Services on Demand
Journal
Article
Indicators
-
 Cited by SciELO
Cited by SciELO -
 Access statistics
Access statistics
Related links
-
 Cited by Google
Cited by Google -
 Similars in
SciELO
Similars in
SciELO -
 Similars in Google
Similars in Google
Share
Caldasia
Print version ISSN 0366-5232
Caldasia vol.36 no.1 Bogotá Jan./June 2014
https://doi.org/10.15446/caldasia.v36n1.43893
http://dx.doi.org/10.15446/caldasia.v36n1.43893
FREDY PALACINO-RODRÍGUEZ
ENRIQUE GONZÁLEZ-SORIANO
CARLOS EDUARDO SARMIENTO
Laboratorio de Artrópodos del Centro Internacional de Física, Universidad Nacional de Colombia. Departamento de Biología / Universidad El Bosque, Bogotá D.C., Colombia. odonata17@hotmail.com
Departamento de Zoología, Instituto de Biología, Universidad Nacional Autónoma de México, México D.F., México. esoriano@ibunam2.ibiologia.unam.mx
Laboratorio de Sistemática y Biología Comparada de Insectos, Instituto de Ciencias Naturales, Universidad Nacional de Colombia, Bogotá, Colombia. cesarmientom@unal.edu.co
ABSTRACT
Erythemis Hagen, 1861 shows a considerable variation in genitalic characters, body coloration and wing venation. Since it is known that these traits are affected by different kinds of selection that probably blur their phylogenetic signal, we chose the genus Erythemis as a model taxon to analyze and compare the phylogenetic signal of these and other morphologic characters. A cladistic analysis was performed using ten species of the genus plus another seventeen species of Libellulidae as outgroup. Characters were defined following standard criteria and were managed using the software DELTA. Tree search was performed with the software NONA. Partitioned and combined analyses were conducted. Character tracking of characters with ri=100 was used to identify synapomorphies. In agreement with the literature, color characters provided strong phylogenetic signal, meanwhile, genitalia characters offered no synapomorphies. We did not find any character that could support the monophyly of Erythemis. The only clade that has strong support from the morphologic set of characters is (E. vesiculosa, (E. simplicicollis, E. collocata)). Contrary to the results found in other Odonata, wing characters offered synapomorphies for some Erythemis clades.
Key words. Odonata, dragonfly, phylogenetic signal, male genitalia, body coloration.
RESUMEN
Erythemis muestra una considerable variación en caracteres de genitalia, coloración del cuerpo y venación alar. Estos caracteres están afectados por diferentes tipos de selección, lo que puede desdibujar su señal filogenética, por lo que nosotros elegimos el género Erythemis como taxón modelo para analizar y comparar la señal filogenética de éstos y otros caracteres morfológicos. Un análisis cladístico se realizó con las diez especies del género más otras 17 especies de Libellulidae como grupo ajeno. Los caracteres se definieron siguiendo criterios de estandarización y fueron manejados con el software DELTA. La búsqueda de árboles fue ejecutada con el software NONA. Se adelantaron análisis particionados y análisis combinados. El rastreo de caracteres con ri=100 se usó para identificar las sinapomorfías. En coincidencia con la literatura, los caracteres de color proveen fuerte señal filogenética mientras que los caracteres de genitales no ofrecieron sinapomorfías. No se encontró ningún caracter que soporte la monofilia del género. El único clado con fuerte soporte es (E. vesiculosa, (E. simplicicollis, E. collocata)). Contrario a lo reportado para otros Odonata, la venación alar arrojó sinapomorfías para algunos clados de Erythemis.
Palabras clave. Odonata, libélula, señal filogenética, genitales del macho, coloración corporal.
Recibido: 25/06/2013
Aceptado: 30/04/2014
INTRODUCTION
The genus Erythemis Hagen, 1861, is composed by ten species distributed in the Neotropical and Neartic regions, which are found from sea level to 2300 masl. Some species within the genus show territorial behavior and tolerate high temperatures (McVey, 1981), males exhibit continuous signals of interspecific aggression during mating and hunting (Baird & May, 2003). Several authors have studied the phylogenetic relationships in Odonata using different data sets; of these, only a few have included Erythemis in their analysis, but no more than one species of the genus has been included (e.g. Ware et al., 2007; Pilgrim & Von Dohlen, 2008). Specific studies on phylogenetic relationships among Erythemis species, were conducted by Kennedy (1923) and Pinto (2008). Kennedy (1923) established a relationship among E. vesiculosa, E. collocata and E. simplicicollis based on the absence of the posterior lobe of the vesica espermalis. Likewise, this author proposed the grouping of E. peruviana, E. mithroides, and E. attala, separating them from the group E. plebeja, E. carmelita and E. haematogastra considering the narrower abdomen of this last group. Unfortunately, the data of Pinto (2008) have not been published and the characters worked by him are not known.
The phylogenetic signal of a character has been an important topic in systematics, which began for the interest on the evolutionary phenomena that may affect it ( Wilson , 1975). Currently, the phylogenetic signal is a topic used to describe the tendency of related organisms to resemble each other without implications about the mechanisms that might cause it (Blomberg, et al., 2003), and it can be described as the number of homologies that may be found in a particular character set. The amount of phylogenetic signal that provides different systems of characters may depend on the selection pressures and evolutionary rates that the character experiences. For example, some studies on genital characters, across several groups of insects, suggests that their evolution could have been faster due to sexual selection (Córdoba-Aguilar, 2005), and this phenomenon may blur the phylogenetic signal of these characters in comparison with other characters that are not under those selective pressures.
The phylogenetic signal of a character set (a group that includes all the characters of a particular corporal region, i.e. wings or thorax) can be analyzed in two ways: 1. A separate analysis of each character set can be conducted and the consensus analysis between the trees obtained may indicate the level of congruency between each proposal; it has been argued that in this way the properties and the selective pressures of each character set are included in every analysis and are shown by the tree that better reflects the information in each analysis (Kluge, 1989). For instance, odonate wings are under natural selection, related to the aerodynamics of the flight (Kesel, 2000) while odonate genitalia and coloration may be under the selective pressures of species recognition processes and sexual selection (Córdoba-Aguilar & Cordero, 2008), thus, the phylogenetic behavior of those character sets may be different. 2. A combined analysis can be performed and the behavior of each character set is compared; it is believed that this approach maximizes the explanatory power of the characters and may conduct a more rigorous test of homology for the characters (Nixon & Carpenter, 1996). In addition, given the phylogenetic signal of different character sets may add to the solution of conflicts in these analyses, polytomies may become less frequent (Kluge, 1989; Kluge & Wolf, 1993).
A priori down weighting or character removal is frequently used (Wiens, 1995). However, it has been proven that supposedly unreliable characters (i.e. genitalia or coloration) may provide phylogenetic signal, meanwhile character sets traditionally considered reliable, may provide lower phylogenetic signal (Areekul & Quicke, 2006, Song & Bucheli, 2010). In the present study a phylogenetic analysis of the genus Erythemis was conducted to: 1) compare the phylogenetic signal of genitalia and color characters with those of other groups of characters, 2) test whether Erythemis is a monophyletic taxon, and 3) propose a phylogenetic hypothesis of relationships among Erythemis species.
MATERIALS AND METHODS
Taxa
The analysis included 27 species, the ten currently recognized species of Erythemis as ingroup, and 17 species as outgroup, those species were selected according to previous phylogenetic hypotheses (e.g. Pilgrim & Von Dohlen, 2008) and to include representatives of other Libellulidae subfamilies. The species included in this study were: Subfamily Sympetrinae Erythemis attala (Selys in Sagra), E. carmelita Williamson, E. collocata (Hagen), E. credula (Hagen), E. haematogastra (Burmeister), E. mithroides (Brauer), E. peruviana (Rambur), E. plebeja (Burmeister), E. simplicicollis (Say), E. vesiculosa (Fabricius), Rhodopygiacardinalis (Erichson), R. geijskesi Belle, R. hinei Calvert, R. hollandi Calvert, R. pruinosa Buchholz; Subfamily Leucorrhininae Brachymesia herbida (Gundlach); Subfamily Libellulinae Garrisonia aurindae Penalva & Costa, 2007, Libellula herculea Karsch; Subfamily Palpopleurinae Perithemislais (Perty), Perithemis mooma Kirby, Perithemis thais Kirby, Perithemis tenera (Say); Subfamily Trameinae Miathyria marcella (Selys), Pantala flavescens (Fabricius), Tramea calverti Muttkowski, and T. rustica De Marmels and Racenis. Considering the work of Pilgrim & Von Dohlen (2008) Rhodothemis rufa Rambur was used for rooting purposes.
In order to record character variation a total of 3,000 specimens from the following entomological collections were studied. Their acronyms follow Evenhuis & Samuelson (2004) and are provided within brackets: Colección de Entomología, Universidad de los Andes, Bogotá, Colombia (ANDES), Laboratorio de Colecciones Entomológicas de la Universidad de Antioquia (CEUA), Colección Nacional de Insectos, Universidad Nacional Autónoma de México, México D. F., México (CNIN), Florida State Collection of Arthropods, Gainesville, Florida, USA (FSCA), Instituto de Ciencias Naturales, Universidad Nacional de Colombia, Bogotá D.C. (ICN), Museum of Comparative Zoology, Harvard University, Cambridge, Massachusetts, USA (MCZ), Museo de Entomología Francisco Luis Gallego, Universidad Nacional de Colombia, Medellín, Colombia (MEFLG), Museu Nacional, Universidade do Rio Janeiro, Rio de Janeiro, Brazil (MNRJ), Museo de Entomología de la Universidad del Valle, Cali, Colombia (MUSENUV), Rosser W. Garrison personal collection, Sacramento, California, USA (RWG), Museo de Colecciones Biológicas-Universidad del Atlántico Región Caribe (UARC), and Museum of Zoology, University of Michigan, Ann Arbor, Michigan, USA (UMMZ).
Character coding and cladistic analyses
The definition of the characters follows the parameters proposed by previous authors (Vogt et al., 2010); in most cases, the functional components of the character follows Sereno (2007) considering characters as properties of the species observed in the organisms expressed as independent variables with exclusive states. The "absent" state was only considered for neomorphic characters in the sense of a "substance" which is either present or absent in any structure (Sereno, 2007). Morphological terminology follows Borror (1942), Riek & Kukalová-Peck (1984), and Garrison et al. (2006). The characters were recorded in a matrix (Supplementary Material) using the DELTA package (Descriptive Language for Taxonomy) (Dallwitz, 2000). Specimens were examined using stereomicroscope and Scanning Electron Microscopy at low voltage (25-30kV). Gold-coated structures were observed using a Scanning Electron FEI Quanta200 microscope.
Diagnostic characters should be synapomorphies as they should be restricted to the species belonging to a specific taxon (i.e. genus); once a diagnostic character is present in other taxa, its value is questionable. The diagnosis of the genus Erythemis (Garrison et al., 2010) was based on a combination of characters and none of them are unique to Erythemis species. Three of these characters were coded with minor adjustments, to fulfill with character definition criteria described above, these were: origin of CuP in HW attached to posterior angle of triangle (character 93), posterior border of vulvar lamina rounded or acute or truncated (119), and posterior hamule bifid (122). Body color was alternatively coded as presence/absence of pigments (coding 1, Table 1), or as presence/absence of color patterns such as spots or stripes on the skeleton (coding 2, Table 1). Partitioned analyses were conducted to test the effect of these coding schemes. The character posterior femur widened and with 3-4 robust spines located at the external angle of the distal region, as described by Garrison et al. (2010), generates several non-exclusive character states violating the exclusivity principle defined by Sereno (2007). Thus, we proposed seven characters considering separate qualities in each such as femur width, spines thickness, number, size, distribution pattern, and location of spines (characters 69-71, 73, 74, 76, 77).
A total of 131 characters were coded (Table 1): 15 characters belong to the abdomen, thorax, and legs, 34 to the wing venation, 15 to the genitalia (vesica spermalis; vulvar lamina, and cerci), and 67 were color characters. Due to high intraspecific variation, the following five characters were not included in the phylogenetic analyses: Number of postnodal veins between costa and radio veins, previous to first postnodal vein between radio and M1 veins in FW (100), Number of postnodal veins between costa and radio veins previous to first postnodal vein between radio and M1 veins in HW (101), Number of cells between A1 and anal angle in HW (102), Number of rows of cells between MA and Mspl in FW (103), and Number of cells in the anal keel bifurcation in HW (107). Williamson (1923) proposed the character widening of the abdominal basal region with different states to separate some species in his key, however, such definition of the character did show high overlapping between states and no species separation, for this reason this character was recoded (character 80). Some characters correspond to alternative coding strategies to test their effect on the phylogenetic analysis (see table 1).
All the characters were coded as non-additive. Missing data were indicated by a question mark ("?") and inapplicable data were indicated by a hyphen mark ("-"). Phylogenetic analyses were done using the parsimony software NONA under the WinClada package10.00.08 (http://www.cladistics.com/). Given the number of taxa (27), heuristic search based on the Ratchet algorithm (Nixon, 1999) with 10% of characters resampled was applied. Ten trees were retained per replicate and tree-bisection-reconnection (TBR) and branch swapping with the default options of the software were used.
For an assessment of tree search thoroughness, we repeated tree search increasing repetitions up to 100,000. Once every search was completed the number of fundamental trees, their length, Ci and Ri were recorded. If the number of fundamental trees did not increase with replications, this was considered as an indication of exhaustively sampled space tree. However, since the number of fundamental trees may increase as replication increases, due to some clades where no further resolution can be reached with the current data set, we identified these cases by comparing the strict consensus trees of every replicate (Table 2). Strict consensus trees were used in every analysis as a summary of the congruent information obtained from the fundamental trees (Nixon & Carpenter, 1996). We only used characters with retention index of 100 as support for specific clades. This value appears if no trace of homoplasic interpretations can be observed in a character (Patterson, 1982; Farris, 1989a). A flowchart of the procedure described here is presented in figure 1.
Comparison of the phylogenetic signal of different character sets
Characters were grouped into the following sets: wing venation, thorax-legs-abdomen, genitalia, and body coloration. These character sets may be susceptible to different selection pressures. For example, the wing venation is exposed to aerodynamic conditions and thus to natural selection pressures (Kesel, 2000), while genitalia and coloration may be subject of sexual selection (Córdoba-Aguilar & Cordero, 2008). Separate and combined or simultaneous phylogenetic analyses were conducted. The strict consensus tree from the combined analysis using the pigment coding strategy (coding 1) was used as reference, given that a higher number of characters provided a more severe test of homology (Kluge, 1989; Kitching et al., 1998) and in some cases these allowed the recovery of hidden homologies within a character subset (Nixon & Carpenter, 1996). In addition, as it is shown in the results section below, this tree presented higher resolution and retention index
The phylogenetic signal of a character set was analyzed by looking at the retention index of each tree. This index has been traditionally used as a general descriptor of the phylogenetic signal in a tree as this is not affected by matrix size (Farris, 1989a; Farris, 1989b; Kitching et al., 1998; Klingenberg & Gidaszewski, 2010). In this study the character sets ranged in size from 15 characters in the genitalia set up to 112 characters in the combined evidence analysis using the color pigment coding strategy. We also traced each character with retention index of 100 on both, its own subset tree, and on the combined analysis tree. In the latter, their assignation of a character subset was recorded; this strategy identifies the phylogenetic signal of that subset in the context of a more stringent dataset (Farris, 1989a; Song & Bucheli, 2010). A third approach to quantify the informativeness of each character set was recording the percentage of homologies with retention 100, respect to the total number of characters in both partitioned and combined analyses.
RESULTS AND DISCUSSION
Tree search
The analyses with the abdomen-legs-thorax character subset and the combined data set reached a maximum of trees that did not changed after 10,000 and 5,000 replications respectively (Table 2). In the analyses with the character subsets genitalia, wings, and color, the number of trees always increased with the number of replications (Table 2); however, the topology of the strict consensus trees of each replication were identical within these character subsets, indicating that the changes in the number of fundamental trees of each replication were the result of polytomies, where no characters allow subtree resolution. These results lead us to conclude that tree search was thorough in all the character subsets and in the total evidence analyses.
Character coding
The consensus tree from the combined pattern presence/absence coding strategy (coding 2, Fig. 2) presented lower resolution than that of the tree from the combined pigment presence/absence coding strategy (coding 1, Fig. 3). In the latter, several species of Erythemis appear in a single clade, the genus Rhodopygia appeared as monophyletic, and it is the sister group of a large clade that includes species of several genera. Nine characters with retention of 100 appeared on this tree. Similarly, when comparing both color dataset codifications, there was a large difference between the two strategies; the tree from the pattern coding was highly unresolved, and with a single clade (E. vesiculosa, (E. simplicicollis, E. collocata)) (Fig. 4) that is also present in the tree from the pigment coding (Fig. 5). The latter was a more resolved tree. The retention index of both coding strategies was very similar (Table 3).
It has been proposed that proper coding of characters is a crucial step in phylogenetic research especially when using morphologic data, and the compliance with basic requirements of character definition, such as independence, exclusivity, and logical standardization, must be addressed (Sereno, 2007; Vogt et al., 2010). In this study we found a good example of the importance of these requirements; when coding color characters as pattern, or strategy coding 1, these show lower resolution than the pigment coding, or strategy coding 2, analyzed as separate datasets or in the combined analyses.
Partitioned analyses, combined analyses, and phylogenetic signal
The abdomen-legs-thorax and the genitalia subsets offered higher retention indexes (82 and 64 respectively) while the wing venation subset offered lower retention index (Table 3). The retention index values of the two coding strategies for the color subset were a bit higher than those of the total evidence analyses using the two coding strategies (Table 3). The genitalia subset provided a highly unresolved tree with the single clade (Erythemiscollocata,E. simplicicollis) (Fig. 6) supported by the character of suboval shape of the vesica spermalis hook (130, ri=100).
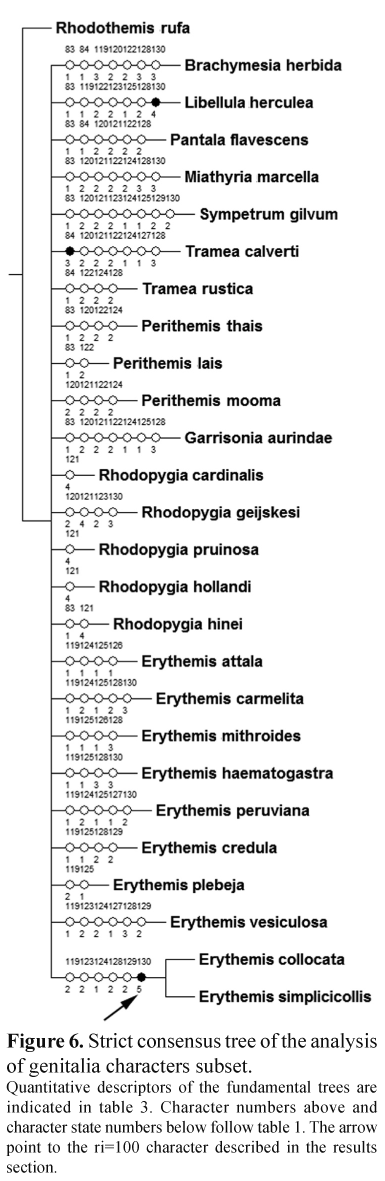
The extensive analysis of the genitalia of several species through Scanning Electron Microscopy revealed a large complexity of structures not observed before but unfortunately their coding was difficult due to variation. A similar situation occurred with the abdomen-legs-thorax subset, where only a clade (Erythemis mithroides,E. haematogastra) was found (Fig. 7).
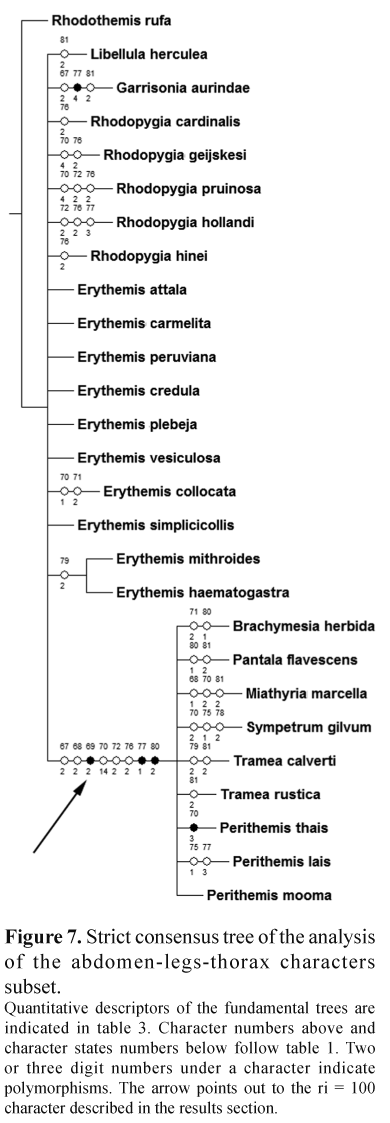
The wing veins subset offered a tree where most of the Erythemis species are located in a large basal polytomy and others are in other sections of the tree (Fig. 8). The presence/absence color pigment subset offered a mostly resolved tree with five polytomies, four of these are composed by three branches while one includes seven branches (Fig. 5); the clades are a mixture of species from different genera. Two characters had a 100 ri value and support the clade (E. vesiculosa, (E. collocata, E. simplicicollis)), these characters refer to the presence of green and red pigment on the epiproct (47 and 49 respectively).
The strict consensus of the 12 fundamental trees found from the combined analysis using the presence/absence of pigment coding strategy presented a basal polytomy composed by Libelulla herculea, four species of Erythemis and two large clades, one which included the other six species of Erythemis, and a large clade with species of several genera (Fig. 3), no characters with 100 ri could be traced at the node of the Erythemis species (Fig. 3). A total of nine characters were found with ri = 100, One belong to the color pattern subset (15), four belong to color pigments character subset (41, 45, 48, 50), two to the wing veins subset (86, 109), one to the thorax-legs-abdomen subset (68), and one to the genitalia subset (122). None of these characters were recovered as synapomorphies with ri = 100, in the analyses using the separate subsets of characters. Only three of the six clades observed in the analysis of the color pigment coding subset were present in the combined analysis.
The thickened long spines in the hind femur present in Erythemis, are also present in the genus Rhodopygia, in the species Libelulla herculea, Rhodothemis rufa and in Garrisonia aurindae. The disposition of the long spines in the external angle of the posterior femur exhibits a large array of variation in the species studied and even variation within species was recorded. The number of long spines in the external angle of the posterior femur also shows large variability and species such as E. haematogastra and E. credula had specimens with a lower or higher number of long spines to those proposed as diagnostic of the genus. In addition, species of other genera such as Perithemis, Rhodopygia, and Libellula exhibit between 3 and 4 long spines in the hind femur. The widened hind femur is also present in Libellula herculea Karsch, 1889 and Garrisonia aurindae Penalva & Costa, 2007 (Penalva & Costa, 2007).
Despite the debate about the use of either combined or partitioned analyses in phylogenetic studies (Lecointre & Deleporte, 2004; Nixon & Carpenter, 1996), our analysis is in agreement with the first as the trees of the combined analyses are more informative than these of the partitioned analyses and also present a larger number of synapomorphies. Moreover, the combined analyses uncover nine homologies that were not observed in the partitioned analyses. Another result that agrees with the literature (Wenzel & Siddall, 1999) points out to the lack of additivity of characters in phylogenetic studies; despite that the color characters were the more abundant of the entire data set (58%), that the color characters were five of the nine synapomorphies found in the combined analysis, and that the phylogenetic analysis of the color pigment subset provided the more resolved tree, this tree agreed only in six out of the 19 nodes observed in the combined analysis. In addition, none of the four characters with ri=100 in the subset analyses was observed as such in the combined analysis. Thus, the role of a character subset and that of a character can only be understood once the analysis is conducted to detect hidden synapomorphies (Nixon & Carpenter, 1996). A single character from the genitalia subset (122) was recovered as synapomorphy in the combined analysis; this result differs from these found by other authors (e.g., Song & Bucheli, 2010) who surveyed a large number of studies and concluded that genitalia characters can be as useful to phylogenetic analysis as any other character set, but they suggest a careful examination in every study. In the present case, the observed variation is expressed as homoplasy in different lineages, agreeing with the low informativeness of the genitalic region, as a consequence of accelerated and divergent sexual selection pressures (Méndez & Córdoba-Aguilar, 2004; Song & Wenzel, 2008; Song & Bucheli, 2010).
The characters from the abdomen-legs-thorax subset offered a highly unresolved tree; however, one of these characters appeared as a homology supporting a clade in the combined analysis (Fig. 3). Because odonate wing venation is complex and full of autopomorphies (Rehn, 2003), the set of wing characters of Erythemis provided a mostly unresolved tree (Fig. 8); however, two characters of this set appeared as homologies in the combined analysis (Fig. 3). Our results do not entirely comply with other authors (e.g., Rehn, 2003; Pilgrim & Von Dohlen, 2008), who proposed that wing venation is a highly variable region and provides very poor phylogenetic information. Despite the strong selection pressures that flight performance exerted over these structures (Kesel, 2000), homologies were recovered from these structures.
Even though Kennedy (1923) proposed the widening of basal region of the abdomen to establish species groups for the genus Erythemis, an analysis of body proportions of this region performed by the authors (unpublished data), showed that its high variation do not allow to recognize the discontinuity and therefore the character states can not be acknowledged. The relation between E. simplicollis and E. collocata proposed by Kennedy (1923) based on the absence of posterior lobe in the vesica spermalis, was corroborated by this study, but using the shape of the hook of the vesica spermalis.
Despite that color varies intraspecifically due to environment, ontogeny, and diet (Winston, 1999), and that museum specimens are often discolored, our results agreed with others who provided evidence that color characters may be useful for phylogenetic analysis in several insect groups (Areekul & Quicke, 2006). These results supports that color characters are involved in strongly conserved patterns (Song & Bucheli, 2010), perhaps as a consequence of their role on sexual recognition in Erythemis, doing that color characters may show a strongly structured evolution as a whole, that may lead to a strong phylogenetic signal (Song & Bucheli, 2010). As it was demonstrated above, coding is important when including traits, to avoid violations to logic precepts in the characters such as character interdependence, conjunction of character states, or character correlation (Sereno, 2007). The results on wing and color character subsets also points at the importance of looking at the data before proceeding with preventive subtraction (Wenzel & Siddall, 1999).
As it has happened in other odonate taxa (e.g. Dijkstra & Vick, 2006; Ware et al., 2007; Pilgrim & von Dohlen, 2008; Blanke et al., 2013), Erythemis was not found as a monophyletic group due to the extensive homoplasy and structural variability observed in its diagnostic characters (Dijkstra et al., 2014). In Ertythemis case, aside from the high intra and interspecific variation that most of the characters showed, a large number of the character states are shared with other genera.
Some authors have approached to the high variation and complexity of Odonate morphology (e.g. Pilgrim & Von Dohlen, 2008) and they have studied wing venation along with many autopomorphies (Rehn, 2003), showing that the developmental process as larvae may influence this variation (Martinov, 1930). In addition it has been proposed that this variation, might respond for the strong differences in the capability of wing flexion among some odonates such as Aeshna Fabricius, 1775 and Pachydiplax Brauer, 1868 (Combes & Daniel, 2003).
The Erythemis morphology may be an example of the interaction between stochastic evolutionary processes altering the genetic homogeneity of the species (Clegg et al., 2002) and the adaptation to habitat heterogeneity inhabited by their species. Studies in other odonates have suggested that selective pressures such as landscape structure (Taylor & Merriam, 1995), food and predation stress (Svensson & Friberg, 2007), wind and high acidification of the larvae biotopes (Marinov & McHugh, 2010), and sexual selection (Outomuro & Johansson, 2011), can affect the evolution of wing and abdomen characters. For example, Johansson & Samuelson (1994) found that the action of predators might influence the length of dorsal and lateral thorns in Leucorrhinia dubia (Vander Linden, 1825) larvae. Giacomini & De Marco Jr. (2008) found a relationship among the variation of body length in larvae of several Anisoptera species and the habitat portion used by these. The authors stated that species like E. peruviana shows a narrower abdomen associated to the possibility of easy camouflage in macrophytes as a defense against predators. According to Giacomini & De Marco Jr. (2008) the presence and reproduction of the organisms, is related to the variation of their morphology with the environment and its usage that they might do of their habitat.
ACKNOWLEDGEMENTS
We are grateful to the following people for access to different entomological collections: Rosser W. Garrison (RWG), Emilio Realpe and Melissa Sánchez (ANDES-E), Martha Wolff and Cornelio Bota (CEUA); Rafael Borja and León A. Pérez (UARC), Sergio Orduz and John Jairo Quiroz (MEFLG), Nancy Carrejo, Carmen E. Posso and Christian Bermúdez (MUSENUV), Bill Mauffray (FSCA), Janira M. Costa (MNRJ), Philip D. Perkins (MCZ), and Mark F. O'Brien (UMMZ). We thank Natalia von Ellenrieder for her corrections and suggestions on this manuscript. Two referees are thanked for useful advice. We thank the Laboratorio de Sistemática y Biología Comparada de Insectos, and Laboratorio de Artrópodos del Centro Internacional de Física, Universidad Nacional de Colombia, Departamento de Biología, Universidad El Bosque. This research was supported by a fellowship from Departamento Administrativo de Ciencia, Tecnología e Innovación COLCIENCIAS, and by the Dirección de Investigación-Facultad de Ciencias, Universidad Nacional de Colombia (Bogotá Branch). We thank the electronic microscopy laboratory at the Universidad Nacional de Colombia for support with the SEM images.
LITERATURE CITED
1. Areekul, B. & D.L.J. Quicke. 2006. The use of colour characters in phylogenetic reconstruction. Biological Journal of the Linnean Society 88: 193-202. [ Links ]
2. Baird, J.M. & M.L. May. 2003. Fights at the dinner table: agonistic behavior in Pachydiplax longipennis (Odonata: Libellulidae) at feeding sites. Journal of Insect Behavior 16: 189-216. [ Links ]
3. Blanke, A., C. Greve, R. Mokso, F. Beckman & B. Misof. 2013. An updated phylogeny of Anisoptera including formal convergence analysis of morphological characters. Systematic Entomology 38: 474-490. [ Links ]
4. Blomberg, S. P., T.JR. Garland & A.R. Ives. 2003. Testing for phylogenetic signal in comparative data: behavioral traits are more labile. Evolution 57(4): 717-745. [ Links ]
5. Borror, D. 1942. A revision of the Libellulinae genus Erythrodiplax Odonata. Contributions in Zoology and Entomology, No. 4. Biological Series. The Ohio State University , USA . [ Links ]
6. Clegg, S.M., S.M. Degnan, C. Moritz, A. Estoup, J. Kikkawa & I.P.F. Owens. 2002. Microevolution in island forms: the roles of drift and directional selection in morphological divergence of a passerine bird. Evolution 56: 2090-2099. [ Links ]
7. Combes, S.A. & T.L. Daniel. 2003. Flexural stiffness in insect wings. I. Scaling and the influence of wing venation. The Journal of Experimental Biology 206: 2979-2987. [ Links ]
8. Córdoba-Aguilar, A. 2005. Possible coevolution of male and female genital form and function in a calopterygid damselfly. Journal of Evolutionary Biology 18: 132-137. [ Links ]
9. Córdoba-Aguilar, A. & A. Cordero. 2008. Cryptic female choice and sexual conflict. pp. 189-202. In: A. Córdoba Aguilar. (ed.). 2008. Dragonflies and Damselflies: Model organisms for ecological and evolutionary research. Oxford University Press, Oxford . [ Links ]
10. Dallwitz, M.L. 2000. Onwards. Data requirements for naturallanguaje descriptions and identification. http://deltaintkey.com (Consulted: September 12 2010). [ Links ]
11. Dijkstra, K.D.B. & G.S. Vick. 2006. Inflation by venation and the bankruptcy of traditional genera: the case of Neodythemis and Micromacromia, with keys to the continental African species and the description of two new Neodythemis species from the Albertine Rift (Odonata: Libellulidae). International Journal of Odonatology 9, 51-70. [ Links ]
12. Dijkstra, K.D.B., V.J. Kalkman, R.A. Dow, F.R. Stokvis & J. Van Tol. 2014. Redefining the damselfly families: a comprehensive molecular phylogeny of Zygoptera (Odonata). Systematic Entomology 39: 68-96. [ Links ]
13. Evenhuis, N.L. & G.A. Samuelson. 2004. Abbreviations for Insect and Spider Collections of the World. http://hbs.bishopmuseum.org/codens/codens-inst.html (accessed 17 September 2011). [ Links ]
14. Farris, J. S. 1989a. The retention index and homoplasy excess. Systematic Zoology 38: 406- 407. [ Links ]
15. Farris, J.S. 1989b. The retention index and rescaled consistency index. Cladistics 5: 417-419. [ Links ]
16. Garrison, R.W., N. Von Ellenrieder & J.A. Louton. 2006. Dragonfly genera of the New World , an illustrated and annotated key to the Anisoptera. The Johns Hopkins University Press. USA . 384 pp. [ Links ]
17. Garrison R.W., N. Von Ellenrieder & J.A. Louton. 2010. Damselfly genera of the New World . An Illustrated and Annotated Key to the Zygoptera. The Johns Hopkins University Press, Baltimore , xiv + 490 pp, + 24 color plates. [ Links ]
18. Giacomini, H.C. & P. De Marco Jr. 2008. Larval ecomorphology of 13 Libellulidae (Anisoptera, Odonata) of the Middle Rio Doce Valley, Minas Gerais, Brazil. Brazilian Journal of Biology 68(1): 211-219. [ Links ]
19. Johansson, F. & L. Samuelson. 1994. Fish-induced variation in abdominal spine length of Leucorrhinia dubia (Odonata) larvae? Oecologia 100: 74-79. [ Links ]
20. Kennedy, C.H. 1923. The phylogeny and the distribution of the genus Erythemis (Odonata). Miscellaneous Publications of the Museum of Zoology University of Michigan 11: 19-21. [ Links ]
21. Kesel, A. B. 2000. Aerodynamic characteristics of dragonfly wing sections compared with technical aerofoils. The Journal of Experimental Biology 203: 3125-3135. [ Links ]
22. Kitching, I. , P. Forey, C. Humphries & D. Williams. 1998. Cladistics, the theory and practice of parsimony analysis. The systematics association publication No. 11. Oxford University Press. 226pp. [ Links ]
23. Klingenberg, C. & N. Gidaszewski. 2010. Testing and Quantifying Phylogenetic Signals and Homoplasy in Morphometric Data. Systematic Biology 59(3): 245-261. [ Links ]
24. Kluge, A. G. 1989. A concern for evidence and a phylogenetic hypothesis of relationships among Epicrates (Boidae, Serpentes). Systematic Zoology 38: 7-25. [ Links ]
25. Kluge, A. G., & A.J. Wolf. 1993. Cladistics: What's in a word? Cladistics 9: 183-199. [ Links ]
26. Lecointre G. & P. Deleporte. 2004. Total evidence requires exclusion of phylogenetically misleading data. Zoologica Scripta 34: 101-117. [ Links ]
27. Marinov, M. & P. Mchugh. 2010. Comparative study of the Chatham Islands Odonata: Morphological variability, behaviour and demography of the endemic Xanthocnemis tuanuii Rowe, 1987. International Dragonfly Fund-Report 30: 1-44. [ Links ]
28. Martynov, A.V. 1930. Wing venation of the Odonata and Agnatha. The interpretation of the wing venation and tracheation of the Odonata and Agnatha. Psyche 37: 245-280. [ Links ]
29. Mcvey , M.E. 1981. Lifetime reproductive tactics in a territorial dragonfly, Erythemis simplicicollis (Odonata, Libellulidae). Anonymous. Anonymous. Rockefeller University . 1147, OA 6068. Ph. D. Thesis. [ Links ]
30. Méndez, V. & A. Córdoba-Aguilar. 2004. Sexual selection and animal genitalia. Trends in Ecology and Evolution 19: 224-225. [ Links ]
31. Nixon, K.C. 1999. The Parsimony Ratchet, a New Method for Rapid Parsimony Analysis. Cladistics 15: 407-414. [ Links ]
32. Nixon, K.C. & J.M. Carpenter. 1996. On consensus, collapsibility and clade concordance. Cladistics 12: 305-321. [ Links ]
33. Outomuro, D. & F. Johansson. 2011. The effects of latitude, body size, and sexual selection on wing shape in a damselfly. Biological Journal of the Linnean Society 102: 263-274. [ Links ]
34. Patterson, C. 1982. Morphological characters and homology. pp. 21-74. In: K. A. Joysey & A.E. Friday. (eds). Problems of Phylogenetic Reconstruction. Academic Press, London and New York . [ Links ]
35. Penalva, R. & J. M. Costa. 2007. Garrisonia aurindae gen. and spec. nov. from the State of Bahia , Brazil (Anisoptera: Libellulidae). Zootaxa 1453: 33-40. [ Links ]
36. Pilgrim, E.M. & C.D. Von Dohlen. 2008. Phylogeny of the Sympetrinae (Odonata: Libellulidae): further evidence of the homoplasious nature of wing venation. Systematic Entomology 33: 159-174. [ Links ]
37. Pinto, A. 2008. Análise cladística de Erythemis Hagen, 1861 com avaliação de seu posicionamento filogenético em Sympetrinae (Insecta, Odonata, Libellulidae). Dissertação apresentada ao Programa de PósGraduação em Ciências Biológicas (Zoologia), Museu Nacional, da Universidade Federal do Rio de Janeiro/UFRJ, Brasil. [ Links ]
38. Rehn, A.C. 2003. Phylogenetic analysis of higher level relationships of Odonata. Systematic Entomology 28: 181-239. [ Links ]
39. Riek, E.F. & J. Kukalová-Peck. 1984. A new interpretation of dragonfly wing venation based upon Early Upper Carboniferous fossils from Argentina (Insecta: Odonatoidea) and basic character states in pterygote wings. Canadian Journal of Zoology 62(6): 1150-1166. [ Links ]
40. Sereno, P.C. 2007. Logical basis for morphological characters in phylogenetics. Cladistics 23: 565-587. [ Links ]
41. Song, H. & J.W. Wenzel. 2008. Mosaic pattern of genital divergence in three populations of Schistocerca lineata Scudder (Orthoptera: Acrididae: Cyrtacanthacridinae). Biological Journal of the Linnean Society 94: 289–301. [ Links ]
42. Song, H. & S.R. Bucheli. 2010. Comparison of phylogenetic signal between male genitalia and nongenital characters in insect systematics. Cladistics 26: 23-35. [ Links ]
43. Svensson, E.I. & M. Friberg. 2007. Selective predation on wing morphology in sympatric damselflies. The American Naturalist 170, 101-112. [ Links ]
44. Taylor, P.D. & G. Merriam. 1995. Wing morphology of a forest damselfly is related to landscape structure. Oikos 73: 43-48. [ Links ]
45. Vogt, L., T. Bartolomaeus & G. Giribet. 2010. The linguistic problem of morphology: structure versus homology and the standardization of morphological data. Cladistics 26: 301-325. [ Links ]
46. Ware, J., M. May & K. Kjer. 2007. Phylogeny of the higher Libelluloidea (Anisoptera: Odonata): An exploration of the most speciose superfamily of dragonflies. Molecular Phylogenetics and Evolution 45: 289-310. [ Links ]
47. Wenzel, J.W. & M.E. Siddall. 1999. Noise. Cladistics 15: 51-64. [ Links ]
48. Wiens, J.J. 1995. Polymorphic characters in phylogenetic systematics. Systematic Biology 44: 482-500. [ Links ]
49. Wilson , E.O. 1975. Sociobiology. The New Synthesis. Harvard University Press, Cambridge . 720pp. [ Links ]
50. Winston, J.E. 1999. Describing species. Practical taxonomic procedure for biologists. New York, NY. Columbia University Press. 518pp. [ Links ]













