Services on Demand
Journal
Article
Indicators
-
 Cited by SciELO
Cited by SciELO -
 Access statistics
Access statistics
Related links
-
 Cited by Google
Cited by Google -
 Similars in
SciELO
Similars in
SciELO -
 Similars in Google
Similars in Google
Share
Revista Colombiana de Cancerología
Print version ISSN 0123-9015
rev.colomb.cancerol. vol.19 no.4 Bogotá Oct./Dec. 2015
https://doi.org/10.1016/j.rccan.2015.08.002
http://dx.doi.org/10.1016/j.rccan.2015.08.002
REVIEW ARTICLE
Circulating microRNAs as potential cancer biomarkers
MicroRNA circulantes como potenciales biomarcadores en cáncer
Javier Ardila-Molanoa,b, Milena Vizcaínoa,c, Martha Lucía Serranoa,d,*
a Grupo Área de Investigaciones, Instituto Nacional de Cancerología, Bogotá, D.C., Colombia
b Instituto de Genética, Facultad de Medicina, Universidad Nacional de Colombia, Bogotá, D.C., Colombia
c Departamento de Biología, Facultad de Ciencias, Universidad Nacional de Colombia, Bogotá, D.C., Colombia
d Departamento de Química, Facultad de Ciencias, Universidad Nacional de Colombia, Bogotá, D.C., Colombia
* Corresponding author. E-mail address: mlserranol@unal.edu.co (M.L. Serrano).
Received 22 December 2014; accepted 11 August 2015 Available online 17 October 2015
Abstract
Cancer is currently recognized as a major cause of mortality worldwide, and, as such, has a significant impact on public health. Successful early detection strategies are linked to a decrease in mortality; however, because they are invasive, highly complex, and have low specificity, their application has been limited. MicroRNAs (miRNAs) are short, non-coding RNA sequences capable of regulating gene expression. Molecular mechanisms of miRNAs in cancer are not fully understood, but specific patterns of miRNAs expression have been associated with many tumour types. MicroRNAs as biomarkers can help to identify tumour origin and allow their early detection. They can also be used to monitor progression and therapeutic response. The presence and stability of miRNAs in blood, as circulating miRNAs, are major factors that contribute to their use as potential biomarkers in a clinical context. The aim of this article is to present a review of miRNAs as circulating biomarkers in cancer.
Keywords: Circulating microRNA; Early detection; Cancer; Biomarkers; Prognosis.
Resumen
El cáncer es reconocido como una de las principales causas de mortalidad a nivel mundial teniendo así un impacto significativo en la salud pública. Una disminución en la mortalidad por esta enfermedad se ha asociado con estrategias exitosas de detección temprana; sin embargo, debido a que estas son invasivas, altamente complejas y registran baja especificidad, su aplicación ha sido limitada. Los microRNA (miRNA) son RNA no codificantes de secuencias cortas, capaces de regular la expresión génica. Los mecanismos moleculares de los miRNA en cáncer aún no están totalmente clarificados, pero patrones específicos de la expresión de estos han sido asociados con muchos tipos de tumores. Los miRNA como biomarcadores pueden ayudar a identificar el origen de un tumor, realizar detección temprana, predecir la progresión de la enfermedad y la respuesta al tratamiento. La presencia y estabilidad de los miRNA en la sangre, como miRNA circulantes, son los principales factores que contribuyen a su uso potencial como biomarcadores en un contexto clínico. El objetivo de este artículo es realizar una revisión sobre los miRNA circulantes en cáncer.
Palabras clave: MicroRNA circulantes; Detección temprana; Cáncer; Biomarcadores; Pronóstico.
Introduction
Ribonucleic acids (RNA) are molecules that carry out crucial functions in gene expression, with some containing information for protein synthesis (messenger RNA) or are part of the translational machinery (ribosomal and transfer RNA), and others are able to regulate gene expression (microRNAs). MicroRNAs are non-coding RNAs, 18 to 22 nucleotides in length, which participate in gene regulation and are capable of altering the expression of their gene targets.1,2 In 1993, the Ambros group, discovered initial evidence of miRNA in the Caenorhabditis elegans nematode; subsequent studies revealed that miRNA was present in an assortment of organisms, plants and animals.3 MicroRNA genes are embedded in a variety of genome regions, and to date, at least 2500 human miRNAs have been identified.4
MicroRNAs play an important role in the development and various cellular processes, such as differentiation, cell growth, proliferation and cell death.5 They have differential expression profiles in different healthy tissues,6 and their altered expression has been documented in chronic diseases, such as diabetes,7,8 coronary disease,9,10 and cancer.6,11,12
The underlying molecular mechanisms of miRNAs in cancer are not fully understood, but specific patterns of miRNA expression have been associated with many tumour types. MicroRNAs have caught the attention of cancer researchers, mainly due to the fact that they are tissue-specific and deregulate in neoplastic tissue,6 and because of this they can help to identify tumour origin, classify human cancers,13 and also be used as markers of progression14 and future therapeutic targets.15-18
The presence and stability of miRNA in blood are major factors that contribute to being used as tumour diagnostic and progression biomarkers in a clinical context.19,20 In this review, a summary is presented of the main findings related to circulating microRNAs as potential cancer biomarkers.
Overview of miRNA
miRNA biogenesis and regulation of gene expression
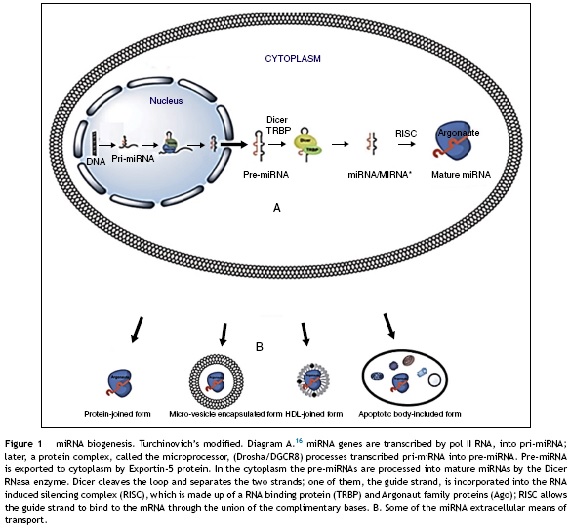
The miRNA biogenesis process is shown in Figure 1. Briefly, miRNA genes produce a pri-miRNA (1-3 kb hairpins), which then turn into a pre-miRNA (70-100 nucleotide hairpins) that are exported to the cytoplasm, where they are converted into mature miRNAs (18-22 nucleotides). Later, the two strands separate and the complementary RNA messenger strand (miRNA), known as the guide strand is incorporated into the RNA induced silencing complex (RISC), which is made up of a RNA binding protein (TRBP) and Argonaut family proteins (Ago).21
Regulation of genetic expression takes place in the RISC-miRNA complex, when the nucleotides 2-8 from the end 5' of miRNA join to nucleotides of end 3' untranslated region of mRNA (3'UTR). Once this union occurs, genetic expression is inhibited in three important ways: I) by reinforcing mRNA degradation, II) by inhibiting mRNA translation, or III) by inducing modification in the chromatin in the target gene, which silences its transcription. If the sequence is totally complementary, mRNA is degraded by direct cleavage, but if the union is partial, mRNA translation is inhibited.1 A single miRNA can influence the expression of hundreds of target genes, and many miRNAs regulate the expression of a single gene.
miRNAs transit from cell to blood stream
MicroRNAs possess mechanisms that preserve their integrity and allow them to transit through the blood stream. These mechanisms include: microvesicles, high and low density lipoproteins, and protein components. The major circulating miRNA transport mechanisms are detailed in Figure 1:
- Microvesicles are small, spherical, membrane-covered bodies of endosome origin, which harbour nucleic acids and proteins. There are different types of microvesicles: I) apoptotic bodies, particles that measure 1-4 µm, II) spill over vesicles (0.1-1 µm) that transport substances to the cell exterior by fusion with the plasma membrane, and III) exosomes, the smallest particles (30-100 nm), that reside within multi-vesicular bodies (MVBs).22,23
- High (HDL) and low (LDL) density lipoproteins are complexes that take part in transporting miRNAs through serum and plasma. Experiments have confirmed that differences exist between miRNA expression patterns in healthy individuals versus those with familial hypercholes-terolemia. MicroRNAs transported by HDL molecules are able to guide miRNAs to host cells and foster their modular genetic effect. These particles can measure from 8 to 12 nm in size.24
- In blood, miRNAs have been linked to different proteins including Argonaut 2 (Ago2), which is an effector of RNA Induced Silencing Complex (RISC).
Circulating miRNAs
In 2010, Weber et al. found that miRNAs are present in biofluids such as: blood, saliva, tears, urine, amniotic fluid, colostrum, breast milk, bronchial secretions, cerebrospinal fluid, peritoneal fluid, pleural fluid, and seminal fluid.19 The stability of circulating miRNAs in biofluids, such as serum and plasma, is a main factor that contributes to their use as potential tumour diagnostic and progression biomarkers in a clinical context.22,23,25 However, in blood it is not easy to separate the levels of truly secreted miRNAs from tumour cells from the circulating cell-free miRNA derived from platelets, erythrocytes, lymphocytes, or normal cell death in blood.
Although many miRNAs have been found in circulation, it remains to be shown if they could functionally down-regulate target mRNAs in target cells.26 Many studies support the idea that exosomal miRNAs can act as regulators of gene expression in distant cells. Exosomes have multiple functions, particularly in cancer, including promotion of local and systemic processes that lead to cell growth and dissemination, or impairment of the immune system response.27 The complete picture of the location and source of miRNAs in the circulation system awaits further clarification.
miRNA analysis
The literature currently reports various analyses based upon two major principles: a search or exploration phase, followed by a validation phase. High performance technologies, such as miRNA sequencing and expression microarrays, have been used to evaluate miRNA profiles in a single assay, starting with small amount of RNA at a relatively low cost and with a high reproducibility.28-34
In the validation phase, specific technology is used for a lower number of selected miRNAs, depending upon the results obtained from previous estimations. Polymerase chain rection (PCR) is usually used to confirm that the findings from a high performance platform are reproducible in a specific miRNA assay.35 Identification of miRNAs with differential expression may help to identify those that are involved in carcinogenesis and could be used as biomarkers. Before the translation of circulating miRNA signatures into a clinical test, all findings require validation and standardisation of all pre-analytical and analytical procedures in order to control for potential technical biases that could have high repercussions on miRNA results, and also limits comparison between different users.30,36,37
Bioinformatics analysis enriches the results obtained from experiments, by supplying information on molecular pathways where miRNAs and their potential targets act. A sampling of databases includes: the Kyoto gene and genome encyclopaedia (KEEG http://www.genome.jp/kegg/), Tar-getScan (http://www.targetscan.org), mirTarget2 (http://www.mirdb.org). The most frequently cited databases on miRNA prediction and targets are: miRanda (http://www.microRNA.org) and miRDB (http://www.mirdb.org). Recently, miRGate(http://mirgate.bioinfo.cnio.es/miRGate/) has been publicly available.38 Oztemur et al39 developed a ranking-based meta-analysis approach using available public databases from miRNA and mRNA microarray studies to discern candidate miRNA biomarkers (meta-miRNAs). This approach identified let-7 family members as breast cancer grade classifiers, and this finding was confirmed by Real Time-PCR studies performed with independent breast tumours.
miRNAs and cancer
The first evidence of involvement of miRNAs in human cancer was when mir-16-1 and/or mir-15a were found down-regulated in 50% to 60% of human chronic lymphocytic leukaemia (CLL) by cytogenetic mechanisms, as deletions and translocations. These two miRNAs activated tumour suppression by inhibiting Bcl-2, an anti-apoptotic effector expression. In the same study, 98 miRNAs were located in fragile, cancer-related, genome regions.40 Since then, several studies have documented the relationship between carcinogenesis and miRNA.41,42
Molecular mechanisms of miRNAs in cancer
Expression of miRNA is deregulated in cancer. Some miRNAs may play a role as an oncogene (oncomiRs), whereas others function as tumour suppressor genes. OncomiRs are overexpressed in cancer, for example, miR-21, mir-17, mir-92a, mir-19b, and mir-106a are oncomiRs responsible for increased proliferation and repression of apoptosis.13 On the contrary, miRNA as tumour suppressor genes are down-regulated in cancer, as miR-18a miR143, miR145 and let-7.43-45 Furthermore, a particular miRNA can exploit both tumour-suppressive and oncogenic functions, depending on the cellular context of its target genes in different cancers.46 In general it has been shown that miRNAs are associated with signalling pathways involved in: I) cell proliferation, as is the case of miR-18a regulation in MAPK pathway genes, II) cell survival pathways, as observed in the case of miR-29a and its regulation in the PI3K1/AKT/MDM2/p53 pathways, associated with cellular apoptosis activity, iii) DNA damage repair pathways, as observed in the case of miR-155 which regulates RAD51 activity, an important protein in DNArepair, following ionic radiation45,47 and iv) cancer cell migration and invasion, as observed in miRNA-29a.10,48-50
miRNAs as diagnostic and prognosis cancer biomarkers
The potential use of miRNAs as biomarkers, beginning with the work of Lu et al. 6 which shows that, with a group of 200 miRNAs it would be possible to identify tumour origin and classify human cancers. In 2008, Schetter et al. 14 identified miRNA expression patterns associated with adenoma to adenocarcinoma progression in colon cancer. Five miRNAs were able to distinguish tumour from non-tumour samples and were linked to poor prognosis. miR-21 expression was low in adenoma and high in stage IV tumours, suggesting that this miRNA probably plays a role as a colorectal tumour progression marker.14 In breast cancer, miRNA profiles were correlated with specific breast cancer features, related to prognosis51-53 and with chemoresistance.54,55 In colorectal cancer, overexpression of miR-21 has been shown to dramatically reduce the therapeutic efficacy of 5-FU and EGFR inhibitor Cetuximab (monoclonal antibody to EGFR) decreases its expression.56
With the rise in the development of research on miRNAs, clinical tests have been developed that provide miRNA profiles, which indicate their possible relationship to cancer. In 2010, Rosenwald et al. 57 developed a platform able to detect primary tumour tissue origin, which can be applied as a clinical test for 25 tumour types. Meiri et al. 11, in 2012, with 64 miRNAs was able to identify 42 tumour types, and identified 52 patients whose primary tumour origin had, until then, been unknown. showing 88% concordance when compared with clinical-pathological evaluations. The above mentioned studies pioneered miRNA diagnostic test development.
Today, commercial companies perform tests using realtime miRNA and PCR microarray technology. Rosetta Genomics® (http://www.rosettagenomics.com) currently offers four miRNA-based tests: I) cancer origin test that evaluates 64 miRNAs able to differentiate 49 cancer origins, designed to help determine the origin for cancer with an unknown primary, II) lung cancer test for the identification of the four main subtypes of lung tumours that significantly influences treatment decisions, III) a kidney cancer test that differentiates three renal cell carcinomas types, and IV) a mesothelioma test that uses microRNA to differentiate malignant pleural mesothelioma from peripheral adenocar-cinomas of the lung, or metastatic carcinomas involving the lung and pleura.
Circulating miRNAs as diagnostic and prognosis cancer biomarkers
In theory, miRNAs in plasma or serum can originate from the tumours or from inflammatory host responses, and their easy access and stability make them ideal for use as biomarkers.58-73 Although in some cancers there is a correlation between specific miRNA profiles in tissue and blood, in other types of cancer a correlation has not been detected. MicroRNA correlation between tissue and blood could depend on the type of tumour, tumour stage, and the specific miRNA type that are being evaluated. Several studies have shown differences between plasma/serum and tissue miRNA profiles in cancer, and miRNAs deregulated in tissue specimens were rarely detected in plasma samples. This suggests that a predictive role of circulating miRNAs is independent from tissue.13,74-77
In 2008, Lawrie et al. found that serum miR-21 and miR-155 have differential expressions between B-cell lymphoma patients and healthy individuals.78 Waters et al. 79 evaluated miRNA expression in a murine model of breast cancer and found low expression levels of miR-195 and miR-497 in serum and tumour tissue three weeks after tumour induction. These results provided evidence of altered circulating miRNA expression levels during tumour development.79 Other studies compared circulating miRNAs with tumour tissue miRNAs; for example, miR-18a in pancreatic cancer shows a good correlation between serum, tissue and tumour cell lines.80 Similarly, Si et al 81 found that miR-21 was overexpressed in serum and tumour tissue in breast cancer patients, but not in controls.
The miRNAs panel in plasma may serve as a potential non-invasive biomarker in detecting the early-stage and prognosis of different cancer types such as gastric,59,82-85 cervical,86 pancreatic,87 colorectal,88-91 lung,74 breast,32,55,92,93 and prostate.83,94-98 These studies need further large-scale validation before circulating miRNAs can be introduced into the clinical setting.
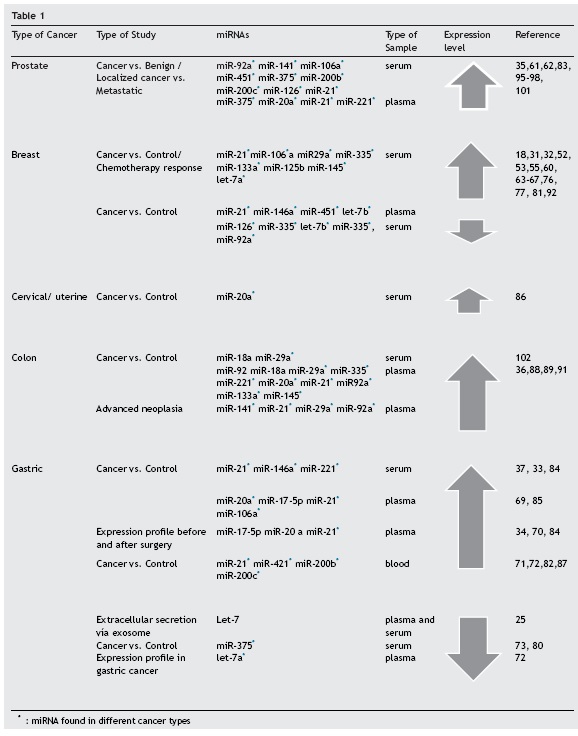
Table 1 shows the circulating miRNAs that modify their expression, in at least two studies, in the five cancers (prostate, breast, cervical, stomach and colon cancer) with the highest incidence and mortality rates in Colombia,99 or miRNAs that show alterations in at least two different cancer types. The review was performed using Medline, Scopus and Science Direct data bases with the search terms "Circulating" + "miRNAs" + i.e. "prostate cancer".
We have decided to only show miRNAs found in at least two different studies, as there are many discordant results among those reviewed. The different ethnic groups, data normalisation methods, and confounding factors in the pre-analytical variables, as well as the entire analytical process may contribute to the discrepancy between research studies on the same disease.26
This review shows there are many circulating miRNAs that have been reported in more than one study for the same cancer type. Such are the cases of miR-141 in prostate cancer, miR-155 in breast cancer, and miR-18a and miR-200c in gastric cancer (Table 1). These miRNAs are of diagnostic importance due to the fact that they have been evaluated and validated using reliable methods.58,75,100-102 Furthermore, there are miRNAs with contradictory results, as some studies found overexpression and others low expression for the same cancer type; for example: miR-16 in prostate cancer,101,103 miR-10b and miR-145 in breast cancer,60,75,101 and let-7 in gastric cancer.25 From a biological point of view it could be possible that they participate in different molecular pathways; however, they cannot be used as a diagnostic biomarker for a specific cancer type.
Additionally, there are more miRNAs that altered their expressions in various cancer types (miR-21, miR-92a, miR-106a, miR-221, miR-375, and miR-29a) (Table 1). miR-21 is overexpressed in prostate, colon, and gastric cancer, and when miR-21 expression is inhibited in glioblastoma cell lines, the activity of caspases 3 and 7 rises, and thus, apo-ptosis, one of the principal molecular signatures in cancer, increases.104 miR-21 is also associated with chemoresis-tance.
The overexpression of miR-106a has been linked to three of the cancers reviewed (prostate, breast, and stomach). At the functional level in gastric cancer, miR106a participates in the negative regulation of TIMP2, an inhibitor of the matrix metalloproteinase (MMPs), which are responsible for degrading extracellular matrix and could play a role in tumour metastasis.41 In many cancer types, these altered miRNAs are not useful as diagnostic biomarkers; but the fact that they alter expression in many cancer types suggest that they play an important role in carcinogenic pathways.
miR-29a is a highly interesting circulating miRNA, due not only to the fact that it has been detected in different types of cancer, but also to its dual role acting as a tumour suppressor gene (osteosarcoma, cervical carcinoma, gastric cancer and lung adenocarcinoma) and an oncogene (breast and colorectal cancer).31,42,48,50,102,105 miR-29a can regulate expression of tumour necrosis factor receptor associated with factor 4 (TRAF4), anti-apoptotic genes, and decreases tumour growth; but in colorectal cancer it promotes metastasis by regulation of E-cadherin and MMP2.50
Circulating miRNA as cancer biomarkers clinical trials
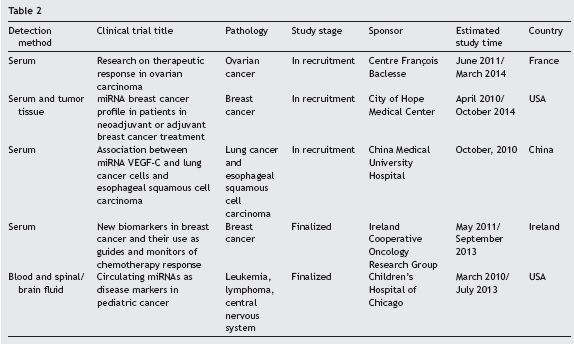
The US Federal Drug Administration (FDA) has approved several clinical trials, which use miRNAs as cancer biomarkers. A sampling of these studies is summarized in Table 2. This table was based upon the clinicaltrials.gov link which summarizes a number of studies in which miRNAs are evaluated for use in diagnosis of specific cancers. These clinical trials included patient cohorts in which differential expression of miRNAs are correlated with tumour stage. The interest in applying these markers in circulating blood is to develop new diagnostic strategies for cancer that will be effective, reproducible and non-invasive for patients.
This document presents a review of evidence on the potential role of circulating miRNAs as non-invasive biomarkers for the early diagnosis of various cancers due to their accessibility and long term stability. Carrying out further studies on circulating miRNA profiles would widen the cancer biomarker research and make it possible to develop diagnostic strategies and examinations using a sensitive and simple peripheral blood test.
References
1. Ambros V. The functions of animal microRNAs. Nature. 2004;431(7006):350-5. [ Links ]
2. Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell. 2004;116(2):281-97. [ Links ]
3. Lee RC, Feinbaum RL, Ambros V. The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell. 1993;75(5):843-54. [ Links ]
4. Kozomara A, Griffiths-Jones S. miRBase: annotating high confidence microRNAs using deep sequencing data. Nucleic Acids Res. 2014;42(Database issue):D68-73. [ Links ]
5. Bartel DP, Chen CZ. Micromanagers of gene expression: the potentially widespread influence of metazoan microRNAs. Nat Rev Genet. 2004;5(5):396-400. [ Links ]
6. Lu J, Getz G, Miska EA, Alvarez-Saavedra E, Lamb J, Peck D, et al. MicroRNA expression profiles classify human cancers. Nature. 2005;435(7043):834-8. [ Links ]
7. El Ouaamari A, Baroukh N, Martens GA, Lebrun P, Pipeleers D, van Obberghen E. miR-375 targets 3'-phosphoinositidedependent protein kinase-1 and regulates glucose-induced biological responses in pancreatic beta-cells. Diabetes. 2008;57(10):2708-17. [ Links ]
8. Wang X, Sundquist J, Zöller B, Memon AA, Palmér K, Sundquist K, et al. Determination of 14 Circulating microRNAs in Swedes and Iraqis with and without Diabetes Mellitus Type 2. PLoS One. 2014;9(1):e86792. [ Links ]
9. Sun X, Zhang M, Sanagawa A, Mori C, Ito S, Iwaki S, et al. Circulating microRNA-126 in patients with coronary artery disease: correlation with LDL cholesterol. Thromb J. 2012;10(1):16. [ Links ]
10. Xiao J, Shen B, Li J, Lv D, Zhao Y, Wang F, et al. Serum microRNA-499 and microRNA-208a as biomarkers of acute myocardial infarction. Int J Clin Exp Med. 2014;7(1): 136-41. [ Links ]
11. Meiri E, Mueller WC, Rosenwald S, Zepeniuk M, Klinke E, Edmonston TB, et al. A second-generation microRNAbased assay for diagnosing tumor tissue origin. Oncologist. 2012;17(6):801-12. [ Links ]
12. Gilad S, Meiri E, Yogev Y, Benjamin S, Lebanony D, Yerushalmi N, et al. Serum microRNAs are promising novel biomarkers. PLoS One. 2008;3(9):e3148. [ Links ]
13. Boeri M, Verri C, Conte D, Roz L, Modena P, Facchinetti F, et al. MicroRNA signatures in tissues and plasma predict development and prognosis of computed tomography detected lung cancer. Proc Natl Acad Sci U S A. 2011;108(9): 3713-8. [ Links ]
14. Schetter AJ, Leung SY, Sohn JJ, Zanetti KA, Bowman ED, Yanaihara N, et al. MicroRNA expression profiles associated with prognosis and therapeutic outcome in colon adenocarcinoma. JAMA. 2008;299(4):425-36. [ Links ]
15. Schwarzenbach H, Nishida N, Calin GA, Pantel K. Clinical relevance of circulating cell-free microRNAs in cancer. Nat Rev Clin Oncol. 2014;11(3):145-56. [ Links ]
16. Cheng CJ, Bahal R, Babar IA, Pincus Z, Barrera F, Liu C, et al. MicroRNA silencing for cancer therapy targeted to the tumour microenvironment. Nature. 2015;518(7537):107-10. [ Links ]
17. Ling H, Fabbri M, Calin GA. MicroRNAs and other non-coding RNAs as targets for anticancer drug development. Nat Rev Drug Discov. 2013;12(11):847-65. [ Links ]
18. Cho WC. MicroRNAs: potential biomarkers for cancer diagnosis, prognosis and targets for therapy. Int J Biochem Cell Biol. 2010;42(8):1273-81. [ Links ]
19. Weber JA, Baxter DH, Zhang S, Huang DY, Huang KH, Lee MJ, et al. The microRNA spectrum in 12 body fluids. Clin Chem. 2010;56(11):1733-41. [ Links ]
20. Mostert B, Sieuwerts AM, Martens JW, Sleijfer S. Diagnostic applications of cell-free and circulating tumor cell-associated miRNAs in cancer patients. Expert Rev Mol Diagn. 2011;11(3):259-75. [ Links ]
21. Turchinovich A, Samatov TR, Tonevitsky AG, Burwinkel B. Circulating miRNAs: cell-cell communication function? Front Genet. 2013;4:119. [ Links ]
22. Hessvik NP, Sandvig K, Llorente A. Exosomal miRNAs as Biomarkers for Prostate Cancer. Front Genet. 2013;4:36. [ Links ]
23. Turchinovich A, Weiz L, Burwinkel B. Extracellular miRNAs: the mystery of their origin and function. Trends Biochem Sci. 2012;37(11):460-5. [ Links ]
24. Vickers KC, Palmisano BT, Shoucri BM, Shamburek RD, Remaley AT. MicroRNAs are transported in plasma and delivered to recipient cells by high-density lipoproteins. Nat Cell Biol. 2011;13(4):423-33. [ Links ]
25. Ohshima K, Inoue K, Fujiwara A, Hatakeyama K, Kanto K, Watanabe Y, et al. Let-7 microRNA family is selectively secreted into the extracellular environment via exosomes in a metastatic gastric cancer cell line. PLoS One. 2010;5(10):e13247. [ Links ]
26. Margue C, Reinsbach S, Philippidou D, Beaume N, Walters C, Schneider JG, et al. Comparison of a healthy miRNome with melanoma patient miRNomes: are microRNAs suitable serum biomarkers for cancer? Oncotarget. 2015. [ Links ]
27. Falcone G, Felsani A, D'Agnano I. Signaling by exosomal microRNAs in cancer. J Exp Clin Cancer Res. 2015;34(1):32. [ Links ]
28. Huang X, Yuan T, Tschannen M, Sun Z, Jacob H, Du M, et al. Characterization of human plasma-derived exosomal RNAs by deep sequencing. BMC Genomics. 2013;14:319. [ Links ]
29. Williams Z, Ben-Dov IZ, Elias R, Mihailovic A, Brown M, Rosenwaks Z, et al. Comprehensive profiling of circulating microRNA via small RNA sequencing of cDNA libraries reveals biomarker potential and limitations. Proc Natl Acad Sci U S A. 2013;110(11):4255-60. [ Links ]
30. Tiberio P, Callari M, Angeloni V, Daidone MG, Appierto V. Challenges in Using Circulating miRNAs as Cancer Biomarkers. Biomed Res Int. 2015;2015:731479. [ Links ]
31. Wu Q, Lu Z, Li H, Lu J, Guo L, Ge Q. Next-generation sequencing of microRNAs for breast cancer detection. J Biomed Biotechnol. 2011;2011:597145. [ Links ]
32. Wu X, Somlo G, Yu Y, Palomares MR, Li AX, Zhou W, et al. De novo sequencing of circulating miRNAs identifies novel markers predicting clinical outcome of locally advanced breast cancer. J Transl Med. 2012;10:42. [ Links ]
33. Liu H, Zhu L, Liu B, Yang L, Meng X, Zhang W, et al. Genome-wide microRNA profiles identify miR-378 as a serum biomarker for early detection of gastric cancer. Cancer Lett. 2012;316(2):196-203. [ Links ]
34. Liu N, Liu X, Zhou N, Wu Q, Zhou L, Li Q. Gene expression profiling and bioinformatics analysis of gastric carcinoma. Exp Mol Pathol. 2014. [ Links ]
35. Moltzahn F, Olshen AB, Baehner L, Peek A, Fong L, Stöppler H, et al. Microfluidic-based multiplex qRT-PCR identifies diagnostic and prognostic microRNA signatures in the sera of prostate cancer patients. Cancer Res. 2011;71(2):550-60. [ Links ]
36. Giráldez MD, Lozano JJ, Ramírez G, Hijona E, Bujanda L, Castells A, et al. Circulating microRNAs as biomarkers of colorectal cancer: results from a genome-wide profiling and validation study. Clin Gastroenterol Hepatol. 2013;11(6):681-8, e3. [ Links ]
37. Kim SY, Jeon TY, Choi CI, Kim DH, Kim GH, Ryu DY, et al. Validation of circulating miRNA biomarkers for predicting lymph node metastasis in gastric cancer. J Mol Diagn. 2013;15(5):661-9. [ Links ]
38. Andrés-León E, González Peña D, Gómez-López G, Pisano DG. miRGate: a curated database of human, mouse and rat miRNAmRNA targets. Database (Oxford). 2015;2015, bav035. [ Links ]
39. Oztemur Y, Bekmez T, Aydos A, Yulug IG, Bozkurt B, Dedeoglu BG. A ranking-based meta-analysis reveals let-7 family as a meta-signature for grade classification in breast cancer. PLoS One. 2015;10(5):e0126837. [ Links ]
40. Calin GA, Dumitru CD, Shimizu M, Bichi R, Zupo S, Noch E, et al. Frequent deletions and down-regulation of micro- RNA genes miR15 and miR16 at 13q14 in chronic lymphocytic leukemia. Proc Natl Acad Sci U S A. 2002;99(24):15524-9. [ Links ]
41. Zhu M, Zhang N, He S. Similarly up-regulated microRNA-106a in matched formalin-fixed paraffin-embedded and fresh frozen samples and the dynamic changes during gastric carcinogenesis and development. Pathol Res Pract. 2014. [ Links ]
42. Zhang W, Qian JX, Yi HL, Yang ZD, Wang CF, Chen JY, et al. The microRNA-29 plays a central role in osteosarcoma pathogenesis and progression. Mol Biol (Mosk). 2012;46(4):622-7. [ Links ]
43. Michael MZ, O' Connor SM, van Holst Pellekaan NG, Young GP, James RJ. Reduced accumulation of specific microRNAs in colorectal neoplasia. Mol Cancer Res. 2003;1(12):882-91. [ Links ]
44. Takamizawa J, Konishi H, Yanagisawa K, Tomida S, Osada H, Endoh H, et al. Reduced expression of the let-7 microRNAs in human lung cancers in association with shortened postoperative survival. Cancer Res. 2004;64(11):3753-6. [ Links ]
45. Tsang WP, Kwok TT. The miR-18a* microRNA functions as a potential tumor suppressor by targeting on K-Ras. Carcinogenesis. 2009;30(6):953-9. [ Links ]
46. Chen PS, Su JL, Hung MC. Dysregulation of microRNAs in cancer. J Biomed Sci. 2012;19:90. [ Links ]
47. Feng Z, Zhang C, Wu R, Hu W. Tumor suppressor p53 meets microRNAs. J Mol Cell Biol. 2011;3(1):44-50. [ Links ]
48. Yamamoto N, Kinoshita T, Nohata N, Yoshino H, Itesako T, Fujimura L, et al. Tumor-suppressive microRNA-29a inhibits cancer cell migration and invasion via targeting HSP47 in cervical squamous cell carcinoma. Int J Oncol. 2013;43(6):1855-63. [ Links ]
49. Han HS, Son SM, Yun J, Jo YN, Lee OJ. MicroRNA-29a suppresses the growth, migration, and invasion of lung adenocarcinoma cells by targeting carcinoembryonic antigen-related cell adhesion molecule 6. FEBS Lett. 2014;588(20):3744-50. [ Links ]
50. Tang W, Zhu Y, Gao J, Fu J, Liu C, Liu Y, et al. MicroRNA-29a promotes colorectal cancer metastasis by regulating matrix metalloproteinase 2 and E-cadherin via KLF4. Br J Cancer. 2014;110(2):450-8. [ Links ]
51. Iorio MV, Ferracin M, Liu CG, Veronese A, Spizzo R, Sabbioni S, et al. MicroRNA gene expression deregulation in human breast cancer. Cancer Res. 2005;65(16):7065-70. [ Links ]
52. Chan M, Liaw CS, Ji SM, Tan HH, Wong CY, Thike AA, et al. Identification of circulating microRNA signatures for breast cancer detection. Clin Cancer Res. 2013;19(16):4477-87. [ Links ]
53. Wang PY, Gong HT, Li BF, Lv CL, Wang HT, Zhou HH, et al. Higher expression of circulating miR-182 as a novel biomarker for breast cancer. Oncol Lett. 2013;6(6):1681-6. [ Links ]
54. Ouyang M, Li Y, Ye S, Ma J, Lu L, Lv W, et al. MicroRNA profiling implies new markers of chemoresistance of triple-negative breast cancer. PLoS One. 2014;9(5):e96228. [ Links ]
55. Wang H, Tan G, Dong L, Cheng L, Li K, Wang Z, et al. Circulating MiR-125b as a marker predicting chemoresistance in breast cancer. PLoS One. 2012;7(4):e34210. [ Links ]
56. Yu Y, Nangia-Makker P, Farhana L, Rajendra GS, Levi E, Majumdar AP. miR-21 and miR-145 cooperation in regulation of colon cancer stem cells. Mol Cancer. 2015;14(1):98. [ Links ]
57. Rosenwald S, Gilad S, Benjamin S, Lebanony D, Dromi N, Faerman A, et al. Validation of a microRNA-based qRT-PCR test for accurate identification of tumor tissue origin. Mod Pathol. 2010;23(6):814-23. [ Links ]
58. Valladares-Ayerbes M, Reboredo M, Medina-Villaamil V, Iglesias-Díaz P, Lorenzo-Patiño MJ, Haz M, et al. Circulating miR-200c as a diagnostic and prognostic biomarker for gastric cancer. J Transl Med. 2012;10:186. [ Links ]
59. Chen X, Ba Y, Ma L, Cai X, Yin Y, Wang K, et al. Characterization of microRNAs in serum: a novel class of biomarkers for diagnosis of cancer and other diseases. Cell Res. 2008;18(10):997-1006. [ Links ]
60. Zhu W, Qin W, Atasoy U, Sauter ER. Circulating microRNAs in breast cancer and healthy subjects. BMC Res Notes. 2009;2:89. [ Links ]
61. Yaman Agaoglu F, Kovancilar M, Dizdar Y, Darendeliler E, Holdenrieder S, Dalay N, et al. Investigation of miR-21, miR-141, and miR-221 in blood circulation of patients with prostate cancer. Tumour Biol. 2011;32(3):583-8. [ Links ]
62. Mahn R, Heukamp LC, Rogenhofer S, von Ruecker A, Müller SC, Ellinger J. Circulating microRNAs (miRNA) in serum of patients with prostate cancer. Urology. 2011;77(5):1265, e9-16. [ Links ]
63. Asaga S, Kuo C, Nguyen T, Terpenning M, Giuliano AE, Hoon DS. Direct serum assay for microRNA-21 concentrations in early and advanced breast cancer. Clin Chem. 2011;57(1): 84-91. [ Links ]
64. van Schooneveld E, Wouters MC, Van der Auwera I, Peeters DJ, Wildiers H, Van Dam PA, et al. Expression profiling of cancerous and normal breast tissues identifies microRNAs that are differentially expressed in serum from patients with (metastatic) breast cancer and healthy volunteers. Breast Cancer Res. 2012;14(1):R34. [ Links ]
65. Liu J, Mao Q, Liu Y, Hao X, Zhang S, Zhang J. Analysis of miR-205 and miR-155 expression in the blood of breast cancer patients. Chin J Cancer Res. 2013;25(1):46-54. [ Links ]
66. Sun Y, Wang M, Lin G, Sun S, Li X, Qi J, et al. Serum microRNA- 155 as a potential biomarker to track disease in breast cancer. PLoS One. 2012;7(10):e47003. [ Links ]
67. Kumar S, Keerthana R, Pazhanimuthu A, Perumal P. Overexpression of circulating miRNA-21 and miRNA-146a in plasma samples of breast cancer patients. Indian J Biochem Biophys. 2013;50(3):210-4. [ Links ]
68. Huang Z, Huang D, Ni S, Peng Z, Sheng W, Du X. Plasma microRNAs are promising novel biomarkers for early detection of colorectal cancer. Int J Cancer. 2010;127(1):118-26. [ Links ]
69. Tsujiura M, Ichikawa D, Komatsu S, Shiozaki A, Takeshita H, Kosuga T, et al. Circulating microRNAs in plasma of patients with gastric cancers. Br J Cancer. 2010;102(7):1174-9. [ Links ]
70. Wang M, Gu H, Wang S, Qian H, Zhu W, Zhang L, et al. Circulating miR-17-5p and miR-20a: molecular markers for gastric cancer. Mol Med Rep. 2012;5(6):1514-20. [ Links ]
71. Zheng Y, Cui L, Sun W, Zhou H, Yuan X, Huo M, et al. MicroRNA- 21 is a new marker of circulating tumor cells in gastric cancer patients. Cancer Biomark. 2011;10(2):71-7. [ Links ]
72. Zhou H, Xiao B, Zhou F, Deng H, Zhang X, Lou Y, et al. MiR- 421 is a functional marker of circulating tumor cells in gastric cancer patients. Biomarkers. 2012;17(2):104-10. [ Links ]
73. Zhang WH, Gui JH, Wang CZ, Chang Q, Xu SP, Cai CH, et al. The identification of miR-375 as a potential biomarker in distal gastric adenocarcinoma. Oncol Res. 2012;20(4):139-47. [ Links ]
74. Wozniak MB, Scelo G, Muller DC, Mukeria A, Zaridze D, Brennan P. Circulating MicroRNAs as Non-Invasive Biomarkers for Early Detection of Non-Small-Cell Lung Cancer. PLoS One. 2015;10(5):e0125026. [ Links ]
75. Wang F, Zheng Z, Guo J, Ding X. Correlation and quantitation of microRNA aberrant expression in tissues and sera from patients with breast tumor. Gynecol Oncol. 2010;119(3):586-93. [ Links ]
76. Schwarzenbach H, Milde-Langosch K, Steinbach B, Müller V, Pantel K. Diagnostic potential of PTEN-targeting miR-214 in the blood of breast cancer patients. Breast Cancer Res Treat. 2012;134(3):933-41. [ Links ]
77. Cookson VJ, Bentley MA, Hogan BV, Horgan K, Hayward BE, Hazelwood LD, et al. Circulating microRNA profiles reflect the presence of breast tumours but not the profiles of microRNAs within the tumours. Cell Oncol (Dordr). 2012;35(4): 301-8. [ Links ]
78. Lawrie CH, Gal S, Dunlop HM, Pushkaran B, Liggins AP, Pulford K, et al. Detection of elevated levels of tumour-associated microRNAs in serum of patients with diffuse large B-cell lymphoma. Br J Haematol. 2008;141(5):672-5. [ Links ]
79. Waters PS, McDermott AM, Wall D, Heneghan HM, Miller N, Newell J, et al. Relationship between circulating and tissue microRNAs in a murine model of breast cancer. PLoS ONE. 2012;7(11):e50459. [ Links ]
80. Morimura R, Komatsu S, Ichikawa D, Takeshita H, Tsujiura M, Nagata H, et al. Novel diagnostic value of circulating miR- 18a in plasma of patients with pancreatic cancer. Br J Cancer. 2011;105(11):1733-40. [ Links ]
81. Si H, Sun X, Chen Y, Cao Y, Chen S, Wang H, et al. Circulating microRNA-92a and microRNA-21 as novel minimally invasive biomarkers for primary breast cancer. J Cancer Res Clin Oncol. 2013;139(2):223-9. [ Links ]
82. Zhu C, Ren C, Han J, Ding Y, Du J, Dai N, et al. A five-microRNA panel in plasma was identified as potential biomarker for early detection of gastric cancer. Br J Cancer. 2014;110(9):2291-9. [ Links ]
83. Bryant RJ, Pawlowski T, Catto JW, Marsden G, Vessella RL, Rhees B, et al. Changes in circulating microRNA levels associated with prostate cancer. Br J Cancer. 2012;106(4):768-74. [ Links ]
84. Song MY, Pan KF, Su HJ, Zhang L, Ma JL, Li JY, et al. Identification of serum microRNAs as novel non-invasive biomarkers for early detection of gastric cancer. PLoS One. 2012;7(3): e33608. [ Links ]
85. Komatsu S, Ichikawa D, Tsujiura M, Konishi H, Takeshita H, Nagata H, et al. Prognostic impact of circulating miR-21 in the plasma of patients with gastric carcinoma. Anticancer Res. 2013;33(1):271-6. [ Links ]
86. Zhao S, Yao D, Chen J, Ding N. Circulating miRNA-20a and miRNA-203 for screening lymph node metastasis in early stage cervical cancer. Genet Test Mol Biomarkers. 2013;17(8): 631-6. [ Links ]
87. Schultz NA, Dehlendorff C, Jensen BV, Bjerregaard JK, Nielsen KR, Bojesen SE, et al. MicroRNA biomarkers in whole blood for detection of pancreatic cancer. JAMA. 2014;311(4):392-404. [ Links ]
88. Ng EK, Chong WW, Jin H, Lam EK, Shin VY, Yu J, et al. Differential expression of microRNAs in plasma of patients with colorectal cancer: a potential marker for colorectal cancer screening. Gut. 2009;58(10):1375-81. [ Links ]
89. Pu XX, Huang GL, Guo HQ, Guo CC, Li H, Ye S, et al. Circulating miR-221 directly amplified from plasma is a potential diagnostic and prognostic marker of colorectal cancer and is correlated with p53 expression. J Gastroenterol Hepatol. 2010;25(10):1674-80. [ Links ]
90. Luo X, Stock C, Burwinkel B, Brenner H. Identification and evaluation of plasma MicroRNAs for early detection of colorectal cancer. PLoS One. 2013;8(5):e62880. [ Links ]
91. Cheng H, Zhang L, Cogdell DE, Zheng H, Schetter AJ, Nykter M, et al. Circulating plasma MiR-141 is a novel biomarker for metastatic colon cancer and predicts poor prognosis. PLoS One. 2011;6(3):e17745. [ Links ]
92. Mar-Aguilar F, Mendoza-Ramírez JA, Malagón-Santiago I, Espino-Silva PK, Santuario-Facio SK, Ruiz-Flores P, et al. Serum circulating microRNA profiling for identification of potential breast cancer biomarkers. Dis Markers. 2013;34(3): 163-9. [ Links ]
93. Ng EK, Li R, Shin VY, Jin HC, Leung CP, Ma ES, et al. Circulating microRNAs as specific biomarkers for breast cancer detection. PLoS One. 2013;8(1):e53141. [ Links ]
94. Mihelich BL, Maranville JC, Nolley R, Peehl DM, Nonn L. Elevated Serum MicroRNA Levels Associate with Absence of High-Grade Prostate Cancer in a Retrospective Cohort. PLoS One. 2015;10(4):e0124245. [ Links ]
95. Selth LA, Townley SL, Bert AG, Stricker PD, Sutherland PD, Horvath LG, et al. Circulating microRNAs predict biochemical recurrence in prostate cancer patients. Br J Cancer. 2013;109(3):641-50. [ Links ]
96. Brase JC, Johannes M, Schlomm T, Fälth M, Haese A, Steuber T, et al. Circulating miRNAs are correlated with tumor progression in prostate cancer. Int J Cancer. 2011;128(3): 608-16. [ Links ]
97. Shen J, Hruby GW, McKiernan JM, Gurvich I, Lipsky MJ, Benson MC, et al. Dysregulation of circulating microRNAs and prediction of aggressive prostate cancer. Prostate. 2012;72(13):1469-77. [ Links ]
98. Chen ZH, Zhang GL, Li HR, Luo JD, Li ZX, Chen GM, et al. A panel of five circulating microRNAs as potential biomarkers for prostate cancer. Prostate. 2012;72(13):1443-52. [ Links ]
99. Ferlay J, Soerjomataram I, Dikshit R, Eser S, Mathers C, Rebelo M, et al. Cancer incidence and mortality worldwide: Sources, methods and major patterns in GLOBOCAN 2012. Int J Cancer. 2015;136(5):E359-86. [ Links ]
100. Mitchell PS, Parkin RK, Kroh EM, Fritz BR, Wyman SK, Pogosova- Agadjanyan EL, et al. Circulating microRNAs as stable bloodbased markers for cancer detection. Proc Natl Acad Sci U S A. 2008;105(30):10513-8. [ Links ]
101. Watahiki A, Macfarlane RJ, Gleave ME, Crea F, Wang Y, Helgason CD, et al. Plasma miRNAs as Biomarkers to Identify Patients with Castration-Resistant Metastatic Prostate Cancer. Int J Mol Sci. 2013;14(4):7757-70. [ Links ]
102. Brunet Vega A, Pericay C, Moya I, Ferrer A, Dotor E, Pisa A, et al. microRNA expression profile in stage III colorectal cancer: circulating miR-18a and miR-29a as promising biomarkers. Oncol Rep. 2013;30(1):320-6. [ Links ]
103. Lodes MJ, Caraballo M, Suciu D, Munro S, Kumar A, Anderson B. Detection of cancer with serum miRNAs on an oligonucleotide microarray. PLoS One. 2009;4(7):e6229. [ Links ]
104. Chan JA, Krichevsky AM, Kosik KS. MicroRNA-21 is an antiapoptotic factor in human glioblastoma cells. Cancer Res. 2005;65(14):6029-33. [ Links ]
105. Chen L, Xiao H, Wang ZH, Huang Y, Liu ZP, Ren H, et al. miR-29a suppresses growth and invasion of gastric cancer cells in vitro by targeting VEGF-A. BMB Rep. 2014;47(1):39-44. [ Links ]