Services on Demand
Journal
Article
Indicators
-
 Cited by SciELO
Cited by SciELO -
 Access statistics
Access statistics
Related links
-
 Cited by Google
Cited by Google -
 Similars in
SciELO
Similars in
SciELO -
 Similars in Google
Similars in Google
Share
Colombia Médica
On-line version ISSN 1657-9534
Colomb. Med. vol.43 no.1 Cali Jan./Mar. 2012
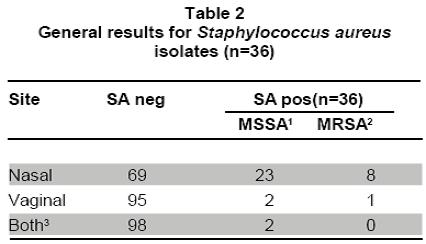
Nasal and vaginal colonization of methicillin-resistant Staphylococcus aureus in pregnant women in Cartagena, Colombia Colonización nasal y vaginal por Staphylococcus aureus resistente a meticilina en mujeres embarazadas en Cartagena, Colombia Oscar Correa, MD1, Kelly Delgado, MD2, Carla Rangel, MD3, Ana Bello, MD4, Niradiz Reyes, PhD5 1Research Assistant, Genetics and Molecular Biology group, Department of Basic Sciences. School of Medicine, Universidad de Cartagena, Cartagena, Colombia. e-mail: oscarleo27@gmail.com Received for publication November 17, 2010 Accepted for publication July 27, 2011 Summary Introduction: The host niche for Staphylococcus aureus (SA) are the anterior nares; however, vaginal colonization rates between 14% and 17.1% in pregnant women have been recently reported, raising interest about the potential risk in postpartum women and in neonates from colonized mothers. Keywords: Staphylococcus aureus; Methicillin-resistance; Maternal colonization; Virulence factor; Pregnancy. Resumen Introducción: Staphylococcus aureus (SA) ha adoptado como nicho habitual en las narinas anteriores; sin embargo, recientemente se han reportado tasas de colonización del tracto genital de mujeres embarazadas entre 14% y 17.1%, lo que aumenta el interés respecto al riesgo asociado para el neonato y la madre en el posparto. Palabras clave: Staphylococcus aureus; Resistencia a meticilina; Factores de virulencia; Embarazo; Colonización. Staphylococcus aureus (SA) is an important human pathogen responsible for nosocomial and communityacquired infections, which also behaves as commensal flora in healthy individuals â mainly colonizing the anterior nares1. Carriers of SA, in particular methicillinresistant Staphylococcus aureus (MRSA), have a higher risk for developing clinical infections, being the infections caused by MRSA strains the most important at the clinical level because they are more difficult to treat2. In recent decades, MRSA strains, especially those associated to the community [communityassociated methicillin-resistant Staphylococcus aureus (CA-MRSA)] have become an important issue in public health, given that they are a cause of increased morbidity and mortality3. These CA-MRSA strains cause severe skin and soft tissue infections, necrotizing pneumonia, and sepsis in otherwise healthy children, teens, and, more recently, in neonates. The increasing number of infections in neonates caused by CAMRSA emphasizes the need to identify the possible environmentally or maternally derived sources of infection4,5. For pregnant women, SA also poses a health risk because it is the main cause of infection of the surgical site, causing between 25% and 50% of infections of the post cesarean surgical site, representing a major cause of morbidity and a cause of puerperal mastitis6. Early epidemiological studies showed that 5% of women were colonized with SA in their genital tract and postpartum women had the highest colonization rates; furthermore, vaginal-rectal carriage of SA has been found associated with development of postpartum fever7. Although risk factors associated to colonization with MRSA strains during pregnancy have not been fully characterized, associations with race, parity, type of birth, and colonization with group B streptococci have been suggested7. It is known that incidence of CA-MRSA infections varies among different communities and populations, and apparently, pregnant women are more susceptible and have risk factors that predispose them to developing these infections6,8. Nevertheless, there is a scarcity of epidemiologic reports about MRSA infections present in pregnant and puerperal women6. More recently, several studies, mainly from the United States, have reported vaginal colonization rates for SA in pregnant women from 14% to 17.1%, and although the risk of vertical transmission has been suggested or even demonstrated9-11, the association of such colonization with infant outcome has not been clearly established12. Thus, there is a need to increase knowledge in this area of research. A literature review of the subject reveals few published studies in this area, which are mainly reviews of clinical cases13, making clear that more epidemiology-based studies are needed, given the increasing importance of CA-MRSA strains in serious neonatal infections14 and in women in puerperal stage8. Antibiotic resistance of MRSA strains is due to the acquisition of mecA gene, which codes for PBP-2a, a penicillin-binding protein with low binding affinity for beta-lactam antibiotics. The MecA gene is located in a mobile genetic element, the chromosomal staphylococcal cassette (SCCmec) of which seven major types have been identified, from I to VII. SCCmec types IV, V and VII are the most frequently found in CA-MRSA isolates, which also frequently carry the genes for Panton-Valentine leukocidin (PVL)15. Up to 96% of MRSA infections in pregnant women mainly affect the skin and soft tissue, although several cases of pneumonia in the puerperal period have also been attributed to MRSA strains6. Because PVL-carrying MRSA strains have been implicated in the development of serious skin and soft tissue infections and in necrotizing pneumonia16, in the present study besides determining the SA nasal and vaginal colonization rates and antibiotic profiles of the isolates, we evaluated the presence of PVL genes and SCCmec types of MRSA isolates by multiplex PCR. Methods Design, study population, and inclusion criteria. This was a pilot study performed by the Genetics and Molecular Biology Research Group at Universidad de Cartagena. The study site, the Clínica de Maternidad Rafael Calvo (CMRC), in Cartagena, Colombia, is a university-affiliated maternity clinic attending pregnant women from the urban and rural areas around Cartagena, and whose obstetric service delivers between 8,000 and 8,500 infants per year. Following a research protocol approved by the Ethics Committee of Universidad de Cartagena and the Institutional Review Board of CMRC, women with at least 35 weeks of gestational age, who attended the outpatient or emergency departments during the months of January through June 2009 were enrolled in the study after signing an informed consent. Women with ruptured membranes and those under antibiotic treatment or with clinical evidence suggesting a current staphylococcal infection were excluded from the study. By following approaches previously described by Pinter et al.13 and Beigi et al.9 nasal and vaginal swabs were obtained from the pregnant women enrolled (n=100) with vaginal samples taken from the outer third portion of the vagina. Although the objective of the study was to determine SA nasal and vaginal colonization rates and antibiotic susceptibility, we tried to ascertain any possible associations of colonization with complications related to staphylococcal infection either in the mothers or their newborns implementing a post discharge weekly follow-up by telephone for up to four weeks. Mothers were contacted by telephone at least once a week during the babyâs first month of life. They were asked of the general health of their infants, as well as of their own health. Specific questions were asked to determine whether any fevers, rashes, or other skin lesions had occurred in the infant. Additionally, participants were asked to file a questionnaire about social demographic data and medical records. Laboratory methods. Nasa l and vaginal specimens were transported to the microbiology laboratory of the School of Medicine at Universidad de Cartagena and processed within 8 to 18 h, according to protocols described by Bettin et al.17 Antibiotic susceptibility of isolates confirmed as SA was performed by the disc diffusion method following recommendations of the Clinical and Laboratory Standards Institute (CLSI). The antibiotics evaluated were: rifampin (5 µg), clindamycin (2 µg), erythromycin (15 µg), gentamicin (10 µg), vancomycin (30 µg), cefoxitin (30 µg), and oxacillin (1 µg). The SA strain ATCC 33591 was used as control. The D-zone test for inducible clindamycin resistance was performed for each isolate according to CLSI method. Isolates were classified as MRSA if they demonstrated resistance to cefoxitin and MSSA if they were susceptible. MRSA were further confirmed by PCR amplification of mecA gene. Both MSSA and MRSA isolates were stored at -35°C for subsequent molecular studies. Molecular analysis Genomic DNA extraction. Genomic DNA from each isolate was obtained according to the protocol described by Millar et al.18 with some modifications. Briefly, each SA isolate was sub-cultured in nutrient agar for 24 h at 37oC. Around five colonies were resuspended in 1 ml of Tris 0.5 M, centrifuged at 13,000 rpm x 5 min. Supernatant was discarded and the pellet resuspended in 500 µl buffer TE (10 mM Tris; 1 mM EDTA, pH: 8.0) and boiled at 100°C for 30 min, and then incubated at -35°C for 20 min, thawed at 65°C and finally centrifuged at 13,000 rpm for 15 min. Supernatant containing bacterial DNA was collected in a clean tube and stored at -20°C for subsequent PCR assays. Multiplex PCR assays. Multiplex polymerase chain reaction assays for assessments of presence of the lukF-PV (encoding part of the PVL toxin), mecA, and nuc genes were performed for all SA isolates by using previously described primers19-21. SA strains ATCC 33591 (mecA+; nuc+; PVL) and ATCC 25923 (mecA were used as amplification controls and pure water as negative controls. All SA isolates were subjected to multiplex PCR by using a set of three primer pairs previously reported: MecA1FMecA2R that amplifies a 147-bp fragment of mecA gene; Nuc1FNuc2R that amplifies a 300-bp fragment of nuc gene specific for SA, and LukPV1FLukPV2R that amplifies a 437-bp fragment of PVL gene. Additionally, all confirmed MRSA isolates were subjected to SCCmec typing by using a multiplex PCR assay, according to the protocol described by Zhang et al.19 by using SA strains NCTC10442 for SCCmec type I, N315 for SCCmec type II, and JCSC4744 for SCCmec type IV. PCR products were visualized in a 2% agarose gel stained with ethidium bromide under UV transillumination. Statistical analysis. Statistical analysis was performed with SPSS version 13.0 for Windows. Based on microbiological results, the study population was classified as carrier and non-carrier and subclassified according to the colonization site (vagina/ nares/both) and the frequency of colonization was determined for each of the two anatomical sites studied. Univariate analysis was applied to determine association of colonization to potential risk factors by using the Fisher exact test using a p value d»0.05 for statistical significance. Results and discussion Table 1 shows demographic data from our study population and Table 2 shows the general results of SA isolates from this study. One hundred pregnant women were enrolled in this pilot study during a 6month period and 34 of them were colonized with SA. Twenty nine of them were colonized only in the nares, three were colonized only in the vagina, and two harbored S. aureus at both sites. Colonization of pregnant women with SA was more common in the nares than in the vagina or at both sites [29/34 (85.3%) vs 3/34 (8.8%) and 2/34 (5.9%); p<0.05]. From the total of participants, we obtained 36 SA isolates, nine of which (25%) were MRSA, one was from the vagina; thus, the overall MRSA colonization rate among pregnant women was 9%. The nasal colonization rate for SA found in our pilot study (29%) was similar to rates previously reported for the general population2,22. However, the nasal colonization by MRSA (8% of the overall study population, 27.6% of the nasal SA isolates) was higher than the frequencies previously described both at international and national levels22. Albeit, at local level there is a report of similar nasal colonization rates for MRSA in school-age children23; in other local reports, MRSA nasal isolates were not detected among elderly individuals residing in a nursing-home17. To our knowledge, the colonization for SA in pregnant women had not been evaluated previously in Colombia. We found in this pilot study 3% vaginal and 2% nasal and vaginal colonization rates for SA, comparable to rates reported in similar studies in geographically different populations9,12, and a 29% nasal colonization rate, which is in the range described in other similar studies9,13. For MRSA, nasal colonization rates in this group of pregnant women were much higher (8%) than the rates reported for the general population15,24 and for pregnant women in different geographical areas13. For example, a pilot study carried out in Cleveland (USA) reported that 22% of the study population were nasal carriers of SA9. The authors of that study referenced a former report published in 1978 where the nasal colonization of SA in pregnant women was 4%. In a study similar to ours, Pinter et al.13, reported that from 304 pregnant women, 34 (11.2%) of them were colonized in the nares, seven of whom (2.3%) were colonized with MRSA13 Most recent data on SA colonization in obstetrics (both MSSA and MRSA) are derived mainly from the USA5,9-11,13 or Europe12, and until now they provide little evidence that either universal or targeted screening is beneficial for mothers or babies5,11,13,24,25. Higher colonization rates with SA are observed when the samples are from rectal-vaginal samples10,11 compared to vaginal12,13, which may reflect the fact that SA colonizes frequently the gastrointestinal tract. In the study published by Andrews et al.11 genital tract colonization by MRSA among pregnant women was evaluated and correlated with infant outcome. From 5732 pregnant women participating in the study, 833 were colonized by SA (14.5%), 202 of whom were MRSA positive (3.5% overall MRSA colonization). The authors reported that no cases of early-onset invasive neonatal infection by MRSA occurred among infants in their study. In the pilot study presented here, from 100 pregnant women screened, only five vaginal isolates were obtained, one of which was MRSA. Social demographic characteristics of our study population are summarized in Table 1. Statistical analysis of this data did not show significant associations with carrier state. This may reflect the fact that our sample population was small, although similar studies with larger populations have also failed to identify risk factors for SA colonization in pregnant women and the implications for maternal and neonatal outcomes5,11,12. In our study, follow up could only be completed for 52 mother-infant pairs, and we did not observe SA-related complications in mothers or in newborns; thus, we could not identify factors associated to a higher risk of colonization or development of complications either in mothers or infants. To date, rectal vaginal colonization of SA has been associated to race, parity, route of birth, and colonization by Streptococcus agalactiae7. In fact, Chen et al.7, reported that patients colonized by S. agalactiae were also more likely to be colonized with SA, but those colonized by MRSA were not colonized with S. agalactiae.
2General Practitioner, Unidad Intermedia San Francisco de Asís, Sincelejo, Colombia. e-mail: kellyosbourne02@hotmail.com
3General Practitioner, ESE San Francisco Javier, Margarita, Colombia. e-mail: caparajim18@hotmail.com
4Auxiliary Professor. Department of Gynecology and Obstetrics, School of Medicine. Universidad de Cartagena, Cartagena, Colombia. e-mail: amabet2001@hotmail.com
5Associate Professor, Group of Genetics and Molecular Biology, Department of Basic Sciences, School of Medicine, Universidad de Cartagena, Cartagena, Colombia. e-mail: nreyesr@unicartagena.edu.co
Objectives: To determine the prevalence of nasal and vaginal colonization of SA and the antibiotic susceptibility of the isolates in pregnant women attending a maternity hospital in Cartagena, Colombia.
Methods: Nasal and vaginal swabs were obtained from participants and subjected to microbiological and molecular assays. A post discharge follow-up was performed for up to four weeks.
Results: From 100 pregnant women enrolled in the study, 34 were colonized with SA; 29 only in the nares, threeonly in the vagina, and two at both sites. Colonization of pregnant women with SA was more common in the nares than in the vagina or at both sites [29/34 (85.3%) vs 3/34 (8.8%) and 2/34 (5.9%); p<0.05]. We obtained 36 SA isolates, nine (25%) of which were methicillin-resistant Staphylococcus aureus (MRSA), one was from the vagina; thus, the overall MRSA colonization rate among pregnant women was 9%. Molecular analysis showed that Panton-Valentine leukocidin (PVL) genes were carried by the vaginal MRSA, seven of the nasal MRSA, and two of the Methicillinsensitive Staphylococcus aureus (MSSA) isolates. Two MRSA isolates carried SCCmec type I and seven carried SCCmec type IV.
Conclusions: Nasal colonization rate for SA in the study population was similar to previous reports. However, the frequency of nasal colonization of MRSA was higher while vaginal colonization of SA was lower than previously reported in other studies for similar populations. The MRSA isolates obtained showed a community profile.
Objetivo: Determinar la prevalencia de colonización nasal y vaginal por SA y los patrones de susceptibilidad a antibióticos de los aislamientos en una comunidad obstétrica de Cartagena, Colombia.
Métodos: Se obtuvieron hisopados nasales y vaginales de las participantes y s e sometier on a ensayos microbiológicos y moleculares. Se realizó seguimiento a la madre y al neonato durante cuatro semanas.
Resultados: De 100 embarazadas participantes, 34 estuvieron colonizadas con SA; 29 solo en fosas nasales, tres solo vaginal y dos en ambos sitios. La colonización fue más común en fosas nasales que en vagina o en ambos sitios [29/34 (85.3%) vs 3/34 (8.8%) y 2/34 (5.9%); p<0.05]. Se obtuvieron 36 aislamientos de SA de los cuales 9 fueron MRSA (25%), proviniendo uno de vagina; la tasa de colonización total por MRSA en embarazadas fue 9%. Los genes para la leucocidina de Panton-Valentine (PVL) se detectaron en la cepa MRSA vaginal, siete MRSA nasales, y dos MSSA. Dos aislamientos MRSA portaban el elemento SCCmec tipo I y siete el tipo IV. No se detectó resistencia a otros antibióticos en los aislamientos MRSA; tres aislamientos susceptibles a meticilina (MSSA) fueron resistentes a eritromicina.
Conclusiones: Aunque la colonización nasal por SA en la población estudiada estuvo dentro del rango reportado previamente, la colonización nasal por MRSA fue mayor, mientras que la colonización vaginal fue más baja que las informadas previamente en otros estudios para poblaciones similares. Los aislamientos MRSA obtenidos presentaron un perfil comunitario.
Although asymptomatic nasal, vaginal, and rectal colonization with MRSA has been reported to occur in some pregnancy-related clinical cases in which SA colonization has been identified as risk factor for serious systemic infection after delivery8, epidemiological studies show that universal screening and decolonization efforts in pregnant women are currently non-cost effective12,25. Similarly, there is no current evidence that vertical transmission of SA constitutes a significant risk for neonatal complications, although vertical transmission of SA have been documented in some studies4,13.
In our pilot study, antibiotic susceptibility of the SA isolates found that all MRSA isolates had the resistance profile associated to the community, with exclusive resistance to methicillin. Also, three MSSA isolates had erythromycin resistance; one of them had the erythromycin inducible clindamycin resistance phenotype, detected by the D-test. We did not find resistance to other antibiotics in MRSA or MSSA isolates.
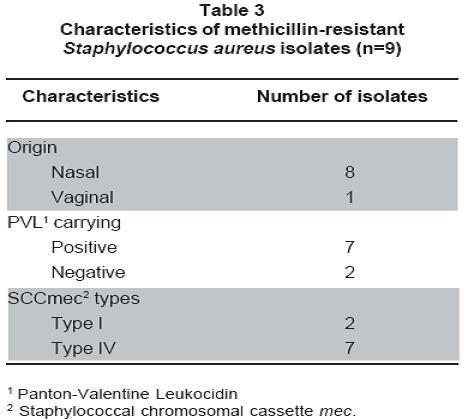
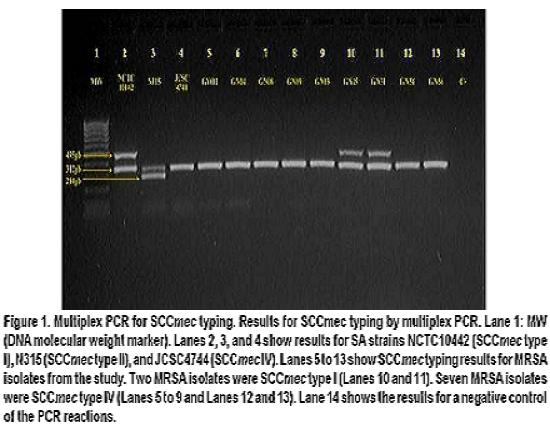
Table 3 summarizes the characteristics of MRSA isolates. SCCmec typing by multiplex PCR assay (Figure 1) showed that among the MRSA isolates from this study, two had SCCmec type I and seven had SCCmec type IV, data consistent with communityassociated MRSA, given that it has been previously reported that MRSA isolates commonly found in community settings frequently bear SCCmec type IV (and V)15.
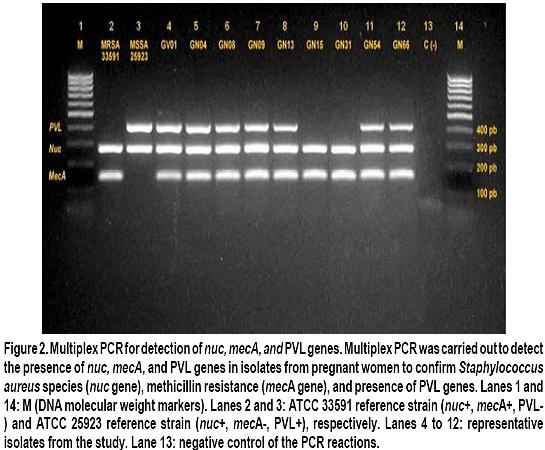
Although controversial, Panton-Valentine leukocidin has been widely regarded as a major virulencedeterminant associated to epidemic clones of CAMRSA in the world16. In the current study, we determined the presence of PVL genes among SA isolates, both MSSA and MRSA. Among MRSA isolates, six nasal isolates and the vaginal isolate were PVL positive (Figure 2). In contrast, among the MSSA isolates, only two were PVL positive; thus, PVL genes were more frequently found in MRSA isolates compared to MSSA [7/9 (77.7%) vs 2/27 (7.4%); p<0.01].
Conclusions
Although this was a small pilot study and it may not be generalized to the entire pregnant population, the colonization rates found for MRSA were high, as were the frequency of PVL-positive MRSA isolates, emphasizing the need for further epidemiologic studies addressed to evaluate the impact of this colonization in puerperal and neonatal morbidity in our geographic region and to identify subpopulations of pregnant women that may benefit from SA screening, given that morbidity and associated costs for MRSA infections in this population are increasingly reported6,8.
We could not find statistical significance among socio demographic factors, colonization status, and development of complications, which could be attributed to the low number of participants able to complete the follow up. Thus, additional studies with more epidemiologic power are needed to ascertain the role of SA colonization in pregnant women from our region.
Antibiotic susceptibility profiles, SCCmec typing, and presence of PVL genes of MRSA isolates points out to their community origin.
Conflict of interest.
The authors declare having no conflict of interest related to this study.Acknowledgements
We are thankful to Javier Escobar from the Laboratory of Bacterial Molecular Genetics at Universidad El Bosque for technical assistance in the multiplex PCR assays; to Enrique Jiménez for collaboration in sample taking, to Alfonso Bettin and Juan Rebollo for assistance in laboratory protocols.
References
1. Peacock SJ, de Silva I, Lowy FD. What determines nasal carriage of Staphylococcus aureus? Trends Microbiol. 2001; 9:605-10. [ Links ]
2. Safdar N, Bradley EA. The risk of infection after nasal colonization with Staphylococcus aureus. Am J Med. 2008; 121:310-5. [ Links ]
3. Zetola N, Francis JS, Nuermberger EL, Bishai WR.Community-acquired meticillin-resistant Staphylococcus aureus: an emerging threat. Lancet Infect Dis. 2005; 5: 275-86. [ Links ]
4. Huang YC, Chao AS, Chang SD, et al. Association of Staphylococcus aureus colonization in parturient mothers and their babies. Pediatr Infect Dis J. 2009; 28: 742-4. [ Links ]
5. Reusch M, Ghosh P, Ham C, Klotchko A, Singapuri S, Everett G. Prevalence of MRSA colonization in peripartum mothers and their newborn infants. Scand J Infect Dis. 2008;40: 667-71. [ Links ]
6. Laibl VR, Sheffield JS, Roberts S, McIntire DD, Trevino S, Wendel GD. Clinical presentation of communityacquired methicillin-resistant Staphylococcus aureus in pregnancy. Obstet Gynecol. 2005; 106: 461-5. [ Links ]
7. Chen KT, Campbell H, Borrell LN, Huard RC, Saiman L, Della-Latta P. Predictors and outcomes for pregnant women with vaginal-rectal carriage of communityassociated methicillin-resistant Staphylococcus aureus. Am J Perinatol. 2007; 24: 235-40. [ Links ]
8. Stumpf PG, Flores M, Murillo J. Serious postpartum infection due to MRSA in an asymptomatic carrier: case report and review. Am J Perinatol. 2008; 25: 413-5. [ Links ]
9. Beigi R, Hanrahan J. Staphylococcus aureus and MRSA colonization rates among gravidas admitted to labor and delivery: a pilot study. Infect Dis Obstet Gynecol. 2007; 2007: 70876. [ Links ]
10. Chen KT, Huard RC, Della-Latta P, Saiman L. Prevalence of methicillin-sensitive and methicillin-resistant Staphylococcus aureus in pregnant women. Obstet Gynecol. 2006; 108: 482-7. [ Links ]
11. Andrews WW, Schelonka R, Waites K, Stamm A, Cliver SP, Moser S. Genital tract methicillin-resistant Staphylococcus aureus: risk of vertical transmission in pregnant women. Obstet Gynecol. 2008; 111: 113-8. [ Links ]
12. Bourgeois-Nicolaos N, Lucet JC, Daubié C, et al. Maternal vaginal colonisation by Staphylococcus aureus and newborn acquisition at delivery. Paediatr Perinat Epidemiol. 2010; 24: 488-91. [ Links ]
13. Pinter DM, Mandel J, Hulten KG, Minkoff H, Tosi MF. Maternal-infant perinatal transmission of methicillinresistant and methicillin-sensitive Staphylococcus aureus. Am J Perinatol. 2009; 26: 145-51. [ Links ]
14. Dehority W, Wang E, Vernon PS, Lee C, Perdreau-Remington F, Bradley J. Community-associated methicillinresistant Staphylococcus aureus necrotizing fasciitis in a neonate. Pediatr Infect Dis J. 2006; 25: 1080-1. [ Links ]
15. Deurenberg RH, Stobberingh EE. The molecular evolution of hospitaland community-associated methicillin-resistant Staphylococcus aureus. Curr Mol Med. 2009; 9: 100-15. [ Links ]
16. Diep BA, Otto M. The role of virulence determinants in community-associated MRSA pathogenesis. Trends Microbiol. 2008; 16: 361-9. [ Links ]
17. Bettin A, Suárez P, Bedoya A, Reyes N. Staphylococcus aureus in residents from a nursing-home in Cartagena. Rev Salud Publica. 2008; 10: 650-7. [ Links ]
18. Millar BC, Jiru X, Moore JE, Earle JA. A simple and sensitive method to extract bacterial, yeast and fungal DNA from blood culture material. J Microbiol Methods. 2000; 42: 139-47. [ Links ]
19. Zhang K, McClure JA, Elsayed S, Louie T, Conly JM. Novel multiplex PCR assay for characterization and concomitant subtyping of staphylococcal cassette chromosome mec types I to V in methicillin-resistant Staphylococcus aureus. J Clin Microbiol. 2005; 43: 5026-33. [ Links ]
20. Brakstad OG, Aasbakk K, Maeland JA. Detection of Staphylococcus aureus by polymerase chain reaction amplification of the nuc gene. J Clin Microbiol. 1992; 30: 1654-60. [ Links ]
21. Lina G, Piemont Y, Godail-Gamot F, et al. Involvement of Panton-Valentine leukocidin-producing Staphylococcus aureus in primary skin infections and pneumonia. Clin Infect Dis. 1999; 29: 1128-32. [ Links ]
22. Olarte NM, Valderrama IA, Reyes KR, et al. Methicillinresistant Staphylococcus aureus colonization in a Colombian hospital intensive care unit: phenotypic and molecular characterization. Biomedica. 2010; 30: 353-61. [ Links ]
23. Castro-Orozco R, Villafane-Ferrer LM, Alvarez-Rivera E, De Arco MM, Rambaut-Donado CL, Vitola-Heins GV. Methicillin-resistant Staphylococcus aureus in children attending school in Cartagena, Colombia. Rev Salud Publica. 2010; 12: 454-63. [ Links ]
24. Gray J, Patwardhan SC, Martin W. Meticillin-resistant Staphylococcus aureus screening in obstetrics: a review. J Hosp Infect. 2010; 75: 89-92. [ Links ]
25. Beigi RH, Bunge K, Song Y, Lee BY. Epidemiologic and economic effect of methicillin-resistant Staphylococcus aureus in obstetrics. Obstet Gynecol. 2009; 113: 983-91. [ Links ]