Services on Demand
Journal
Article
Indicators
-
 Cited by SciELO
Cited by SciELO -
 Access statistics
Access statistics
Related links
-
 Cited by Google
Cited by Google -
 Similars in
SciELO
Similars in
SciELO -
 Similars in Google
Similars in Google
Share
Actualidades Biológicas
Print version ISSN 0304-3584
Actu Biol vol.30 no.88 Medellín Jan./June 2008
ARTÍCULOS DE INVESTIGACIÓN
DNA EXTRACTION FROM HEAVY OIL CONTAMINATED MICROCOSMS AND RPOB GENE PCR AMPLIFICATION
EXTRACCIÓN DE ADN A PARTIR DE MICROCOSMOS CONTAMINADOS CON PETRÓLEO PESADO Y AMPLIFICACIÓN DEL GEN RPOB
Lucía C. Lozano–A.1; María A. Bautista1, Jenny Dussan–G.1, 2, Martha J. Vives–Flórez1
1 Centro de Investigaciones Microbiológicas (CIMIC). Departamento de Ciencias Biológicas, Universidad de los Andes. A. A. 4976. Bogotá, Colombia.
2 Correo electrónico:<jdussan@uniandes.edu.co>
Abstract
This work describes a new method for extracting genomic DNA from heavy oil–contaminated soils. This method combines soil washes using three washing solutions with enzymatic lysis (SW–EL method). The suitability of the SW–EL method was validated by extracting DNA from samples as small as 0.5 g from model microcosms comprising soil artificially contaminated with 1 and 10% (w/w) heavy petroleum and seeded with a bacterial consortium. Compared with an established protocol for lysis and DNA purification using the commercial kit GeneReleaserTM (Bioventures) (GRL method), the novel SW–EL method provided higher DNA yields. With the addition of GeneReleaser to the PCR reaction to function as a chelating resin, rpoB genes (serving as model genes) could be successfully amplified. The SW–EL method has potential to be of use for DNA–based analysis of microbial consortia present in heavy–oil contaminated soils.
Key words: DNA extraction, heavy oil, microcosms, PCR amplification, rpoB.
Resumen
Este trabajo describe un nuevo método de extracción de ADN genómico de suelo contaminado con petróleo pesado. El método combina lavados del suelo usando tres soluciones de lavado con lisis enzimática de las células (método SW–EL). El método SW–EL fue validado mediante la extracción de ADN de muestras de 0,5 g tomadas de microcosmos preparados con suelo artificialmente contaminado con petróleo pesado al 1 y 10% (p/p) e inoculados con un consorcio bacteriano. Comparado con un protocolo establecido de lisis y obtención de ADN usando el kit comercial Gene ReleaserTM (Bioventures) (método GRL), el nuevo método SW–EL permitió obtener mayor cantidad de ADN. Con la adición de Gene Releaser como resina quelante a la reacción de PCR, los genes rpoB (usados como modelo) fueron amplificados, indicando que el método SW–EL puede ser usado para analizar los consorcios microbianos presentes en suelos contaminados con petróleos pesados.
Palabras clave: extracción de ADN, amplificación PCR, microcosmos, petróleo pesado, rpoB.
INTRODUCTION
Characterization of soil microbial communities is often accomplished using molecular methods (Dallhof et al., 2000). These techniques are useful for overcoming the inability to culture most of the microorganisms found in environmental samples and the lack of sensitivity of traditional microbiological methods (Yeates et al., 1998). However, molecular methods are very susceptible to contaminants that may interfere considerably with DNA recovery from environmental samples. A common problem with DNA obtained from soil is the presence of humic and fulvic acids, which have similar size and charge characteristics as DNA, leading to the co–extraction of these compounds (LaMontagne et al., 2002; Yeates et al., 1998). These acids reduce the efficiency of DNA–DNA hybridization and inhibit Taq polymerase activity (Fortin et al., 2004; Zhou et al., 1996). DNA extraction from hydrocarbon–contaminated soils presents an even greater challenge because, in addition to the usual soil contaminants, traces of organic substances and heavy metals remain in the DNA extracts and reduce PCR amplification efficiency by degrading or capturing nucleic acids or by inactivating DNA polymerase (Fortin et al., 2004). Contaminants such as polycylic aromatic hydrocarbons (PAH's) and heavy metals like copper and lead (which are known to interfere with enzymatic activity) have often been found bound to clay and humic materials in hydrocarbon–contaminated soil (Fortin et al., 2004) and are often co–extracted with DNA. In order to study the diversity of a complex environment like a hydrocarbon–contaminated soil through culture–independent techniques, efficient protocols that eliminate these contaminants and yield good quality DNA must be developed.
Soil washing is a chemical treatment often used to solubilize hydrocarbons and heavy metals, in order to remove those contaminants from the soil. Washing solutions usually contain chelating or surfactant agents, acids, salts and/or redox substances (Dermont et al., 2008). EDTA is recognized as an effective chelating agent to extract metals such as aluminum (Al), copper (Cu), chromium (Cr), lead (Pb) and zinc (Zn) (Dermont et al., 2008; Elsan et al., 2006). EDTA can also enhance diesel fuel removal from diesel fuel–contaminated soil (Zhang et al., 2007). Triton X–100 can increase the solubilization of hydrophobic organic compounds such as PAH`s (Luning and Pritchard, 2002).
Existing DNA extraction methods employ different approaches to eliminate PCR inhibitors. Washing steps using hexadecylmethylammonium bromide (CTAB) and polyethylene glycol (PEG) precipitation are necessary to obtain better quality DNA (LaMontagne et al., 2002; Zhou et al., 1996). Washing steps with surfactants such as Triton X–100 and chelating agents prior to cell lysis improve the recovery of nucleic acids and are beneficial to obtain efficient PCR amplification of PAHs, PCBs and heavy metals contaminated sediments (Fortin et al., 2004). However, these steps tend to make DNA extraction protocols time–demanding, tedious and costly. Commercial extraction kits have been developed promising to alleviate inhibition problems with straight–forward and simplified sample processing procedures. GeneReleaserTM (BioVentures) accomplishes cell lysis, can remove up to 75 ng of humic acids (Menking et al., 1999) and sequesters PCR inhibitors increasing PCR yield and specificity simplifying DNA amplifications by avoiding the lengthy protocols of DNA purification.
In the past, our group has isolated several microbial consortia successfully used for large–scale bioremediation processes of heavy oil–contaminated soils. However, monitoring changes in bacterial communities during the biodegradation process has been hampered by limited technology. To further understand these bacterial communities using molecular methods such as PCR–DGGE, it would be desirable to develop an effective method for extracting amplifiable bacterial DNA from heavy oilcontaminated soil. The aim of this study was to compare two protocols: the first uses a combination of soil washes (Fortin et al., 2004) and enzymatic lysis (Yeates et al., 1998; Zhou et al., 1996), followed by standard purification steps using organic solvents. The second one is a modification of Marquez–Rocha et al. (2005), which includes treatment with GeneReleaserTM (BioVentures), a commercial product designed for release genomic DNA and, according to the fabricant, that allows cell lysis directly in the amplification tube on the thermocycler, sequesters cell lysis products which might inhibit polymerases and improves amplification yield and specificity. DNA quality and purity was validated by comparing the PCR amplification efficacies of the DNA obtained by the two methods. Results showed that the combination of soil washes and enzymatic lysis is suitable for DNA extraction from 0.5 g of petroleum–contaminated soil, resulting in good DNA yields. However, to obtain efficient PCR amplification, an additional DNA purification step was necessary, using a chelating agent in the PCR reaction.
MATERIALS AND METHODS
Bacterial strains and microcosms inoculation. The strains Pseudomonas sp. PS60 and Bacillus cereus group O20B used to inoculate the microcosms were isolated during a previous study from a hydrocarbon–contaminated soil in Casanare (Colombia). Both strains were identified on the basis of partial 16S rRNA gene sequence (data not published).
Microcosms were set up as follows: equal amounts (65 g each) of soil, sand and poultry manure were mixed and distributed into three different plastic containers; 1% (v/w) heavy oil–16oAPI from Casanare, Colombia (5º 30' 30.4' N, 71º 45' 37' W) was added to three microcosms, which were then separately inoculated with Pseudomonas sp. PS60, B. cereus O20B, or rested tap water (as control). 10 ml rested tap water were added to each microcosm to moisten and homogenize the system. Bacteria used for inoculation were grown on plates of Plate Count Agar PCA (Scharlau) at 30 °C for 16 hours; the confluent growth obtained from a plate was collected, and cell concentration was adjusted to obtain a final cell count of 5 x 108 CFU/g in the microcosms. The procedure was repeated for the other three microcosms containing 10% (v/w) heavy oil. All microcosms were kept at 30 ºC and samples of 0.5 g were taken at day 3 for DNA extraction and determination of bacterial culturable population in PCA medium.
Total DNA extraction. DNA extraction using SW–EL (Soil Washes–Enzymatic Lysis). This method combines two previously reported protocols, which were modified and will be referred to as the SW–EL protocol. Soil washes prior to cell lysis were modified from Fortin et al. (2004). A soil sample of 0.5 g was washed twice with 1 ml solution 1 (50 mM Tris–HCl pH 8.0, 200 mM NaCl, 5 mM Na2EDTA, 0.05% Triton X–100) by vortexing 2 min and centrifuging for 3 min at 3,000 x g. This was followed by washing with 1 ml solution 2 (50 mM Tris–HCl pH 8.0, 200 mM NaCl, 5 mM Na2EDTA) and two washes with solution 3 (10 mM Tris–HCl pH 8.0, 0.1 mM Na2EDTA).
The DNA extraction method was based on Yeates et al. (1998) and Zhou et al. (1996). One ml of extraction buffer [CTAB 1%, 100 mM Tris–HCl (pH 8.0), 100 mM sodium EDTA (pH 8.0), 1.5 M NaCl, proteinase K 0.05 mg/ml] was added to the soil pellets and incubated at 37 °C for 30 min with shaking at 180 rpm. SDS (sodium dodecyl sulfate) was added (0.2 ml; 10%), and the samples were incubated at 65 ºC for 90 min, and centrifuged at 6,000 x g for 10 min at room temperature.
The supernatants were collected, and the soil pellets were re–extracted with 1 ml of extraction buffer, incubated at 65 ºC for 10 min and centrifuged as above. Half–volume of polyethylene glycol (30%)/sodium chloride (1.6 M) was added to the supernatants, incubated at room temperature for 2 h and centrifuged at 10,000g for 20 min. The nucleic acid pellets were resuspended in 0.2 ml of TE (10 mM Tris–HCl, 1 mM sodium EDTA, pH 8.0), and potassium acetate was added to a final concentration of 0.5 M. Samples were transferred to ice for 5 min then centrifuged at 16,000 x g for 30 min at 4° C. The DNA was purified with phenol/chloroform 1:1 and chloroform/isoamyl alcohol 24:1 and precipitated with 0.6 volume of isopropanol and left overnight at –20 °C. DNA was pelleted by centrifugation at 16,000 x g for 30 min and resuspended in 20µl of ultrapure water. Three µl of purified DNA were loaded onto a 0.7% agarose gel stained with ethidium bromide and quantified using Gel–Doc Imaging SystemTM (BioRad) with High Mass LadderTM (Invitrogen, cat. 10496–016). The quality of the extracted DNA was evaluated via the absorbance ratios at 260nm/280nm and 260nm/230 nm.
DNA extraction using GRL (Gene Releaser Lysis). The other protocol we assayed was modified from Márquez–Rocha et al. (2005), and will be here referred to as the GRL protocol. Ten ml 0.2M NaCl were added to 0.5 g of soil and incubated at 30 oC for 15 min with shaking at 145 rpm. Particles were allowed to settle and 1 ml supernatant was centrifuged at 16,000 x g for 5 min. The pellets obtained from the last centrifugation step were first washed with 50 µl 0.2 M NaCl and then with 50 µl distilled water and resuspended in 20 µl TE buffer (10 mM Tris–HCl, pH 8.0; 1 mM EDTA, pH 8.0) and 20 µl GeneReleaserTM (BioVentures), and vortexed for 30 s. The samples were overlaid with 10 µl of sterile mineral oil and heated in a microwave oven for 2 min, then centrifuged for 30 s. The supernatant (DNA–containing fraction) was transferred to a sterile eppendorf tube. The quality and the concentration of the extracted DNA were analyzed as described for SW–EL protocol.
Preparation of crude extracts. Crude extracts of Pseudomonas sp. PS60 and B. cereus O20B were obtained from cultures in Luria–Bertani both incubated at 30 oC for 14h and 140 rpm, which were boiled for five min, and centrifuged. 0.9 µl from the supernatant was used as the source of DNA.
PCR amplification. DNA obtained with SW–EL protocol, GRL protocol, or Pseudomonas sp. PS60 and Bacillus cereus O20B crude extracts was used for PCR amplification of the rpoB gene. In the case of SW–EL method, 0.5 µl of either undiluted DNA, 1/10 or 1/50 dilutions were used as template with and without 4.5 µl of GeneReleaserTM (BioVentures), used here as a chelating agent according to Yeates et al. (1998); reactions were overlaid with 10 ml of sterile mineral oil. The tubes were heated on a microwave oven for 7 min. A beaker containing 100 ml of water was included as a microwave sink. Tubes were incubated for 10 min at 80 oC in MyCyclerTM (Bio–Rad) before adding the 20 µl PCR mix (to obtain a final reaction volume of 25ul). For the GRL method, following manufacturer's instructions, 24 µl PCR mix were added to 1 ml of the supernatant (DNA–containing fraction).
Primers rpoB1698f (5–AACATCGGTTTGATCAAC–3) and rpoB2014r (5–GTTGCATGTTGGTACCCAT–3) (Dahllöf et al., 2000) were used to amplify the rpoB gene. 25 µl reactions were prepared containing 0.2 mM each dNTP´s, 0.5 µM of each primer, 2.5 mM MgSO4, 2U Tucan Taq polymerase (Corpogen), 1X PCR buffer and 1X PCR Enhancer solution (Invitrogen).
The amplification program consisted of a denaturing step of 94 °C for 5 min, 6 cycles of denaturing for 30 s at 94 °C, annealing for 1.5 min at 40 °C, and a 1.5 min extension at 72 °C, followed by 19 cycles at 94 °C for 30 s, 1.5 min at 50 °C, and a 1.5 min at 72 °C. A final extension step of 72°C for 10 min was then performed (Dahllöf et al., 2000). 3 µl of PCR product were run on a 1.2% agarose gel containing ethidium bromide and visualized using the Bio–Rad Gel Doc imaging system.
Sequencing of PCR products. PCR products after rpoB gene amplification from the strains Pseudomonas sp. PS60 and B. cereus O20B were sequenced. PCR products were purified (Wizard SV gel and PCR clean–up systemTM, Promega) and sequenced using the BigDye Terminator v3.1 Cycle Sequencing kit (ABI Prism 310 sequencer, Applied Biosystems). Resulting sequences were analyzed in GeneBank using the Blast program (Altschul et al., 1997).
RESULTS AND DISCUSSION
Bacterial counts from microcosms in Plate Count agar were as follows: control microcosm 1% oil 2 x 107 CFU/g, microcosm 1% oil + B. cereus O20b 1.6 x 108 CFU/g, microcosm 1% oil + Pseudomonas sp. Ps60 9.6 x 107 CFU/g, control microcosm 10% oil 1.5 x 107 CFU/g, microcosm 10% oil + B. cereus O20b 1.6 x 108 CFU/g, microcosm 10% oil + Pseudomonas sp. Ps60 3.4 x 107 CFU/g. In all cases, and according to what was expected, seeded microcosms showed higher bacterial populations.
Many DNA extraction methods for soils are available, but very few are suitable for DNA extraction from hydrocarbon contaminated soils. DNA recovery from substrates contaminated with hydrocarbons typically requires more extensive purification strategies (Evans et al., 2004; Fortin et al., 2004; Kasai et al., 2005) and/or utilisation of purification resins (Márquez–Rocha et al., 2005) in order to obtain high molecular weight DNA free from inhibitors for PCR amplification. We used microcosms artificially contaminated with heavy petroleum to evaluate two DNA extraction protocols: the SW–EL protocol employed a series of soil washes before the enzymatic lysis of the cells; the GRL protocol, consisted in a step of NaCl washing followed by lysis using the commercial product GeneReleaserTM. We obtained better results with the SW–El protocol. With this method DNA was obtained from all the microcosms (table 1). In contrast, with the GRL protocol we recovered DNA only from the microcosms that were inoculated with bacteria and contained 10% heavy oil (table 2).
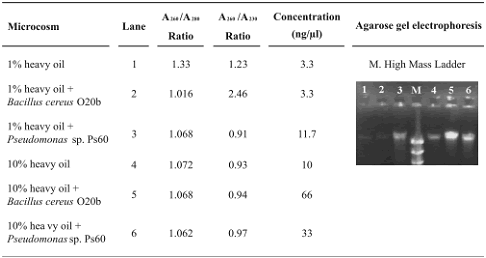
Table 1. DNA obtained from heavy oil–contaminated microcosms using the SW–EL (Soil Washes–Enzymatic Lysis) method
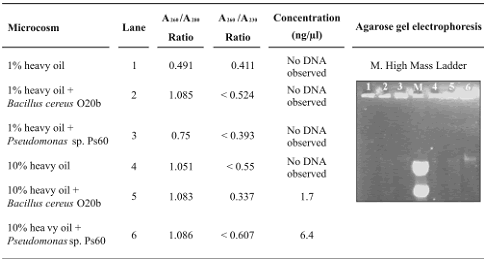
Table 2. DNA obtained from heavy oil–contaminated microcosms using the GRL (Gene Releaser Lysis)method
DNA quantities obtained with the SW–EL method were higher when compared with those obtained with the extraction protocol GRL (tables 1 and 2). The A260/A230 ratios for the DNA from SW–EL method were also better and the A260/A280 ratios were similar for both methods (tables 1 and 2). Higher A260/A230 ratios indicate that the DNA is contaminated with lower amounts of humic acid–like compounds that probably were removed via CTAB solution and ammonium acetate precipitation in protocol SW–EL (Fortin et al., 2004; Zhou et al., 1996). Washing steps increase the quantity and quality of the resulting DNA from soils (Fortin et al., 2004; Purohit et al., 2003); solution 1 used in the SW–EL protocol contains Triton X–100, which increases the solubility of oils (Fortin et al 2004), and probably allows their removal.
It is interesting that the 10% hydrocarbon microcosms resulted in a larger quantity of DNA than in 1% oil soil; these results cannot be attributed to higher bacteria populations because bacterial cell counts were similar in the two petroleum concentrations. Fortin et al. (2004) reported that when the washes were incorporated before the lysis, the amount of DNA recovered was reduced in the less contaminated sediments possibly due to the desorption of bacteria from the soil hydrocarbons (Stelmack et al., 1999).
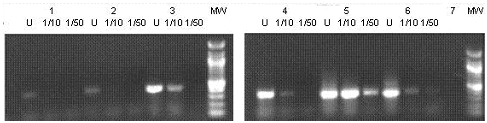
The presence of inhibitors in the recovered DNA was evaluated by amplification of a 368 bp rpoB fragment. rpoB PCR products were obtained from all undiluted DNA obtained with the SW–El protocol, but only when GeneReleaser was present in the PCR mix (figure 1); without GeneReleaser we did not obtain amplicons with undiluted DNA (we did obtain amplification using DNA diluted 1/10, data not shown). Failure to amplify undiluted DNA was probably due to the presence of inhibitors (PAHs, PCBs and heavy metals), according to previous results reported by Fortín et al. (2004), PCR amplification was only successful when DNA dilutions were used as templates.
Figure 1. Agarose gel electrophoresis of rpoB amplification products from undiluted (U), dilution 1/10 (1/10), or 1/50 (1/50) DNA obtained from oil–contaminated microcosms using the SW–EL (Soil Washes–Enzymatic Lysis) method. 1. 1% heavy oil; 2. 1% heavy oil + Bacillus. cereus O20b; 3. 1% heavy oil + Pseudomonas sp. Ps60.; MW. 100pb ladder. 4. 10% heavy oil; 5. 10% heavy oil + Bacillus. cereus O20b; 6. 10% heavy oil + Pseudomonas sp. Ps60; 7. Control without DNA; MW. 100pb ladder.
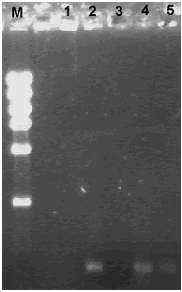
It was possible to amplify the 368 bp rpoB fragment from the DNA extracted using the GRL protocol (figure 2, lines 5 and 6); with this protocol the limitation was not the presence of inhibitors but the low amount of DNA obtained (table 2). Previously, Márquez–Rocha et al. (2005) obtained amplification of the 16S rRNA gene from all 13 soil samples (obtained from locations near to oil storage tanks in Mexico) assayed using the same protocol. In our case, maybe it is possible to enhance DNA recovery by increasing the amount of soil sample used.
The minimal DNA concentration required to obtain PCR products was 0.5 ng (we assayed 0.033–33 ng).The PCR product was sequenced to verify its identity. For all cases it corresponded to the rpoB gene sequence and confirmed the genera of the bacteria used (table 3).Lysis strategies should be selected based on the kind of DNA analyses that will follow the extraction; for PCR amplification of a particular gene the GRL method may be used, but for microbial diversity studies the SW–EL showed to be more reliable since it provides higher DNA amounts and lower humic acids contamination (represented by a lower 260/230 ratio).
Figure 2. Agarose gel electrophoresis of rpoB amplification products from DNA obtained from oil–contaminated microcosms using the GRL (Gene Releaser Lysis) method. M. DNA High mass ladder; 1. Control without DNA; 2. Bacillus cereus O20b; 3. 10% heavy oil; 4. 10% heavy oil + Bacillus cereus O20b; 5. 10% heavy oil + Pseudomonas sp. Ps60.
CONCLUSIONS
The combination of soil washes and enzymatic lysis is suitable for the DNA extraction from small samples of heavy petroleum contaminated soil (0.5 g) and, with the addition of a chelating resin, the rpoB gene could be amplified from landfarming treatments in order to determine the presence of the oil degrading bacteria by DGGE or other molecular analysis.
ACKNOWLEDGEMENTS
This work was supported by grants from the Research Committee of Science Faculty, Andes University. We thank Engineer Hugo Vladimir Ramirez and Perenco Limited S. A. for providing us with heavy oil from Oil Field Station, Casanare, Colombia.
REFERENCES
1. Altschul SF, Madden TL, Schäffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ. 1997. Gapped BLAST and PSI–BLAST: a new generation of protein database search programs. Nucleic Acids Research, 25(17):3389–3402. [ Links ]
2. Dahllöf I, Baillie H, Kjelleberg S. 2000. rpoB–based microbial community analysis avoids limitations inherent in 16S rRNA gene intraspecies heterogeneity. Applied and Environmental Microbiology, 66(8):3376–3380. [ Links ]
3. Dermont G, Bergeron M, Mercier G, Richer–Laflèche M. 2008. Soil washing for metal removal: A review of physical/chemical technologies and field applications. Journal of Hazardous Materials, 152(1):1–31. [ Links ]
4. Ehsan S, Prasher SO, Marshall WD. 2006. A washing procedure to mobilize mixed contaminants from soil. II. Heavy metals. Journal of Environmental Quality, 35(6):2084–2091. [ Links ]
5. Evans FF, Seldin L, Sebastian GV, Kjelleberg S, Holmström C, Rosado AS. 2004. Influence of petroleum contamination and biostimulation treatment on the diversity of Pseudomonas spp. in soil microcosms as evaluated by 16S rRNA based–PCR and DGGE. Letters in Applied Microbiology, 38(2):93–98. [ Links ]
6. Fortín N, Beaumier D, Lee K, Greer CW. 2004. Soil washing improves the recovery of total community DNA from polluted and high organic content sediments. Journal of Microbiological Methods, 56(2):181–191. [ Links ]
7. Kasai Y, Takahata Y, Hoaki T, Watanabe K. 2005. Physiological and molecular characterization of a microbial community established in unsaturated, petroleum–contaminated soil. Environmental Microbiology, 7(6):806–818. [ Links ]
8. LaMontagne MG, Michel FC, Holden PA, Reddy CA. 2002. Evaluation of extraction and purification methods for obtaining PCR–amplifiable DNA from compost for microbial community análisis. Journal of Microbiological Methods, 49(3):255–264. [ Links ]
9. Luning Prak D, Pritchard PH. 2002. Solubilization of polycyclic aromatic hydrocarbon mixtures in micellar nonionic surfactant solutions. Water Research, 36(14):3463–3472 [ Links ]
10. Márquez–Rocha FJ, Olmos–Soto J, Rosano–Hernández MC, Muriel–García M. 2005. Determination of the hydrocarbon–degrading metabolic capabilities of tropical bacterial isolates. International Biodeterioration & Biodegradation, 55(1):17–23. [ Links ]
11. Menking DE. Emanuel PA, Valdes JJ, Kracke SK. 1999. Rapid cleanup of bacterial DNA from field samples. Resources Conservation and Recycling, 27(1–2):179–186. [ Links ]
12. Purohit HJ, Kapley A, Moharikar AA, Narde G. 2003. A novel approach for extraction of PCR–compatible DNA from activated sludge samples collected from different biological effluent treatment plants. Journal of Microbiological Methods, 52(3):315–323. [ Links ]
13. Stelmack PL, Gray MR, Pickard MA. 1999. Bacterial adhesion to soil contaminants in the presence of surfactants. Applied and Environmental Microbiology, 65(1):163–168. [ Links ]
14. Yeates C, Gillings MR, Davison AD, Altavilla N, Veal DA. 1998. Methods for microbial DNA extraction from soil for PCR amplification. Biological Procedures Online, 1(1):40–47. [ Links ]
15. Zhang W, Tsang DCW, Lo IMC. 2007. Removal of Pb and MDF from contaminated soils by EDTA– and SDS–enhanced washing. Chemosphere, 66(11):2025–2034 [ Links ]
16. Zhou J, Bruns MA, Tiedje J. 1996. DNA Recovery from soils of diverse composition. Applied and Environmental Microbiology, 62(2):316–322. [ Links ]