas production wells that cut coal beds, allowing the migration of gas from the coal beds to the well.
Analysis of 16S rRNA gene sequences of metagenome samples from coal bed cores or aquifers has enlarged knowledge on the microbial diversity in coal reservoirs throughout world. Coal beds showed a high prokaryotic diversity represented by species of Firmicutes, Spirochetes, Bacteroidetes, and all subgroups of Proteobacteria; as well as methanogens, including Methano-sarcinales, Methanomicrobiales and Methanobacteriales species, which represent all the known methanogenic pathways (Strapoc et al., 2011; Meslé et al., 2013).
Coal methanogenesis is a process involving complex consortia that degrade fossil organic matter present in coal beds. Briefly, hydrolytic and fermentative bacteria hydrolyze complex organic compounds to more simple monomers and oligomers. Then the fermenters, syntrophs and/or acetogens ferment and/or convert these monomers and oligomers mainly to hydrogen (H2), carbon dioxide (CO2) and acetate (Wang et al., 2010). Finally, methanogens produce methane by hydrogen-otrophic ( CO2 reduction), acetoclastic, or methylotrophic methanogenic pathways. Ex situ coal-enrichment cultures studies showed the Methanosarcina, Methanocorpusculum and Methanosaeta species as predominant methanogens and a wide diversity of hydrolytic and fermentative bacteria in the methanogenic consortia (Green et al., 2008; Kruger et al., 2008; Strqpoc et al., 2008; Orem et al., 2010; Penner et al., 2010; Barnhart et al., 2013). Methane production by microbial consortia appears to be influenced by coal micronutrient availability (Ünal et al., 2012), coal rank (Robbins et al., 2016) and coal oxidation state (Gallagher et al., 2013).
Because Colombia has the largest coal reserves in South America, CBM exploitation could contribute significantly to increase methane production in the country. The coal-bearing Guaduas formation of Maastichtian to Paleo-cene age is present in the Bogotá Basin, Eastern Cordillera of Colombia. Stable carbon (δ 13C) and deuterium (δ D) isotopic signatures indicate that methane gas in the Guaduas formation has a mixture of thermogenic and biogenic gases (Garcia-Gonzalez, 2010). Since knowledge on coal mine methanogens is essential for the establishment of CBM technologies, the present work aimed to identify the microbial consortia involved in coal biogenic methanogenesis in the "La Ciscuda" coal mine. Using coal-enrichment cultures, 16S rRNA gene metagenome and gas chromatography (GC) analyses, we identified the methanogenic microbial consortia from this coal mine involved in coal degradation and subsequent gas production.
MATERIALS AND METHODS
Coal sampling
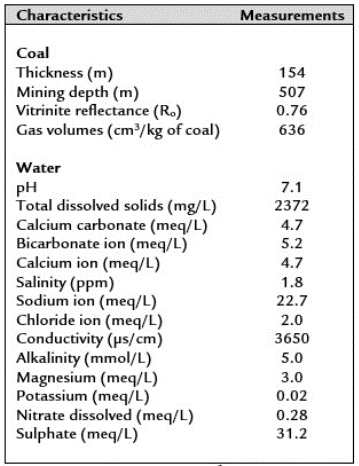
Coal samples were taken from an underground and methane-producing coal mine (La Ciscuda) located in the middle segment (Mantle No. 11, latitude: 5°12'40.08" north; longitude: 73°50'25.60" west) of the Checua-syncline (Figure 1). Underground coal samples were affected by water infiltration from the surface due to their shallow depth (< 200 m deep). The geochemical characteristics of the coal and associated water in La Ciscuda are presented in Table 1.
Coal-enrichment cultures
Cultures inoculated with powdered- coal samples were established using Reinforced Clostridial Medium (RCM) purchased from Oxoid LTD (Basingstoke, England), and gasified per 10 min with CO2 to replace oxygen dissolved in the medium. RCM was used because it allows both growth of anaerobic microbes and provides carbon sources (i.e., dextrose, sodium acetate and soluble starch) and nitrogen sources (beef extract, peptone, and yeast extract) and growth conditions required for methanogenesis, such as osmotic balance (sodium chloride) and low redox potentials (L-cysteine). RCM composition per liter was as follows: beef extract (10 g), peptone (10 g), sodium chloride (5 g), dextrose (5 g), yeast extract (3 g), sodium acetate (3 g), soluble starch (1 g), L-cysteine HCl (0.5 g), agar (0.5 g), pH 6.8 ± 0.2. Gas (CO2) media supplement was purchased from CryoGas Company (Bogotá, Colombia). Briefly, coal portions (0.5 ± 0.2 g) were externally sterilized by immersing in ethanol (70%), dried and pulverized, and then the coal powder was placed in sterile glass canisters containing 20 mL of RCM. The coal-enrichment cultures were grown in triplicate for a month. We always included control assays for non-microbial growth and non-production of biogenic gas, in which the powdered coal samples were placed into sterile glass canisters containing only sterile water.
Gas chromatography analyses
Gas analysis was carried out using the static headspace (S-HS) technique and gas chromatography (GC) coupled to a thermal conductivity detector (TCD) and a flame ionization detector (FID). The GC-TCD-FID analysis was performed in a gas chromatograph AT 7890A (Agilent Technologies, Palo Alto, CA, USA), equipped with TCD and FID. Gas analysis was performed on a HP 7694E static headspace device (Hewlett-Packard, Palo Alto, CA, USA) coupled to the gas chromatograph. The columns used in the analysis were as follows: i) GS-Carbonplot (monolithic carbon, 30 m x 0.53 mm x 3 µm) for H2, O2, N2, CO, CH4 separation; ii) HP-PLOT Molesieve [zeolite (molecular sieve 5 Å), 30 m x 0.53 mm x 50 µm] for CO2, C2H2, C2H4, C2H6, C3H8 separation. A nickel-powder catalytic converter, installed between the TCD and FID, converted CO and CO2 to CH4. FID temperature was maintained at 250 °C. Oven temperature was programmed in the following sequence: from 40 °C (5 min), at 10 °C/min to 100 °C, and then at 10 °C/min to 250 °C. Argon (Linde SA Colombia, Bogotá, Colombia) at a volumetric flow rate of 12 mL/min was used as the carrier gas.
Isolate collection
For microbial isolation, the coal-enrichment cultures were diluted in phosphate buffer supplemented with 1% Triton X-100. Culture dilutions were inoculated (0.1 mL) in glass tubes with fresh CO2-gasified RCM and incubated at either 37 oC or 60 oC, under aerobic and anaerobic conditions. Anaerobic condition was maintained using the Oxoid Atmosphere Generation System and supplements (Oxoid Ltd, Cambridge, UK). For preservation, bacteria were inoculated in glass tubes with semisolid RCM (agar 6 g/L), where the microbial colonies were collected and grown again in fresh RCM. Bacteria isolates were conserved in zeolite (Sigma-Aldrich, St. Louis, USA) with 30% of glycerol at - 80 oC. Bacteria strains and methanogenic consortia were stored in the LMMA-UIS Microbial Collection (http://cepariolmma.uis.edu.co/).
16S rRNA gene metagenome and bacteria isolate amplification
DNA extractions from methanogenic culture and from bacteria isolates were achieved following the methodology proposed by Liu (2009), and their quality and concentration were tested by spectrophotometer. Amplification of the bacteria 16S rRNA gene was performed using the forward 530F (5'-GTCCCAGCMGCCGCGG-3') and reverse 1490R (5'-GGTTACCTTGTTACGACTT-3') universal primers (Wani et al., 2006). In the case of archaea, 16S rRNA gene was amplified using the forward PARCH340f (5'-CCCTACCGGGGYGCASCAG-3') and reverse PREA1100r (5'-YGGGTCTCGCTCGTTRCC-3') primers (Ovreås et al., 1997). Reaction mixture (25 µL) was as follows: 2.5 µl of 10X buffer, 6.2 µL of dNTPs (2 mM), 0.4 µL of each primer (100 µM), 0.4 µL of DreamTaq™ DNA Polymerase (Fermentas, USA), 5 uL of template DNA (5 ng/µL), and 10.1 µL of distilled water. The amplification was carried out using a Thermocycler MasterCycler® Pro-Realplex4 (Eppendorf, Hamburg, Germany). After an initial 3 min denaturation step at 94 oC, 35 PCR cycles were done, each cycle consisting of 45 s at 94 oC, 1 min at 55 oC, and 1 min at 72 oC, ending with an extension at 72 oC for 5 min. PCR products were resolved on a 0.8% agarose gel containing EZ-Vision DNA dye (Amresco, Ohio, USA) and images were recorded using a DigiGenius imaging system (Syngene, Maryland, USA).
Metagenome clone library construction
Using the Clone JET™PCR Cloning Kit" (Thermo Scientific, Massachusetts, USA) or pGEM-T- easy vector (Promega Corp, Wisconsin, USA), we created 16S rRNA gene libraries from each methanogenic culture. PCR products of each 16S rRNA gene were inserted into a pJET1.2/blunt vector and transformed into chemically competent Escherichia coli JM101 cells. Colonies arising on Luria Bertani medium plates (triptone, 10 g, sodium chloride, 10 g, yeast extract, 5 g, pH 7.0) and containing 50 µg/rnL of ampicillin were grown in fresh Luria Bertani (LB) broth and then the plasmids were purified as described by Sambrook and Russell (2001). The archaea PCR products were cloned in pGEM-T-easy vector and transformed into chemically competent E. coli JM109 cells. White colonies arising on LB plates containing 50 mg/mL of ampicillin, IPTG (500 mM) and X-gal (40 mg/mL), were grown in liquid LB broth and then plasmids were purified as described above. The recombinant plasmids were used to amplify rRNA 16S gene clones which were purified with PCR Clean-Up Systems (Promega Corp, Wisconsin, USA) and sequenced with Sanger's method, using the Applied Biosystems Hitachi 3500 Genetic Analyzer (ThermoFisher Scientific, Massachusetts, USA) and manufacturer protocols. Each sample was sequenced at least twice with both forward and reverse primers.
Comparative sequence and phylogenetic analyses
The 16S rDNA partial sequences were first aligned to determine the informative regions and to discard sequence ends with erroneous variability using the BioEdit V7.2.5 software (Hall 1999). The edited sequences were compared with those stored in the National Center for Biotechnology Information (NCBI) database (www.ncbi.nlm.nih.gov). BLAST algorithm (Altschul et al., 1990) was applied for identification of the closest species sequences. Taxon sequence assignments were done using minimum identity values as follows: genera (≥ 95%) and species (≥ 98.7%) (Stackerbrandt and Ebers 2006). Microbial 16S rDNA gene sequences with BLAST query coverage of 100% and an identity value higher than 80% were deposited in GeneBank database with accession numbers as indicated in Table 2. Each operational taxonomic unit (OTU) as defined above was used for phylogenetic tree construction. For comparison, NCBI database related sequences (CP038186, CP042252, CP014793, CP041154, NR1 15692, NR074260, LT996592) were also included. Phylogenetic trees were constructed based on the Tamura-Nei model and the Unweighted Pair Group Method using Arithmetic Averages (UPGMA) method utilizing the Molecular Evolutionary Genetics Analysis (MEGA 5.2) program (Tamura et al., 2011). Bootstrap analysis with 2000 replicates was applied to assign confidence levels to the nodes in the tree.
Table 2 Microbial composition in coal-enriched methanogenic cultures.
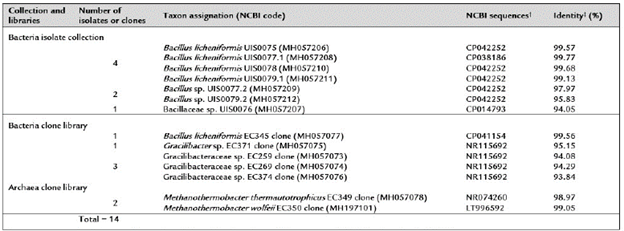
Taxon sequence assignations were done using minimum identity values as follows: genera (¿ 95%) and species 98.7%).
†: The best matching complete genome sequence found in the NCBI database.
‡: Identity refers to the percentage of matches with the aligned NCBI database sequence.
RESULTS
Compared with standard gas profiles (Figure 2a), GC-TCD-FID analysis indicated that, after one-month, cultures at 37 oC (Figure 2b) produced a de novo gas mixture composed mainly of carbon dioxide (CO2), methane (CH4) and carbon monoxide (CO), while cultures at 60 oC (Figure 2c) only produced CO2. As expected, control experiments (coal powder placed in sterile water) did not produce de novo biogenic gas (Figure 2d). These results indicated that a methanogenic consortium obtained from La Ciscuda coal sample was responsible for biogenic gas generation in the cultures.
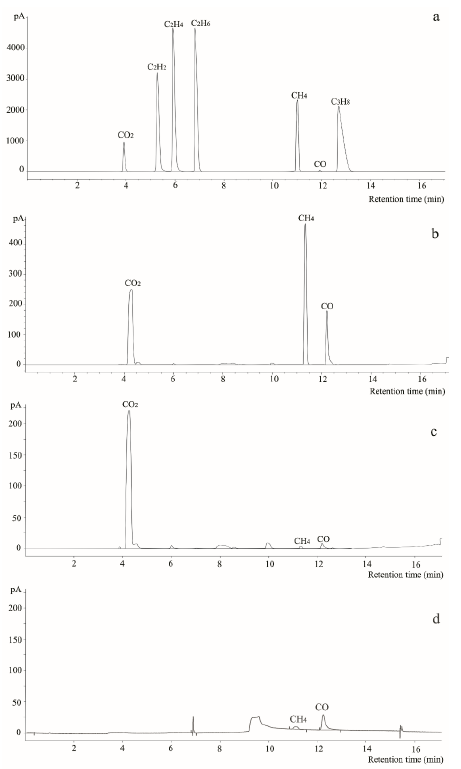
Figure 2 Chromatographic profiles obtained by GC-TCD-FID analysis. a) Standard gas compounds mixture used for comparison; b) Gas mixture produced in the bioreactor with coal-RCM cultures grown at 60 °C; d) Gas mixture recovered from bioreactor with coal poder disolved in only sterile wáter (negative control) instead of RCM.
A total of fourteen 16S rRNA gene sequences were obtained from bacteria isolates (7) and from bacteria libraries (7) developed from cultures (Table 2). BLAST analyses of the isolate sequences showed identity values between 94.05-99.77% with NCBI database Bacillus sequences; four of these (MH057206.1, MH057208.1, MH057210.1 and MH057211.1) matched Bacillus licheniformis sequences with identity values higher than 98.7%. One sequence (MH057077.1) from a bacteria clone library also matched B. licheniformis species sequences with an identity value of 99.56%. Further, BLAST analysis of other bacteria clone library sequences (MH057075.1, MH057073.1, MH057074.1 and MH057076.1) showed identity values (93.84-95.15%) with NCBI database Gracilibacteraceae sequences. One sequence (MH057075.1) matched Gracilibacter thermo-tolerans sequences, the type species of the genus Gracili-bacter (Lee et al., 2006), with identity values higher than 95.0%. Similarly, BLAST analysis of the sequences from the archaea clone libraries (MH057078.1 and MH197101.1) showed high identity values (>98.7%) with NCBI database Methanothermobacter thermautotrophicus (NR074260) and Methanothermobacter wolfeii (LT996592) sequences. In summary, the bacterial isolates and clone libraries obtained from coal-enriched cultures indicated that a minimal methanogenic consortium was formed by specie from two bacteria genera (Bacillus and Gracilibacter) and one archaea genus (Methanothermobacter) species. A UPGMA tree based on all 16S rRNA gene sequences (including type species sequences from the NCBI database) defined the same three main prokaryotic groups (Figure 3).
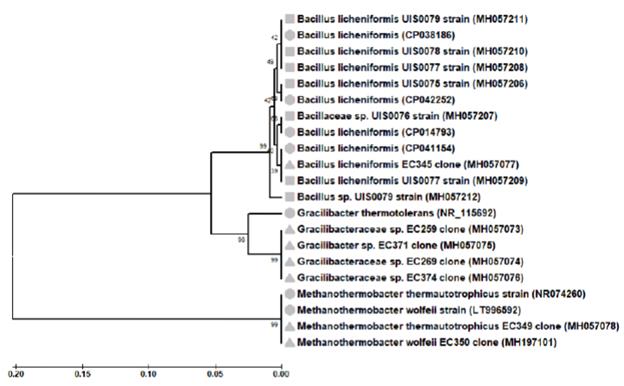
Figure 3 Phylogenetic tree of the strain (C) and clone (p) 16S rRNA gene phylotypes retrieved from coal-enrichment cultures. In parenthesis, the accession number sequences from GeneBank database were given. For comparison, NCBI database related sequences (■▲●) were also included. Alignments were performed with MEGA 5.2 software. The topologies of the tree were obtained with theTamura-Nei model and the UPGMA method. Bootstrap values (n = 2000 replicates) were reported.
DISCUSSION
This work constitutes the first effort to identify the composition of microbial consortia involved in methane production in a coal mine from the Bogotá Basin in Colombia. Our results supported de novo biogenic nature of methane gas produced at the La Ciscuda coal mine as previously indicated using δ13C and δ D isotopic signatures (Garcia-Gonzalez 2010). Further, the study identified a minimal methanogenic consortium that inhabited this coal mine, formed by the bacteria species Bacillus licheniformis and Gracilibacter sp., possibly, G. thermo-tolerans (Lee et al., 2006), and the methanogens Methan-othermobacter thermautotrophicus and M. wolfeii (Wasserfallen et al., 2000). Excepting Gracilibacter, these microbial genera have been previously identified from coal-enrichment cultures experiments (Table 3).
Although methanogens from coal-enrichment cultures were not isolated, they did grow as a methanogenic consortium (Figure 2b). RCM is a very rich medium that provided multiple carbon and nitrogen sources and growth conditions required for methanogenesis such as osmotic balance and low redox potentials. Under these growth conditions, hydrolytic and fermentative bacteria (i.e., B. licheniformis) can enzymatically hydrolyze starch to saccharides such as dextrose (Komolprasert and Ofoli 1991), as well as, can ferment this dextrose through mixed-acid fermentation pathways to organic acids and alcohols (Shariati et al., 1995). Bacillus species, including B. licheniformis, can also solubilize or biodegrade coal lignite into aromatic and aliphatic compounds (Polman et al., 1994). Moreover, G. thermotolerans grows well in medium with similar carbon and nitrogen sources existing in RCM and their growth on media containing glucose produced acetate, lactate, and ethanol as main fermentation end products (Lee et al., 2006). It also has been reported (Sakai et al., 2010) that G. thermotolerans formed a methanogenic consortium with Methanocella arvoryzae, a hydrogenotrophic methanogen isolated from rice field soil. These authors also indicated that G. thermotolerans fermentation products (acetate, H2 and CO2) were required by Methanocella arvoryzae for methane production. We believe that in our study Bacillus and Gracilibacter species, especially the latter, provided substrates (H2 and CO2) to Methanothermobacter species (M. thermautotrophicus and M. wolfeii) for methane production. Gracilibacter thermotolerans cannot grow above 58 oC (Lee et al., 2006), explaining why our methanogenic consortium produced methane at 37 oC, but not at 60 oC.
Table 3 Characterization of methanogenic coal-enriched cultures using 16S rRNA gene analyses.
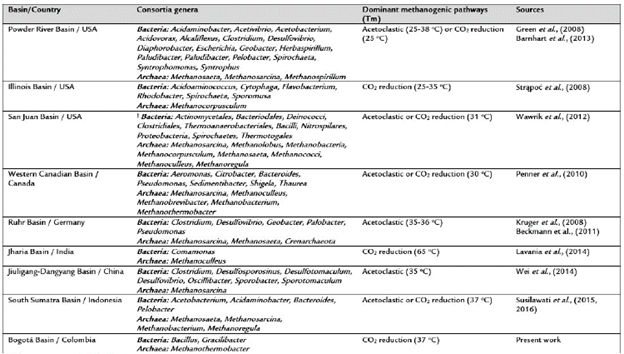
†, The genera were not denned.
Tm, Temperature used for coal-enrichment cultures
Based on our results from coal-enrichment cultures we can speculate on how biogenic methane could be generated in the La Ciscuda coal mine. As indicated in Table 1, this coal mine is located at 200 m depth from surface, where anoxic and saline conditions prevail. Under these conditions the coal mine yields 636 cm3 of methane gas per kg of coal. Parkes et al. (2011) showed that prokaryotes stimulate mineral H2 formation for the deep biosphere and for subsequent microbial activity, including CO2 and CH4 production. We believe that infiltration of meteoric waters into coal mines can stimulate microbial degradation of coal lignite to aromatic and other compounds (Chang et al., 2005), producing H2 and CO2 as final products that are, in this case, the substrates for methanogenesis by Methanothermobacter species. Meth-anothermobacter thermautotrophicus, formerly Methano-bacterium thermoautotrophicum (Smith et al., 1997), and M. wolfei, are representative subsurface methanogen species (Wasserfallen et al., 2000) that previously have been described to produce methane by reduction of CO2 in coal mine methanogenic environments (Ward et al., 2004; Penner et al., 2010).
CONCLUSION
In this work, we identified bacteria (Bacillus and Gracilibacter) and archaea (Methanothermobacter) species forming a minimal methanogenic consortium from La Ciscuda coal mine as a first step for evaluation of CBM generation technologies. Based on this consortium we suggested that methane was produced by hydrogenotrophic or CO2 reduction pathways.