Services on Demand
Journal
Article
Indicators
-
 Cited by SciELO
Cited by SciELO -
 Access statistics
Access statistics
Related links
-
 Cited by Google
Cited by Google -
 Similars in
SciELO
Similars in
SciELO -
 Similars in Google
Similars in Google
Share
Revista Facultad Nacional de Agronomía Medellín
Print version ISSN 0304-2847
Rev. Fac. Nac. Agron. Medellín vol.66 no.1 Medellín Jan./June 2013
Characterization of Carbofuran Degrading Bacteria Obtained from Potato Cultivated Soils with Different Pesticide Application Records
Caracterización de Bacterias Degradadoras de Carbofuran Obtenidas de Suelos Bajo Cultivo de Papa y con Diferente Historia de Aplicación de Plaguicidas
José Castellanos Rozo1; Jimena Sánchez Nieves2; Daniel Uribe Vélez3; Leonardo Moreno Chacón4 and Luz Marina Melgarejo Muñoz5
1 Researcher. Universidad Nacional de Colombia - Sede Bogotá - Facultad de Ciencias - Instituto de Biotecnología. A.A. 14490, Bogotá, Colombia. <jmcastellanosr@unal.edu.co>
2 Assistant Professor. Universidad Nacional de Colombia - Sede Bogotá - Facultad de Ciencias - Departamento de Biología. Carrera 45 No. 26 - 85, Bogotá, Colombia. <jsanchezn@unal.edu.co>
3 Associate Professor. Universidad Nacional de Colombia - Sede Bogotá - Facultad de Ciencias - Instituto de Biotecnología. A.A. 14490, Bogotá, Colombia. <duribev@unal.edu.co>
4 Researcher. Universidad Nacional de Colombia -Sede Bogotá -Laboratorio de Fisiología y Bioquímica Vegetal. A.A. 14490, Bogotá, Colombia. <almorenoch@unal.edu.co>
5 Titular Professor. Universidad Nacional de Colombia - Sede Bogotá - Facultad de Ciencias - Departamento de Biología. Carrera 45 No. 26 - 85, Bogotá, Colombia. <lmmelgarejom@unal.edu.co>
Received: February 9, 2012; accepted: November 13, 2012.
Abstract
Eighty-two bacterial isolates with potential Carbofuran degradation activity (Furadan®3SC) were obtained from soils cultivated with the potato variety Unica (Solanum tuberosum) in Silos, Norte de Santander (Colombia), with different records of pesticide application. The bacteria were selected for their ability to grow at 25 °C for 72 h in media containing 200 mg L-1 of analytical Carbofuran as the sole source of carbon and/ or nitrogen. The results showed that ten isolates, 12% of those obtained, grew in the culture media. Eight of theses isolates were obtained from soils with a high pesticide exposure (eight years of application), and identified, by macroscopic, microscopic and biochemical characteristics, as Sphingomonas paucimobilis. The other two were obtained from soils with three years and one year of application and were identified as Pseudomonas aeruginosa and Yersinia pseudotuberculosis, respectively. Subsequently, these bacteria were tested for their ability of hydrolytic degradation of Carbofuran; the results show that the pesticide was degraded only by the isolates of Sphingomonas paucimobilis for 72 h. The results obtained in the in vitro tests show the bacterial metabolic capacity for the biodegradation of Carbofuran, highlighting the potential use of the bacteria for future field evaluation tests in places where residues of the pesticide may exist, as an alternative to control the impact that N-methyl carbamate pesticides have on the environment and human health.
Key words: Pesticides, Sphingomonas paucimobilis, selection, biodegradation.
Resumen
Ochenta y dos aislamientos bacterianos con actividad potencial de degradación de Carbofuran (Furadan®3SC), fueron obtenidos de suelos, cultivados con papa (Solanum tuberosum) variedad Única, del municipio de Silos, Norte de Santander (Colombia) con diferente historia de aplicación del plaguicida. Las bacterias fueron seleccionadas por su capacidad para crecer a 25 °C durante 72 h, en medios de cultivo conteniendo 200 mg L-1 de Carbofuran analítico como única fuente de carbono y/o nitrógeno. Los resultados mostraron que diez aislamientos correspondientes al 12 % de los obtenidos, crecieron en el medio de cultivo. Ocho de los aislamientos fueron obtenidos del suelo con mayor historia de exposición al plaguicida (ocho años de aplicación), e identificados por características macroscópicas, microscópicas y bioquímicas como Sphingomonas paucimobilis. Los otros dos fueron obtenidos de suelos con tres y un año de aplicación y caracterizados como Pseudomonas aeruginosa y Yersinia pseudotuberculosis, respectivamente. Posteriormente, estas bacterias fueron evaluadas por su capacidad para degradar Carbofuran por vía hidrolítica, demostrando que el plaguicida fue degradado únicamente por los aislamientos de Sphingomonas paucimobilis durante un tiempo de 72 h. Los resultados obtenidos en los ensayos in vitro denotan la capacidad metabólica que tienen estas bacterias para la biodegradación de Carbofuran, destacando su potencial utilización para futuros ensayos de evaluación en campo, en lugares en donde pueda existir residualidad del plaguicida, como alternativa para controlar el posible impacto que tienen los plaguicidas N-metilcarbamatos en el medio ambiente y en la salud humana.
Palabras clave: Plaguicidas, Sphingomonas paucimobilis, selección, biodegradación.
Carbofuran is a widely used N-methyl carbamate mobility in soils (depending on their characteristics) pesticide in global agriculture as an insecticide, and poor management practices associated with acaricide and nematicide (Chelinho et al., 2011). overdose can cause potential risks to public health Its high toxicity to animals and humans, significant with diseases such as lung cancer (Bonner et al., 2005) and reproductive problems, among others (Goad et al., 2004).
Studies by Parekh et al. (1994) determined that bacteria are mainly responsible for the degradation of Carbofuran in acid and neutral soils. Two main routes of bacterial degradation of N-methyl carbamate and especially of Carbofuran have been proposed: oxidative and hydrolytic pathways (Chaudhry and Ali, 1988). However, it has been found that hydrolytic degradation of Carbofuran produces less toxic metabolites compared to the oxidative pathway (Chapalmadugu and Chaudhry, 1991). Carbofuran hydrolysis may occur by two pathways: the instability of the ester bond of the carbonyl group of the N-methylcarbamic acid attached to the phenol or the amide bond of N-methylcarbamic acid, both producing Carbofuran 7-phenol (2,3-dihydro-2,2-dimethyl-7benzofuranol), a less toxic metabolite than Carbofuran, carbon dioxide and methylamine. The latter is used as a source of carbon and/or nitrogen by a diverse group of bacteria that hydrolyzes Carbofuran but does not degrade the aromatic ring (Trabue et al., 2001).
Few studies, describing bacteria that hydrolyze Carbofuran, have shown complete degradation of the aromatic ring structure. Feng et al. (1997) reported that strain CF06 of the genus Sphingomonas is capable of completely mineralizing the aromatic ring of Carbofuran without detection of the metabolites involved. Kim et al. (2004) identified strain SB5a of Sphingomonas sp., isolated from soil with a five year Carbofuran application history, which showed activity on the aromatic ring by hydrolysis of Carbofuran 7-phenol, producing an intermediary metabolite called 2-hydroxy-3[3-methylpropan-2-ol]phenol, which is then converted into red metabolites. Subsequently, Park et al. (2006) characterized, by mass spectrometry and nuclear magnetic resonance analysis (NMR), one of these red metabolites as 5[2-hydroxy-2-methylpropyl]2,2-dimethyl-2,3-dihydro-naphtho [2,3-6] furan-4,6,7,9-tetrone, further suggesting that this red metabolite comes from the condensation of some metabolites of the degradation of 2-hydroxy3[3-methylpropan-2-ol]phenol (Park et al., 2006). On the other hand, Yan et al. (2007) isolated and characterized strain FND-3 of Novosphingobium sp., isolated from a waste water treatment system in a manufacturing company of pesticides in China, finding that this bacteria, in addition to degrading the Carbofuran aromatic ring with the hydrolytic pathway and producing the metabolite 2-hydroxy3[3-methylpropan-2-ol) phenol, had the ability to hydrolyze the ether bond of the Carbofuran furanyl ring, producing the metabolite 2-hydroxy-3[3methylpropan-2-ol) benzene-N-methyl carbamate and Carbofuran, generating the metabolite 5-hydroxyCarbofuran.
The aim of this study was to isolate native aerobic bacteria in the rhizosphere of Solanum tuberosum plants (Unica variety) in Silos, Norte de Santander (Colombia), in agricultural soils with different records of Carbofuran application, characterized by their ability for hydrolytic degradation of Carbofuran (Furadan® 3SC - commercial product).
Materials and methods
Sampling. Samples were collected from farms located in the "La Laguna" district of Silos, Norte de Santander (N-7° 15' 14.76", W-72° 45' 49.068"). Three farms were selected (based on a survey of producers and counseling from ICA Pamplona, Colombia) with soils that had different records of application of Furadan® 3SC (commercial product) of 1, 3 and 8 years, and that were cultivated with the potato (Solanum tuberosum) cultivar Unica and sampled at a commercial maturity stage. Three integrated samples were obtained from each farm, each comprised of 10 subsamples of rhizosphere soil, taken with a zig-zag pattern from different areas of the cultivation, at a depth of 0 to 15 cm, according to Ogram et al. (2000). The samples were taken to the laboratory under refrigerated conditions at 4 °C and processed independently.
Counting and isolation of bacteria with potential degradation activity of Carbofuran. 10.0 g of each soil were weighed and added to 90 mL of a sterile solution of 0.85% NaCl (w/v). The study employed serial dilutions and triplicate surface plating for viable cell count (Madigan et al., 2002) in minimal mineral salt media: M1, M2, M3 and M4 (Bano and Musarrat, 2004). The minimal medium M1 (Biolabs reagent, 98% purity) had the following composition (g L-1): Na2HPO4, 2.1; MgSO4, 0.01; CaCl2.2H2O, 0.1; FeSO4.7H2O, 0.001; CuSO4, 0.04; Na2MoO4, 0.002; (NH4)2SO4, 1.0 and Furadan® 3SC (commercial pesticide), 0.1, as the sole carbon source. The M2 minimal medium was prepared like M1, but without (NH4)2SO4 and with glucose (1%) as the carbon source and Furadan® 3SC (0.1 g L-1) as the sole nitrogen source. The M3 minimal medium was prepared like M1, but without (NH4)2SO4 and with Furadan® 3SC (0.1 g L-1) as the sole source of carbon and nitrogen. The M4 minimal medium (positive growth control) was prepared like M1, but with glucose (1%) and Furadan® 3SC (0.1 g L-1). All of the culture media were prepared by dissolving the components in distilled and deionized HPLC grade water and agar-agar (15 g L-1) and adjusting the pH to 7.0. Subsequently, the inoculated plates were incubated at 25 °C for eight days in darkness for bacterial growth.
Counts for total viable bacteria cells in terms of colony forming units per gram of soil (CFU g-1), and counts for isolates and number of individuals per isolate were carried out. Each bacterial isolate was purified by streaking it on agar (Madigan et al., 2002) in minimal medium M4, performing Gram stain and description of the macroscopic and microscopic characteristics according to Holt et al. (1994). Each isolate was then plated in the minimum medium M4, incubated for 48 h and preserved in cryopreservation vials at -70 °C in a Gherna medium (Gibson and Khoury, 1986).
Selection of bacteria for growth in medium with Carbofuran (analytical reagent). The obtained isolates were replicated in the minimum solid culture media M1, M2, and M3 with 200 mg L-1 of Carbofuran (Aldrich analytical grade, 99% purity), as the sole source of carbon and/or nitrogen; and incubated at 25 °C. Isolates that grew for 72 h were selected.
Characterization of bacterial isolates. Taxonomic identification of the selected bacterial isolates was performed with the commercial identification system: BBL CRYSTAL® (Becton Dickinson, Maryland, USA) pursuant to the manufacturer's instructions, followed by complementary biochemical tests using substrates according to Bergey's Manual for bacteriological determination (Holt et al., 1994).
Evaluation of bacterial growth and degradation of Carbofuran. Each bacterial isolate was massively grown in the solid culture medium LB (Luria Bertani) at 25 °C for 48 h. The LB culture medium was prepared with the following composition (g L-1): yeast extract 5 (Oxoid, 98% purity); tryptone, 10 (Oxoid, 98% purity); sodium chloride, 10 (Biolabs, 98% purity); agar-agar, 15 (Oxoid, 98% purity), pH 7. Subsequently, the microbial biomass was suspended, washed with minimal medium and recovered with centrifugation at 6,000 rpm for 5 min at room temperature for two times, adjusting the bacterial biomass to an optical density of 0.500 ± 0.005 at 600 nm using a spectrophotometer (BioRad® SmartSpec). Then 2 mL of the biomass were inoculated in 50 mL tubes containing 18 mL of minimal medium with 200 mg L-1 of analytical grade Carbofuran as the sole source of carbon and nitrogen; with constant stirring at 150 rpm and 25 °C, according to the modified protocol of Chaudhry and Ali (1988).
To evaluate the growth of bacteria in media containing Carbofuran as the source of carbon and/or nitrogen, samples of 100 microliters (µL) of culture were taken at 0, 72 and 120 h. Each sample was serially diluted, and 100 µL of each dilution were plated on solid LB medium; and incubated at 25 °C for 48 h. Afterwards, a surface plate count of viable cells was carried out (Madigan et al., 2002).
To determine degradation of Carbofuran, 1 mL samples were taken at 0, 72 and 120 h. The pesticide was extracted three times with chloroform (1:1), after which the chloroform was evaporated and the residues were dissolved in methanol. Then, the concentration of the pesticide was determined in µg mL-1 with the spectrophotometric method described by Zhelev et al. (2004). The negative control consisted of minimal mineral salt media with Carbofuran as the sole source of carbon and/or nitrogen without a bacterial inoculum; processed in the same manner described above. The strain ER2 of Aminobacter ciceronei nov., previously described as a Carbofuran degrading bacteria, and Carbofuran, as the source of carbon and/or nitrogen (Topp et al., 1993), were used as a positive control of microbial growth. The strain was kindly provided by Dr. E. Topp and was used as described by McDonald et al. (2005).
Statistical Analysis.Data from growth and degradation for each isolate, for each medium, were analyzed using analysis of variance for repeated measurements, with an α of 0.05 using SPSS for Windows v13. To determine differences between the application treatments and the control treatment, a Dunnett's test was used to find differences between treatments with the obtained isolates; the multiple comparison test LSD (Least Significant Difference) was used.
Results and discussion
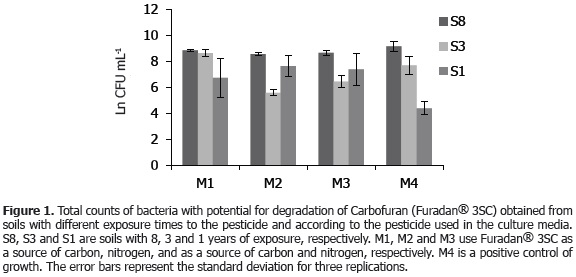
Isolation and count of bacteria with potential degradation activity of Carbofuran. Figure 1 shows the natural log of the CFU g-1 soil of Carbofuran degrading bacteria obtained for the three soil types in the culture media used. The results show that soils with eight years of Carbofuran application (Furadan® 3SC, commercial product) had higher total bacterial counts, using Carbofuran as the sole source of carbon and/or nitrogen; in contrast to soils with one year application of the pesticide. This could be related to that described by Trabue et al. (2001), who, in determining the dynamics of Carbofuran degrading communities with the most probable number technique with radiolabeled Carbofuran in the aromatic ring and lateral chain (14C-NMP) in sites treated with annual applications of Carbofuran for three consecutive years, found increased bacterial populations that hydrolyze Carbofuran to Carbofuran 7-phenol and methylamine, using the latter as a source of carbon and/or nitrogen. However, in the same study, it was found that the Carbofuran aromatic ring degrading communities (degrading Carbofuran 7-phenol) were small and stable because they did not increase in number, suggesting that Carbofuran degradation is also a cometabolic process.
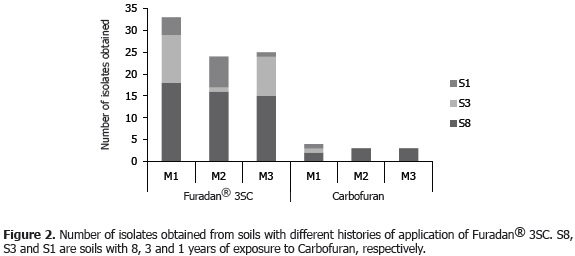
Carbofuran degrading bacterial isolates were obtained from soils with different records of exposure to pesticides. The soil with the longest history of Furadan® 3SC application (eight years) showed the highest number of isolates (49), in contrast to the soils with three years (21) and one year of exposure to the pesticide (12) (Figure 2). Most of the isolates were Gram-negative bacilli with oxidase activity, exopolysaccharide production and catalase positive, similar to the characteristics observed in the isolates described by Chaudhry and Ali (1988); Parekh et al. (1994); Feng et al. (1997); Kim et al. (2004); Wu et al. (2006).
The above results suggest that the increase in Carbofuran degrading bacterial communities and the diversity of the isolates obtained were caused by the adaptation of bacterial communities in an environment continually exposed to the pesticide. This is probably due to horizontal gene transfer. Studies by Desaint et al. (2003) showed that the mcd gene "encoding the Carbofuran hydrolase enzyme" is transferred between microbial populations in the soil, which may explain, at least in part, the high abundance and genetic diversity of Carbofuran degradation. However, some authors suggest that bacterial genera, such as Sphingomonas sp., limit gene transfer to other genera. Probably, this is why strains of Sphingomonas sp. are widely recognized as degrading xenobiotic compounds (Basta et al., 2004; Stolz, 2009; Dong et al., 2012). Horizontal gene transfer in soil has also been described for other pesticide degrading pathways, including the gene transfer of atz that is the operon involved in the degradation of atrazine (Devers et al., 2005; Liang et al., 2012).
Selection of bacteria for growth in medium with Carbofuran (analytical reagent). The results showed that of the 82 isolates obtained with the commercial product (Furadan® 3SC), only 10 isolates, or 12% of the obtained isolates, grew at 25 °C for 72 h in the presence of analytical grade Carbofuran as the sole source of carbon and/or nitrogen (Figure 2).
Characterization of bacterial isolates. The results showed that of the 10 isolates obtained with the analytical grade Carbofuran, 8 had similar macroscopic (yellow colonies, mucoid, defined edge, convex elevation) and microscopic characteristics (short Gram-negative bacilli with cytoplasmic inclusions), and similar biochemical characteristics. The characterization of the bacterial isolates with the commercial identification system: BBL CRYSTALTM® (Becton Dickinson, Maryland, USA), following the manufacturer's instructions and additional biochemical tests with substrates according to Bergey's Manual for bacteriological determination (Holt et al., 1994); determined that the isolates obtained in the three minimal media with Carbofuran (M1, M2 and M3) as the sole source of carbon and/or nitrogen, with soil with eight years of Furadan® 3SC application, belong to the genus Sphingomonas sp., with a reliability percentage between 96 and 99% (Table 1). These results coincide with that described by Feng et al. (1997); Ogram et al. (2000); Kim et al. (2004); Wu et al. (2006), who also isolated and characterized Sphingomonas sp. from soil with a history of Carbofuran application.
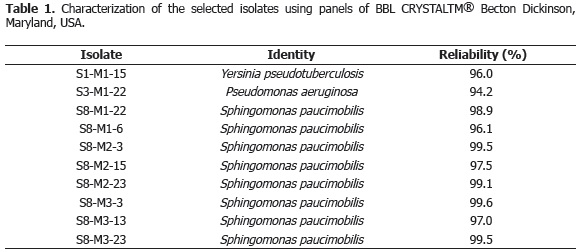
On the other hand, an isolate (S3-M1-22) from the soil with a history of three years of Carbofuran (Furadan® 3SC) and obtained from the minimal medium with Carbofuran (analytical grade) as the sole carbon source (M1) was characterized, using BBL CRYSTALTM® and complementary biochemical tests (grown in cetrimide agar, grown at 42 °C, nitrate positive, gelatin liquefaction and positive mobility), as Pseudomonas aeruginosa, with a reliability percentage of 94.2% (Table 1). Globally, several isolates identified as Pseudomonas sp. have been described as capable of degrading Carbofuran (by the oxidative and/or hydrolytic pathway) used as a carbon source (Chauldry and Ali, 1988; Slaoui et al., 2001; Bano and Musarrat, 2004). Chauldry and Ali (1988) isolated Pseudomonas sp. 50432, which was able to mineralize Carbofuran used as the sole carbon source. They determined that this bacteria was unable to grow from Carbofuran 7-phenol as a carbon source or methylamine as a sole source of carbon and nitrogen, which suggests the oxidative degradation of Carbofuran. Subsequent experiments determined that Pseudomonas sp. 50432 possesses both Carbofuran degradation mechanisms (oxidative and hydrolytic); concluding that the oxidative mechanism, produced by a hydroxylase enzyme, is an inducible characteristic and converts Carbofuran to a metabolite identified as 4-hydroxyCarbofuran, which in turn is degraded to other yet unknown metabolites, achieving complete mineralization (Mateen et al., 2002).
Another isolate, from soil with one year of application (S1-M1-15), which grew in analytical grade Carbofuran as the sole carbon source (medium M1), was characterized as Yersinia pseudotuberculosis with 96% reliability. This bacterium, due to its potential biological risk, was discarded from the study.
Evaluation of the use of Carbofuran as the sole source of carbon and/or nitrogen. Table 2 shows the growth (Ln CFU mL-1) of the selected isolates from analytical Carbofuran, when it was supplied as the sole source of carbon and/or nitrogen in minimal mineral salt media.
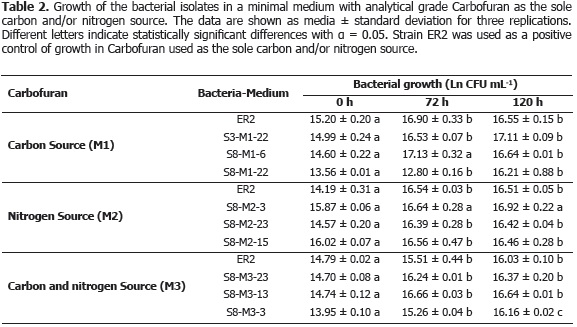
Statistical analysis determined that there were no significant differences between growth of the isolate Aminobacter ciceronei strain ER2 and of the isolates of the present study when Carbofuran was supplied as the sole source of carbon and/or nitrogen. Additionally, nor were there differences between the native isolates obtained, suggesting that they may be colonies of the same microbial population. Moreover, Figure 3 shows the degradation of Carbofuran (µg mL-1) for the isolates obtained when Carbofuran was supplied at a concentration of 200 mg L-1 as the sole source of carbon and/or nitrogen in the minimal mineral salt medium. The Dunnett's test indicated that there were significant differences between the degradation by the isolates of this study and the control treatment (without bacteria). All these results indicate that the selected bacterial isolates degrade the Carbofuran used as the source of carbon and/or nitrogen.
On the other hand, the LSD test indicated that isolate treatments S8-23-M3, M2-23-S8, S8-M3-13, S8-S8 6 and M1-M1-22 presented significant differences in degradation as compared to the ER2 isolate, indicating degradation of Carbofuran via Carbofuran 7-phenol, when used as the sole source of carbon and/or nitrogen in the times of 72 and 120 h (Figure 3).
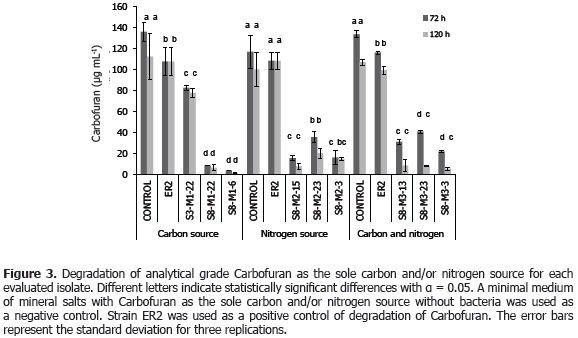
Results reported by Topp et al. (1993) showed that strain ER2 has the capacity to hydrolyze Carbofuran to Carbofuran 7-phenol and methylamine using methylamine as the only source of carbon and nitrogen and accumulating Carbofuran 7-phenol, with a doubling time of three hours. The analytical technique used in this study is based on the formation of a diazonium salt, among the hydrolytic product of Carbofuran (Carbofuran 7-phenol) and aniline, in the presence of sodium nitrite in a basic medium (Zhelev et al., 2004); which explains why Carbofuran degradation cannot be detected for strain ER2. However, the lack of significant differences between the growth of strain ER2 and the isolates obtained in this study in the three culture media (M1, M2 and M3) indicates that the strain ER2 is using Carbofuran as a source of carbon and/or nitrogen (Topp et al., 1993). The results also indicate that there were no significant differences with respect to degradation between the isolates S8-M1-6 and S8-M1-22, S8-M2-3 and S8-M215, and S8-M3-13 and S8-M3-3. It may be possible that there were no differences in their capacity to degrade Carbofuran because all these isolates were identified as Sphingomonas sp. and are from the same type of soil (soil with eight years of Furadan® 3SC application), suggesting that these isolates may be the same bacteria.
Studies by Kim et al. (2004) determined that Sphingomonas sp. strain SB5. Fully hydrolyzed 0.91 mM of Carbofuran to Carbofuran 7-phenol and methylamine in 12 h at 25 °C; and can also hydrolyze Carbofuran 7-phenol as a carbon source in 48 h using methylamine as a source of carbon and nitrogen, which agrees with the results obtained in this study. On the other hand, studies by Jiang et al. (2007) determined that Sphingomonas sp. CDS-1 can degrade 100 mg L-1 of Carbofuran in 32 h at 25 °C, while the strains obtained in this study degraded 200 mg L-1 of Carbofuran in 72 h at 25 °C. Moreover, Jiang et al. (2007) also reported that low concentrations of glucose have no effect on the degradation of the pesticide, which is consistent with the lack of significant differences encountered in the obtained degradation by isolates from Sphingomonas sp. in different tested culture media, when Carbofuran was used as a source of carbon and/or nitrogen.
Studies by Yan et al, (2007) demonstrated that Novosphingobium sp. FND-3 (Gram-negative, oxidase positive, with 97% genetic similarity to Sphingomonas sp. SB5) was able to degrade Carbofuran in a minimal mineral salt medium with 100 mg L-1 of Carbofuran as the sole carbon source at 30 °C at a rate of 28.6 mg L-1 per hour. These results suggest that NDF-3 used several metabolic pathways to degrade Carbofuran; hydrolyzing Carbofuran not only in the ether and ester bonds but also the benzene ring of the pesticide, producing 5-hydroxyCarbofuran, as has been described for other bacteria such as Rhodococcus sp. TEI (Behki et al., 1994).
Another common characteristic observed among isolates obtained in this study identified as Sphingomonas sp. is the production of red pigments in the degradation of Carbofuran (results not shown), which have already been reported in strains that degrade this pesticide (Feng et al.,1997; Kim et al., 2004; Wu et al., 2006). One of these red metabolites has been characterized in Sphingomonas sp. CF06 and Sphingomonas sp. SB5 as 5[2-hydroxy-2methyl-propyl]2, 2-dimethyl-2,3-dihydro-naphtho [2,3-6] furan-4 ,6, 7, 9-tetrone. This red metabolite is formed when Carbofuran 7-phenol is hydrolyzed by the bacteria, producing the intermediate 2-hydroxy-3(3-methylpropan-2-ol) phenol which condenses to the mentioned red metabolites (Park et al., 2006).
The intermediary metabolite 2-hydroxy-3[3- methylpropan2-ol) phenol was also observed in Sphingomonas sp. CDS-1 and Novosphingobium sp. FND-3 (Wu et al., 2006, Yan et al., 2007). Possibly, the isolates obtained in this study possess the same pathway described for the hydrolytic degradation of Sphingomonas sp. (Park et al., 2006) or Novosphingobium sp. FND-3 (Yan et al., 2007), which would be very useful, since these native bacteria not only reduce the deleterious effect of Carbofuran, producing less toxic metabolites through hydrolysis, but can also mineralize various aromatic metabolites produced by Carbofuran degradation in the environment, which would result in the complete removal of the pesticide in conditions where it exists in residues.
Sphingomonas sp. has been reported to have a high metabolic capacity to degrade xenobiotic compounds; and has been found to not only degrade Carbofuran, but also other pesticides. Furthermore, Feng et al. (1997) reported that Sphingomonas sp. CF06, in addition to degrading Carbofuran, Carbofuran 7-phenol and methylamine, has the ability to degrade carbaryl, gentisate and protocatechuic acid. Likewise, Novosphingobium sp. strain FND-3 showed that degrading Carbofuran presented cross degradation activity for isoprocarb, metolcarb and methomyl (Yan et al., 2007). This indicates that it is important to determine the capacity of degradation of other xenobiotics often used in potato cultivation such as carbaryl, isoprocarb, metolcarb and methomyl, to determine the potential that the isolates obtained in this research may possess, and for further studies on the bioremediation of water and soil contaminated with pesticides to be carried out.
Conclusions
The results of this study support the conclusion that the selected isolates characterized as Sphingomonas sp. and obtained from soil with eight years of Furadan® 3SC applications grow and degrade Carbofuran and the aromatic ring of Carbofuran via Carbofuran 7-phenol for 72 h when the pesticide is supplied as the sole source of carbon and/or nitrogen. These results suggest that increased exposure time to a compound such as Carbofuran acts as a factor that stimulates the selection of bacteria that have the capacity to efficiently degrade the pesticide.
Acknowledgements
The authors thank to Research Division of the Universidad Nacional de Colombia, Bogotá for funding the project "Búsqueda y caracterización bioquímica y molecular de microorganismos con potencial para degradar agentes xenobióticos" code 2004803947, the Biotechnology Institute (Laboratory of Agriculture Microbiology), the Biology Department (Research Laboratory of Plant Physiology and Biochemistry), Dr. E. Topp for supplying the ER2 reference strain used as the positive control in this study and Sandra Milena Campos Alba for her valuable contribution to the bacterial characterization.
Bibliography
Bano, N. and J. Musarrat. 2004. Characterization of a novel carbofuran degrading Pseudomonas sp. with collateral biocontrol and plant growth promoting potential. FEMS Microbiology Letters 231(1): 13-17. [ Links ]
Basta, T., A. Keck, J. Klein and A. Stolz. 2004. Detection and characterization of conjugative degradative plasmids in xenobiotics degrading Sphingomonas strains. Journal of Bacteriology 186(12): 3862-3872. [ Links ]
Behki, R.M., E.E. Topp and B.A. Blackwell. 1994. Ring hydroxylation of N-methylcarbamate insecticides by Rhodococcus TE1. Journal of Agricultural and Food Chemistry 42(6): 1375-1378. [ Links ]
Bonner, M.R., W.J. Lee, D.P. Sandler, J.A. Hoppin, M. Dosemeci and M.C. Alavanja. 2005. Occupational exposure to carbofuran and the incidence of cancer in the agricultural health study. Environmental Health Perspectives 113(3): 285-289. [ Links ]
Chaudhry, G.R. and A.N. Ali. 1988. Bacterial metabolism of Carbofuran. Applied and Environmental Microbiology 54(6): 1414-1419. [ Links ]
Chapalmadugu, S. and G.R. Chaudhry. 1991. Hydrolysis of carbaryl by a Pseudomonas sp and construction of a microbial consortium that completely metabolizes carbaryl. Applied and Environmental Microbiology 57(3): 744-750. [ Links ]
Chelinho, S, K. D. Sautter, A. Cachada, I. Abrantes, G. Brown, D.A. Costa and J.P. Sousa. 2011. Carbofuran effects in soil nematode communities: Using trait and taxonomic based approaches. Ecotoxicology and Environmental 74(7): 2002-2012. [ Links ]
Desaint, S, S. Arrault, S. Siblot and J.C. Fournier. 2003. Genetic transfer of the mcd gene in soil. Journal of Applied Microbiology 95(1): 102-108. [ Links ]
Devers, M., S. Henry, A. Hartmann and F.M. Laurent. 2005. Horizontal gene transfer of atrazine-degrading genes (atz) from Agrobacterium tumefaciens St96-4 pADP1: Tn5 to bacteria of maize-cultivated soil. Pest Management Science 61(9): 770-780. [ Links ]
Dong, H.S., D.U. Kim, C.N. Seong, H.G. Song and J.O. Ka. 2012. Genetic and phenotypic diversity of carbofuran-degrading bacteria isolated from agricultural soils. Journal Microbiology Biotechnology 22(4): 448-456. [ Links ]
Feng, X., L.T. Ou and A.V. Ogram. 1997. Plasmidmediated mineralization of carbofuran by Sphingomonas sp. strain CF06. Applied and Environmental Microbiology 63(8): 1332-1337. [ Links ]
Goad, RT; J.T. Goad, B.H. Atieh and R.C. Gupta. 2004. Carbofuran-induced endocrine disruption in adult male rats. Toxicology Mechanisms and Methods 14(4): 233-239. [ Links ]
Gibson, L.F. and J.T. Khoury. 1986. Storage and survival of bacteria by ultra freeze. Letters in Applied Microbiology 3(6): 127-129. [ Links ]
Holt, J., N. Krieg, N. Snealth, J. Staley and S. Williams. 1994. Bergey's manual of determinative bacteriology. Ninth Edition. William and Wilkins Editors, Baltimore. Maryland, USA. 787 p. [ Links ]
Kim, I.S., J.Y. Ryu, H.G. Hur, M.B. Gu, S.D. Kim and J.H. Shim. 2004. Sphingomonas sp. strain SB5 degrades carbofuran to a new metabolite by hydrolysis at the furanyl ring. Journal of Agricultural and Food Chemistry 52(8): 2309-2314. [ Links ]
Liang, B, J. Jiang, J. Zhang, Y. Zhao and S. Li. 2012. Horizontal transfer of dehalogenase genes involved in the catalysis of chlorinated compounds: evidence and ecological role. Critical Reviews in Microbiology 38(2): 95-110. [ Links ]
Madigan, M.T., M. Martinko and J. Parker. 2002. Brock biology of microorganisms. Tenth edition. Prentice Hall, Iberia, Madrid. 986 p. [ Links ]
Mateen, A., G.R. Chaudhry, B. Kaskar, M. Sardessai, M. Bloda, A.R. Bhatti and S.K. Walia. 2002. Induction of carbofuran oxidation to 4-hydroxycarbofuran by Pseudomonas sp. 50432. FEMS Microbiology Letters 214(2): 171-176. [ Links ]
McDonald, I.R., P. Kampfer, E. Topp, K.L. Warner, M.J. Cox, T.L. Hancock, L.G. Miller, M.J. Larkin, V. Ducrocq, C. Coulter, D.B. Harper, J.C. Murrell and R.S. Oremland. 2005. Aminobacter ciceronei sp. nov. and Aminobacter lissarensis sp. nov., isolated from various terrestrial environments. International Journal of Systematic and Evolutionary Microbiology 55(5): 1827:1832. [ Links ]
Ogram, A.V., Y.P. Duan, S.L. Trabue, X. Feng, H. Castro and LT. Ou. 2000. Carbofuran degradation mediated by three related plasmid systems. FEMS Microbiology Ecology 32(3): 197-203. [ Links ]
Parekh, N.R., D.L. Suett, S.J. Roberts, T. McKeown, E.D. Shaw and A.A. Jukes. 1994. Carbofuran degrading bacteria from previously treated field soils. Journal of Applied Bacteriology 76(6): 559-567. [ Links ]
Park, M.R., L. Sunwoo, H. Tae-ho, O. Byung-tack, H.S. Jae and S.K. In. 2006. A new intermediate in the degradation of carbofuran by Sphingomonas sp. strain SB5. Journal of Microbiology and Biotechnology 16(8): 1306-1310. [ Links ]
Slaoui M, Lamrabet, Ouhsssine, Massoui M, Elyachioui M. 2001. Dégradation du carbofuran par une bactérie du genre Pseudomonas sp isolée à partir du sol. Sci. Lett. 3(3). [ Links ]
Stolz, A. 2009. Molecular characteristics of xenobiotic degrading sphingomonads. Applied Microbiology and Biotechnology 81(5):793-811. [ Links ]
Trabue, S.L., A.V. Ogram and L.T. Ou. 2001. Dynamics of Carbofuran degrading microbial communities in soil during three successive annual applications of carbofuran. Soil Biology and Biochemistry 33(1): 75-81. [ Links ]
Topp, E., R.S. Hanson, D.B. Ringelberg, D.C. White and R. Wheatcroft. 1993. Isolation and characterization of an N-methylcarbamate insecticide-degrading methylotrophic bacterium. Applied and Environmental Microbiology 59(10): 3339-3349. [ Links ]
Wu, J., Q. Hong, Y.N. Chen and S.P. Li. 2006. Analysis of the degrading products of carbofuran by Sphingomonas sp. with GC/MS and GC/FTIR. Spectroscopy and Spectral Analysis 26(5): 1716-1719. [ Links ]
Yan, Q., Q. Hong, P. Han, X. Dong, Y. Shen and S. Li. 2007. Isolation and characterization of a carbofuran degrading strain Novosphingobium sp. FEMS Microbiology Letters 271(2): 207-213. [ Links ]
Zhelev, I.Z., V.G. Vrabcheva and M.T.V. Yotsev. 2004. Spectrophotometric determination of the carbamate insecticide carbofuran in biological samples. Bulgarian Journal of Veterinary Medicine 7(3): 189-193. [ Links ]













