RECONSTRUCCIÓN DE ACTIVIDAD NEURONAL MEG/EEG CON MÚLTIPLES PARCHES DISPERSOS Y VARIABLES
JOSÉ D. LÓPEZ
PhD(c).Investigador grupoGAUNAL, Escuela de Mecatrónica. Universidad Nacional de Colombia sede Medellín. Medellín, Colombia. jodlopezhi@unal.edu.co
GARETH R. BARNES
PhD.Wellcome Trust Centre for Neuroimaging, University College London, London, United Kingdom. g.barnes@ucl.ac.uk
JAIRO J. ESPINOSA
PhD. Profesor Asociado.Escuela de Mecatrónica, Universidad Nacional de Colombia sede Medellín. Medellín, Colombia. jespinov@unal.edu.co
]]> Received for review March 15th, 2012, accepted May 31th, 2012, final version June, 19th, 2012
ABSTRACT: MEG/EEG brain imaging has become an important tool in neuroimaging. The reconstruction of cortical current flow is an ill-posed problem, but its uncertainty can be reduced by including prior information within a Bayesian framework. Typically this involves using knowledge of the cortical manifold to construct a set of possible regions of neural source activity. In this work a second stage is proposed to reduce localisation error without severely increasing the computational load. This stage consists of iteratively updating the set of possible regions based on previous reconstructions, in order to focus on those brain regions with a higher probability of being active. The proposed methodology was tested with synthetic MEG datasets giving as a result zero localisation error for single sources and different noise levels. Real data from a visual attention study was used for validation.
KEYWORDS: MEG/EEG inverse problem, Multiple Sparse Priors, Brain imaging
RESUMEN: La reconstrucción de actividad neuronal a partir de datos MEG/EEG se ha convertido en una importante herramienta en neurología. A pesar de ser un problema mal condicionado, su incertidumbre se puede reducir incluyendo información previa en algoritmos basados en inferencia Bayesiana. Típicamente esto implica el uso de conocimiento acerca de la superficie cortical para generar posibles regiones de actividad neuronal. En este trabajo se propone una segunda etapa con el objetivo de reducir el error de localización sin aumentar fuertemente la carga computacional, esta etapa consiste en actualizar iterativamente el conjunto de posibles regiones de activación basándose en las reconstrucciones previas, enfocándose en aquellas regiones del cerebro que tienen más probabilidad de tener actividad. La metodología propuesta fue probada con datos simulados de MEG dando como resultado error cero de localización para fuentes únicas y diferentes valores de ruido, también se realizaron pruebas de validación con datos reales de actividad en la corteza visual.
PALABRAS CLAVE: Problema inverso MEG/EEG, Múltiples fuentes previas dispersas, Imágenes cerebrales.
1. INTRODUCCIÓN
MEG/EEG neural activity reconstruction involves the estimation of the cortical current distribution, which gives rise to the externally measured electromagnetic field. It is based on the assumption that local groups of neurons (around 104) can be modelled as equivalent current dipoles. There are two main ways to reconstruct brain activity based on this dipolar model: (a) Assume a small number of activated regions of arbitrary location and orientation, and fit with a non linear search through the brain [1]. (b) Populate the source space with a large number of dipoles distributed at fixed locations and orientations and estimate their amplitude. Recently major effort has been dedicated to the distributed approach because it is linear, independent of the number and characteristics of activated regions; and because using strategies to reduce the noise and search space, it is robust and computationally feasible [2].
Since the introduction of the well-known minimum norm estimation (also known as total least squares observer) to neuroimaging [3], it has been noted how the inclusion of prior information can reduce the uncertainty of the solution. The use of a smoother proposed by Pascual-Marqui et al. [4] showed how the same minimum norm structure could be improved with more informative priors.
]]> These static approaches have been improved with the inclusion of temporal information with Kalman filters[5]. The Kalman filter is a stochastic optimal estimator that updates the first two moments of random variables (in this case the dipoles) based on the data, providing a robust solution [6]. This Markovian update is highly dependent on the temporal transition among samples. Initially authors treated this problem with quadratic parametric equations based on physiological constraints [5, 7]. An extension to higher order models was presented in [8, 9].The use of Kalman filtering presented two main limitations: the lack of information to form the temporal model of all the neural activity, and the high computational burden with large sets of dipoles. Inclusion of temporal models from the data [10] and good estimation of parameters [9] allowed the reduction of the temporal model uncertainty, but model reduction techniques [5, 11] were unable to allow an increase in the number of dipoles, a requisite to reduce the quantisation error; neural sources between dipoles can only be located in the nearest dipole.
The main improvement of these Kalman filter approaches has been the natural inclusion of the Bayesian framework, already used in economy, engineering and astrophysics. Its implementation in neuroimaging is described within a unifying Bayesian framework compiled in [12]. The Bayesian approaches are inherently static, but given the off-line analysis of MEG/EEG data several dimensional reduction techniques such as principal components and wavelets can be implemented over the time window of interest [13, 14]. This avoids the regularisation based on data commonly used on inverse problems [15].
The main idea of the Bayesian framework is that if the MEG/EEG inverse problem is ill-posed, then it is not possible to perfectly reconstruct the neural activity, but it is possible to provide a probability distribution of the states with a given degree of certainty about their characteristics. It is achieved by using the theory of Empirical Bayes to construct an informative state covariance matrix, formed by the weighted sum of a set of possible covariance components. Each covariance component might, for example, describe the sensor level covariance one would expect due to an active patch of cortex. These weights are obtained by optimising a given cost function. A good example of this implementation is the Multiple Sparse Priors (MSP) algorithm [16], using the negative variational Free energy [17] for that purpose.
The MSP estimation is highly dependent on the set of selected covariance components (or patches). In absence of knowledge about the size, shape and location of the neural current flow, the set of components should ideally be composed of patches of all possible locations and sizes. But this would incur a prohibitively large computational load, conversely too few patches will result in an under sampled solution space.
In Harrison et al. [18] a Green's function based on a graph Laplacian was proposed in order to generate the set of components. This forms a compact set of bell shaped patches of finite cortical extent. Preliminary tests of this work showed that if the neural source is far from the patch centres of the Green's function the estimation fails [19].
In this work we present an improvement for reducing the localisation error due to a poor initial patch set in the MSP reconstruction. We suggest an iterative patch selection approach in which the result of a previous MSP reconstruction is used to generate a new set of patches. This new patch set is seeded close to the reconstructed sources of the previous iteration. This process can be done iteratively using the Free energy of each inversion as a cost function.
This manuscript proceeds as follows. In Section 2 the inverse problem is presented in the Bayesian framework and then the MSP algorithm is explained in terms of the Restricted Maximum Likelihood optimisation. The new stage for reducing the localisation error is also explained. In Section 3 simulation results with noisy synthetic MEG data are presented, these datasets were generated using realistic head models computed with the SPM8 software package (http://www.fil.ion.ucl.ac.uk/spm). In Section 4 the proposed methodology is validated with real MEG data;visual cortex activity is recovered after intentionally removing patches from the region with neural activity. Finally, the results are discussed in Section 5.
2. THEORY
]]> The MEG/EEG data can be related to the neural activity that generates it using the general linear model (GLM):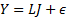
The neural activity 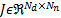 propagates the energy of
propagates the energy of 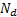 current dipoles through the head, where a set of
current dipoles through the head, where a set of 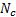 gradiometers/electrodes is used to acquire
gradiometers/electrodes is used to acquire 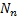 time samples on the dataset
time samples on the dataset 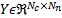 . The fixed location of the dipoles guarantees a linear propagation model that allows the use of a fixed gain matrix
. The fixed location of the dipoles guarantees a linear propagation model that allows the use of a fixed gain matrix 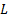 . Finally, the measurements are affected by zero mean Gaussian noise
. Finally, the measurements are affected by zero mean Gaussian noise  . Empty room noise can be introduced for more realistic assumptions [20].
. Empty room noise can be introduced for more realistic assumptions [20].
For a reliable reconstruction of the neural activity it is necessary to define a large number of fixed dipoles inside the search space; these dipoles usually outnumber the sensors ( 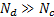 ), making it an ill-posed problem. Under Gaussian assumptions the solution of the GLM can be expressed as the minimisation problem:
), making it an ill-posed problem. Under Gaussian assumptions the solution of the GLM can be expressed as the minimisation problem:
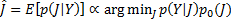 (1)
(1)
where the likelihood is 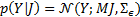 , with
, with 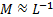 , and the prior source probability distribution is
, and the prior source probability distribution is 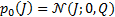 . Assuming a priori that
. Assuming a priori that 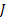 and
and  are zero mean Gaussian processes with covariances
are zero mean Gaussian processes with covariances 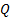 and
and 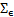 respectively, and
respectively, and 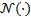 is the multi normal probability density function.
is the multi normal probability density function.
The estimated source activity 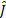 is obtained by minimising Eq. (1) [2]:
is obtained by minimising Eq. (1) [2]:
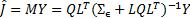
this solution is static but it can be extended to dynamic activity by projecting the temporal information as proposed in[13, 14].
2.1 Computation of the covariance components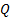 and
and 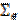 . In absence of sensor noise information the noise is considered independent and uniformly distributed:
. In absence of sensor noise information the noise is considered independent and uniformly distributed: 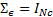 , with
, with 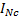 a
a 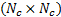 identity matrix. ]]>
The prior source covariance matrix has been traditionally defined as a single smoother [2]. But modern algorithms based on Empirical Bayes such as the Multiple Sparse Priors (MSP) algorithm [16] divide the smoother into a set of
identity matrix. ]]>
The prior source covariance matrix has been traditionally defined as a single smoother [2]. But modern algorithms based on Empirical Bayes such as the Multiple Sparse Priors (MSP) algorithm [16] divide the smoother into a set of 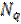 diagonal components
diagonal components 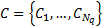 , each component typically (but not always) describing covariance due to a connected patch of cortex. These are subsequently weighted by the algorithm to make an estimate of the sensor covariance Q:
, each component typically (but not always) describing covariance due to a connected patch of cortex. These are subsequently weighted by the algorithm to make an estimate of the sensor covariance Q: 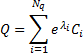
fMRI data can be highly informative when considered as components[21], but in absence of prior information these components are patches generated with a Green's function.
The sources of neural activity are composed by focal regions of neurons with synchronous activity; these focal regions are mostly bell shaped with a maximum in their centre and attenuation in amplitude corresponding to the square of the distance to it [22]. The Green's function is used to generate bell shaped (Gaussian) patches for each covariance component following this behaviour[18].
The Green's function is based on a graph Laplacian 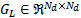 that uses the vertices and faces provided by a structural MRI of the cortical manifold. The Green's function
that uses the vertices and faces provided by a structural MRI of the cortical manifold. The Green's function 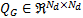 is defined as:
is defined as:
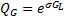 (2)
(2)
with 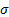 a positive constant value that determines the size of the activated regions. Each patch (covariance component) is generated with a column of
a positive constant value that determines the size of the activated regions. Each patch (covariance component) is generated with a column of 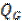 forming a bell centred on the corresponding vertex. The size of the regions, the number of patches, and their locations must be carefully selected in order to avoid empty spaces in the search space. If the set is too large the computational effort to determine which of them are active will be prohibitive.
forming a bell centred on the corresponding vertex. The size of the regions, the number of patches, and their locations must be carefully selected in order to avoid empty spaces in the search space. If the set is too large the computational effort to determine which of them are active will be prohibitive.
With the set of covariance components 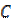 generated, the weights (hyperparameters)
generated, the weights (hyperparameters) 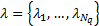 are pruned to those
are pruned to those 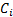 corresponding to activated regions with a non-linear search method, using the Free energy as the cost function.
corresponding to activated regions with a non-linear search method, using the Free energy as the cost function.
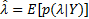 . It can be expressed in terms of known distributions using the Bayes' theorem:
. It can be expressed in terms of known distributions using the Bayes' theorem: 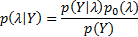
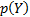 is fixed for a given dataset. The problem is that we only have an approximate posterior: q
is fixed for a given dataset. The problem is that we only have an approximate posterior: q 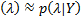 ,and it is necessary to use a cost function to find the optimal set of hyperparameters.
,and it is necessary to use a cost function to find the optimal set of hyperparameters. Let us define the log evidence as:
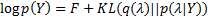
with 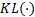 the Kullback-Leibler (KL) divergence. When the approximate posterior
the Kullback-Leibler (KL) divergence. When the approximate posterior 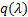 is equal to the true posterior
is equal to the true posterior 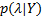 , the KL divergence is zero, and the Free energy is maximised:
, the KL divergence is zero, and the Free energy is maximised: 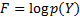 .
.
The Free energy gives a measure between the variance of the data 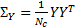 , and the model based variance
, and the model based variance 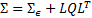 ; while at the same time punishing models with large numbers of hyperparameters[17]:
; while at the same time punishing models with large numbers of hyperparameters[17]:
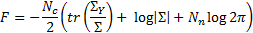
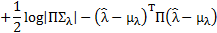
where 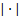 is the matrix determinant operator, and the prior and approximate densities of the hyperparameters are consideredto be Gaussian distributed:
is the matrix determinant operator, and the prior and approximate densities of the hyperparameters are consideredto be Gaussian distributed: 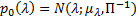 , and
, and 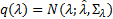 . The optimal combination of hyperparameters is achieved when the maximum Free energy value:
. The optimal combination of hyperparameters is achieved when the maximum Free energy value: 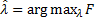 , which is when the Free energy is approximately the log evidence. In absence of prior information the hyperparameters may be selected with zero mean and infinite variance in order to guarantee flat hyperpriors.
, which is when the Free energy is approximately the log evidence. In absence of prior information the hyperparameters may be selected with zero mean and infinite variance in order to guarantee flat hyperpriors.
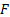 with respect to the hyperparameters:
with respect to the hyperparameters: 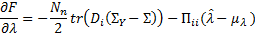
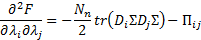
with
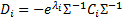
The update of the hyperparameters is obtained for 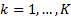 iterations:
iterations:
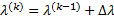
using Fisher scoring as the updating rule:
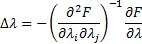
Within the ReML updates, those hyperparameters near to zero are pruned for faster computation. The convergence is achieved for small changes in the Free energy 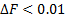 .Note that the total Free energy was not computed with ReML, it is done just once at the end of the iterative process.
.Note that the total Free energy was not computed with ReML, it is done just once at the end of the iterative process.
This procedure is similar to the field of genetic algorithms, where the most probable patches are used as "parents" to generate new "children" in their neighbourhood. The proposed procedure can be implemented as follows:
Note that each iteration is performed over the initial set of patches plus a new set based on those active regions identified in the previous iteration. The new set of patches must cover a region surrounding those active vertices, for example a circle of twice the diameter of a single patch centred on the location of the most active vertex. The initial set of patches is necessary because background activity is expected and it must be explained; i.e. several brain regions are permanently active but may not be of interest.
When a patch is correctly located at the neural source location, the Free energy reaches its maximum; it is expected that further iterations will maintain the maximum Free energy, but it has been observed that sometimes in following iterations more patches become active around the true source, increasing the complexity and consequently reducing the Free energy value. This issue motivates the use of the variation of Free energy as stopping criterion of the proposed algorithm.
In the following sections this stage will be tested with noisy synthetic data and validated with data due to visual cortical activity.
3. SIMULATION RESULTS
]]> Single trial datasets of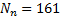 samples over
samples over 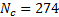 MEG sensors were generated, by projecting a single source in different locations in the cortical surface. A sinusoidal signal of 20 Hz was used for the temporal waveform of the source of neural activity. A Signal to Noise Ratio
MEG sensors were generated, by projecting a single source in different locations in the cortical surface. A sinusoidal signal of 20 Hz was used for the temporal waveform of the source of neural activity. A Signal to Noise Ratio 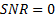 dB was added to the data using white random noise. The sources were focal, Gaussian meshes with full width at half maximum (FWHM) of 10 mm, this size corresponds to common neural activated regions. Figure 1(a) shows an example of a source located in occipital cortex. The translucent glass brains of Figure 1 show the frontal, lateral and superior views of the 512 dipoles with highest variance during the time window of interest (-200 - 500 ms).
dB was added to the data using white random noise. The sources were focal, Gaussian meshes with full width at half maximum (FWHM) of 10 mm, this size corresponds to common neural activated regions. Figure 1(a) shows an example of a source located in occipital cortex. The translucent glass brains of Figure 1 show the frontal, lateral and superior views of the 512 dipoles with highest variance during the time window of interest (-200 - 500 ms). The MSP algorithm was implemented over a grid of 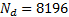 dipoles oriented orthogonal to and distributed over the entire cortical surface, as shown in Figure 2(a). The default MSP inversion was performed with
dipoles oriented orthogonal to and distributed over the entire cortical surface, as shown in Figure 2(a). The default MSP inversion was performed with 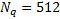 patcheswhose centres were randomly selected from the
patcheswhose centres were randomly selected from the 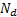 dipoles locations as can be seen in Figure 2(b). The size of the patches was approximately 10 mm. The initial values used in the MSP algorithm optimisation are those of [16].
dipoles locations as can be seen in Figure 2(b). The size of the patches was approximately 10 mm. The initial values used in the MSP algorithm optimisation are those of [16].
The single source shown in Figure 1(a) was located intentionally far from the patch centres of Figure 2(b). The source reconstruction made with the MSP shown in Figure 1(b) is a good example of the algorithm failure when the patches do not match with the sources.
3.1 Results of iterative updates
Figure 1(b) shows that most of the energy is divided in three separated regions, one near the true source, one superficialin the left hemisphere, and another one deep in the left hemisphere. These activated regions were used as seeds about which to create a new set of patch centres. New patch centres were drawn from a Gaussian distribution of FWHM=20 mm around each active peak(Twice the size of a patch, see Figure 3(a)). For simplicity in this example, the default set of patches was not used in the new sets; as the synthetic data do not have extra activity this does not affect the Free energy.
Figure 3(b) shows the MSP source reconstruction obtained with the new set of patches. It is clear that the estimated active cortex is now bounded by the more focally seeded patches. Also the solution is beginning to approach the expected results (Figure 1(a)).
A third iteration was performed again updating the set of patches.The third patch set (shown in Figure 4(a)) became in turn more focal and the reconstruction again approached the (focal) simulated case(Figure 4(b)).
]]> Given that the Free energy continued increasing, a fourth inversion was performed. Figure 5(a) shows the new set of patches filling the region of reconstructed activity. Figure 5(b) shows the fourth MSP reconstruction. The overlapping of patches around the true location meant that several overlapped patches were used to emulate the activity of the true source. The question now remains at which iteration should we have stopped, given that normally we have no knowledge of localisation error. 3.2 ConvergenceThe localisation error of the source of neural activity was defined as the Euclidean distance between its true location: 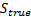 , and the location of the dipole with maximum energy after the estimation,
, and the location of the dipole with maximum energy after the estimation, 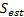 :
: 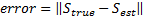 .
.
Preliminary tests with different noise levels (not shown here) and several source locations presented also zero localisation error after three or four iterations.
4. VALIDATION WITH REAL DATA
We used some MEG data acquired in a visual attention task to validate the method. A detailed description of the experimental set-up and previous data analysis were presented in [24]. Averaged single subject data were used.
Figure 6(a) shows the measured activity at the scalp at 151.6 ms of recording. For this experiment the set of patches located within the visual cortex was deliberately sparse, affecting the source reconstruction as shown in Figure 6(b). This first reconstruction had a Free energy value of 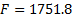 .
.

The second iteration of patches is shown in Figure 7(a). The distribution of these patches was a Gaussian of FWHM=20 mm. Figure 7(b) shows the MSP estimation of the second iteration, there was an increase in the Free energy to 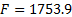 , and the source activity moved closer to the visual cortex. This new region was used to generate the new set of patches, shown in figure 7(c), for a third iteration where physiologically plausible sources in visual cortex can be observed. Figure 7(d) shows the final reconstruction.
, and the source activity moved closer to the visual cortex. This new region was used to generate the new set of patches, shown in figure 7(c), for a third iteration where physiologically plausible sources in visual cortex can be observed. Figure 7(d) shows the final reconstruction.
The Free energy for the third inversion was 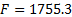 . Given that it continued increasing a forth inversion was performed, but the Free energy maintained its value finishing the iterative process and defining the third inversion as the final one.
. Given that it continued increasing a forth inversion was performed, but the Free energy maintained its value finishing the iterative process and defining the third inversion as the final one.
5. DISCUSSION
In this paperwe have presented a principled and computationally efficient patch update method for the MSP inversion scheme.This patch update is based in two assumptions [23]: Nearest patches of neural activity are active; and a source reconstruction with a patch over the true source has higher Free energy than a reconstruction without it. Theoretically both assumptions are explained by the accuracy of a solution with the right patch distribution, and the complexity of reconstructing the data without correct patches [17].
The ill-conditioned nature of the MEG/EEG inverse problem allows infinite possible solutions for a single dataset, and the noise increases this uncertainty. It is suggested that a fixed initial distribution covering the entire source space in order to allow the optimisation process to explain the non-interesting activity.One possible drawback of this algorithm is that it relies on an approximately correct initial inversion. There are however many similar approaches which (although entailing higher computational load) might in practice prove to be more robust. For example, one could simply use randomly distributed patch centres from the outset [23]. One could classify such inversions based either on that which yielded the highest Free energy (as here) or through Bayesian Model Averaging [25].
Some authors have criticised the convexity assumption on the Free energy [12], or proposed Markov based solutions [5]. But as stated in [25] the quadratic assumptions are comparable to the Euclidean norm estimation, and the source reconstruction used here is the same as Kalman gain with an improvement on priors, the Bayesian framework uses all the data for providing informative priors (Empirical Bayes) avoiding defining dynamics of the neural activity and allowing the reduction of model dimensionality [16].
Finally, note that a very similar procedure to that outlined here could be performed to determine other spatial characteristics such as the true extent of the patches. In this case, the same patch centres would be used but the Green's function modified (Eq. (2)) between iterations.
]]>ACKNOWLEDGMENTS
J.D. López and J.J. Espinosa are supported by ARTICA Research Centre for Excellence, Universidad Nacional de Colombia, and Colciencias, in the project "Procesamiento de Señales". The Wellcome Trust Centre for Neuroimaging is supported by a strategic award from the Wellcome Trust. The authors would like to thank Marcus Bauer for providing the dataset and helpful information for this work.
REFERENCES
[1] Supek, S. and Aine, C.J., Simulation studies of multiple dipole neuromagnetic source localization: model order and limits of source resolution. IEEE transactions on biomedical engineering, Vol 40 (16), pp529 540, 1993. [ Links ]
[2] Grech, R., Cassar, T., Muscat, J., Camilleri, K., Fabri, S., Zervakis, M., Xanthopoulos, P., Sakkalis, V. and Vanrumste, B., Review on solving the inverse problem in EEG source analysis.Journal of NeuroEngineering and Rehabilitation 5, 25, 2008. [ Links ]
[3] Hamalainen, M. and Sarvas, J., Realistic Conductivity Geometry Model of the Human Head for Interpretation of Neuromagnetic Data. IEEE transactions on biomedical engineering, 36, pp165 171, 1989. [ Links ]
[4] Pascual-Marqui, R., Michel, C.M. and Lehmann, D., Low Resolution Electromagnetic Tomography: A new method for localizing electrical activity in the brain. International Journal of psychophysiology, 18, pp. 49-65 1994. [ Links ]
[5] Barton, M., Robinson, P., Kumar, S., Galka, A., Durrant-Whyte, H., Guivant, J. and Ozaki, T., Evaluating the Performance of Kalman-Filter-Based EEG Source Localization. IEEE transactions on biomedical engineering, 56, pp. 122 136, 2009. [ Links ]
[6] Kalman, R.A., New Approach to Linear Filtering and Prediction Problems. Transactions of the ASME, Journal of Basic Engineering, 82, pp. 35 45, 1960. [ Links ]
[7] Kim, J.W., Robinson, P.A. Compact dynamical model of brain activity. Phys. Rev. E, American Physical Society, 75, 10 pages, 2007. [ Links ]
[8] Antelis, J.M. and Minguez, J., Dynamic solution to the EEG source localization problem using Kalman Filters and Particle Filters. Annual International Conference IEEE EMBS, 2009. [ Links ]
[9] Giraldo, E., den Dekker, A. and Castellanos, G., Estimation of dynamic neural activity using a Kalman filter approach based on physiological models. Annual International Conference IEEE EMBS, pp. 2914 2917, 2010. [ Links ]
[10] López, J.D., Valencia, F. and Espinosa, J.J., EEG brain imaging based on Kalman filtering and subspace identification. IEEE Second Latin American Symposium on Circuits and Systems (LASCAS), 4 pages, 2011. [ Links ]
[11] López, J.D., Espinosa, J.J.Spatio-temporal EEG brain imaging based on reduced Kalman filtering. 5th International Conference IEEE EMBS, pp. 64 67. 2011. [ Links ]
[12] Wipf, D., Nagarajan, S.A unified Bayesian framework for MEG/EEG source imaging. Neuroimage, 44, pp. 947 966, 2009. [ Links ]
[13] Phillips, C., Rugg, M. and Friston, K., Anatomically Informed Basis Functions for EEG Source Localization: Combining Functional and Anatomical Constraints. NeuroImage, pp. 16, pp 678 695, 2002. [ Links ]
[14] Trujillo, N.J., Aubert, E. and Penny, W.D. Bayesian M/EEG source reconstruction with spatio-temporal priors. Neuroimage, 39, pp. 318 335, 2008. [ Links ]
[15] Álvarez, A.M., Daza, G., Acosta, C.D. and Castellanos, G., Parameter selection in least squares-support vector machines regression oriented, using generalized cross-validation. Revista DYNA, 171, pp. 23 30, 2011. [ Links ]
[16] Friston, K., Harrison, L., Daunizeau, J., Kiebel, S., Phillips, C., Trujillo, N., Henson, R., Flandin, G. and Mattout, J., Multiple sparse priors for the M/EEG inverse problem. Neuroimage, 39, pp. 1104 1120, 2008. [ Links ]
[17] Friston, K., Mattout, J., Trujillo, N., Ashburner, J. and Penny, W., VariationalFree energy and the Laplace approximation. Neuroimage, 34, pp.220 234, 2007. [ Links ]
[18] Harrison, L., Penny, W., Ashburner, J., Trujillo, N. and Friston, K., Diffusion-based spatial priors for imaging. Neuroimage, 38, pp.677 695, 2007. [ Links ]
[19] López, J.D. and Espinosa, J.J., Single MEG/EEG source reconstruction with multiple sparse priors and variable patches. International Seminar on Medical Image Processing and Analysis (SIPAIM), 2011. [ Links ]
[20] Henson, R., Mouchlianitis, E. and Friston, K., MEG and EEG data fusion: simultaneous localisation of face-evoked responses.NeuroImage 47(2), pp.581 589, 2009. [ Links ]
[21] Henson, R.N., Flandin, G., Friston, K.J. and Mattout, J.A., parametric empirical Bayesian framework for fMRI-constrained MEG/EEG source reconstruction. Human Brain Mapping, 31, pp. 1512 1531, 2010. [ Links ]
[22] Hallez, H., Vanrumste, B., Grech, R., Muscat, J., De Clercq, W., Vergult, A., D'Asseler, Y., Camilleri, K.P., Fabri, S.G., Van Huffel, S. and Lemahieu, I., Review on solving the forward problem in EEG source analysis Journal of Neuro Engineering and Rehabilitation, 4:46. 2007. [ Links ]
[23] López, J., Espinosa, J. and Barnes, G., Random location of Multiple Sparse Priors for solving the MEG/EEG inverse problem. Anual international conference of the IEEE EMBC, 2012.(Accepted). [ Links ]
[24] Bauer, M., Kluge, C., Bach, D., Bradbury, D., JochenHeinze, H., Dolan, R.J. and Driver, J., Cholinergic Enhancement of Visual Attention and Neural Oscillations in the Human Brain. Current Biology, pp. 397 402, 2012. [ Links ]
[25] López, J.D., Penny, W.D., Espinosa, J.J. and Barnes, G.R., A general Bayesian treatment for MEG source reconstruction incorporating lead field uncertainty.Neuroimage, 60, pp.1194 1204, 2012. [ Links ]